What are Silent mutations?
Silent mutations are mutations where the changes in the nucleotide sequence of DNA do not produce any observable effect on the organism.
- The change in nucleotide doesn’t produce a change in the amino acid sequence or the structure and function of the protein.
- Silent mutations are also termed neutral mutations as these do not cause any alteration in the protein product formed.
- Silent mutations are possible due to the fact that multiple codon sequences can code for the same amino acid. As a result, in some cases, the change in nucleotide can still produce the same amino acid.
- Silent mutations are also called synonymous mutations, but all synonymous mutations are not silent. Synonymous mutations can affect transcription, splicing, mRNA transport, and translation, all of which can bring about changes in the phenotype of the organism.
- As silent mutations do not affect the final protein product, these are considered evolutionarily neutral.
- However, it has been studied that organisms usually exhibit codon usage biases where they select a particular codon over the others as a result of translational stability. Thus, silent mutations cannot be considered completely neutral.
- Some silent mutations might exhibit effects when present in homozygous forms. Thus, a mutation is silent in the heterozygous form but produces a considerable effect when converted into the homozygous form.
Causes of Silent Mutation
- Mutations can be caused by a number of different reasons that can range from spontaneous occurring as mistakes during DNA processing or induced by the presence of physical agents.
- Spontaneous mutations occur during the processing of DNA, like DNA replication. The changes can occur due to some mistakes or due to digestion by enzymes like nucleases. In some cases, these mutations can also occur during the repair of the damaged DNA sequence.
- The induced silent mutation occurs when physical agents like radiation or molecules cause the alteration in nucleotide. These mutagens change one nucleotide into another in a way that doesn’t affect the final product.
- In the case of silent mutation, the change in codons doesn’t change the amino acid as both the original codon and the altered codon code for the same amino acid.
- Similarly, some mutations might occur at a point that doesn’t affect the structure and function of the protein, rendering the mutation silent.
- There have been instances where a different amino acid is produced, but it is similar to the original amino acid in its structure and function. As a result, the mutation doesn’t affect the final product, and it can be considered silent.
Mechanism of Silent Mutation
- Silent mutations involve the same mechanism as other mutations where the change in nucleotide sequence occurs as a result of tautomerism or ionization.
- The mechanism of spontaneous mutation is not completely understood, however, it is believed that the mutation occurs either due to the deletion of a nucleotide base or due to changes in the structure and bonding of the base.
- The common mechanism of silent mutation is tautomerism, where the chemical form of the nucleotide changes between keto form and enol form.
- Most of the nucleotides exist in keto form and form hydrogen bonds with other nucleotides. When the keto form is converted to the enol form, the molecule can no longer form a hydrogen bond with other nucleotides. As a result, the nucleotides cannot function to form a complete DNA sequence.
- Besides, some changes can also be observed as a result of ionization in the presence of ionizing radiation like X-ray and UV.
- Some mistakes can occur during replication, where digestive enzymes like nucleases might digest a certain portion of the DNA sequence.
- The change in nucleotides in silent mutation is subtle and doesn’t result in a completely different amino acid sequence.
- The amino acid translated is either the same or different with similar characteristics so that no observable effect is observed in the protein.
Applications of Silent Mutation
The following are some of the applications of silent mutation;
- Even though silent mutations do not produce an observable effect in the resulting protein, the organisms tend to develop codon biases for different codons, indicating a possible evolutionary effect.
- Silent mutations can be used to study the effect of different mutagens on different DNA sequences.
- The use of silent mutation is also essential for the study of different codon and amino acids they code for.
- Silent mutations can be carried from one generation to another in the heterozygous form, where they exhibit no result. However, when present in a homozygous form, these can exhibit a considerable change in the amino acid sequence, affecting the structure and function of the protein.
- Silent mutations can also be used to study the properties of genes and the effects of different processes on the DNA segment.
Examples of Silent Mutation
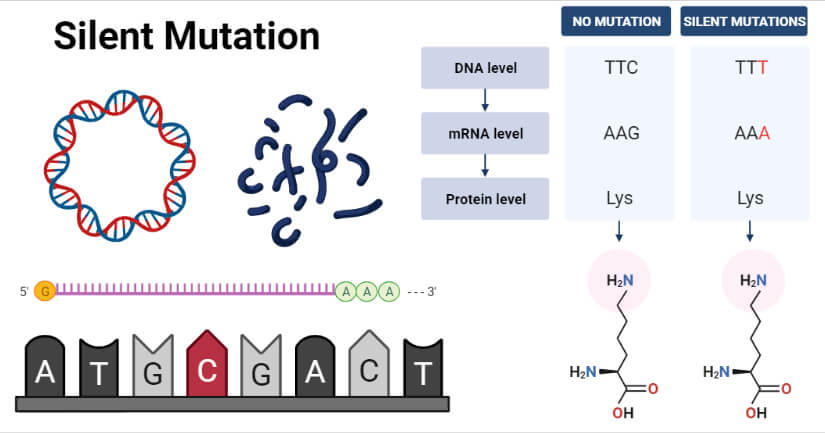
- An example of silent mutation is observed in the case where a thymine nucleotide is replaced by cytosine in a TTC codon, resulting in the formation of a TTT codon.
- The mRNA of the codons are AAG and AAA, respectively. However, both of these mRNAs code for the same amino acid, lysine.
- Thus, the point mutation within a sequence can change the codon, but the resulting amino acid remains the same. In this case, the mutation is silent as it brings no effect on the final product.
- This is possible due to the fact that multiple codons can code for the same amino acids, i.e. degeneracy of the codons.
References
- Lai, Chih-Cheng et al. “The clinical significance of silent mutations with respect to ciprofloxacin resistance in MRSA.” Infection and drug resistance vol. 11 681-687. 3 May. 2018, doi:10.2147/IDR.S159455
- Bali, Vedrana, and Zsuzsanna Bebok. “Decoding mechanisms by which silent codon changes influence protein biogenesis and function.” The international journal of biochemistry & cell biology vol. 64 (2015): 58-74. doi:10.1016/j.biocel.2015.03.011
- Patel, Unnatiben Rajeshbhai et al. “Unraveling the Role of Silent Mutation in the ω-Subunit of Escherichia coli RNA Polymerase: Structure Transition Inhibits Transcription.” ACS omega vol. 4,18 17714-17725. 14 Oct. 2019, doi:10.1021/acsomega.9b02103
- Kashiwagi, Akiko et al. “Contribution of silent mutations to thermal adaptation of RNA bacteriophage Qβ.” Journal of virology vol. 88,19 (2014): 11459-68. doi:10.1128/JVI.01127-14
- Petry F, Loos M. Common silent mutations in all types of hereditary complement C1q deficiencies. Immunogenetics. 2005 Sep;57(8):566-71. doi: 10.1007/s00251-005-0023-z. Epub 2005 Sep 29. PMID: 16086173.
