DNA transcription, also known as RNA synthesis is the process by which genetic information that is contained in DNA is re-written into messenger RNA (mRNA) by an RNA polymerase enzyme.
- The synthesized mRNA is transported out of the cell nucleus where it will later on aid in the synthesis of proteins by the mechanism of translation.
- Regulation of mRNA production in the nucleus, the cell automatically regulated the rate of gene expression.
- The process of transcription is aided by the RNA polymerase enzyme, which copies the right sequences on the DNA template to produce a complementary RNA copy of the gene.
- The basic mechanism of transcription is the same in both eukaryotes and prokaryotes, however, it may differ in a number of ways between them.
DNA Transcription Enzymes and Function
The major enzyme used in DNA transcription is RNA polymerase. In prokaryotes, one type of RNA polymerase enzyme is used, while in eukaryotes, three types of RNA polymerases are used i.e RNA polymerase I, II, and III.
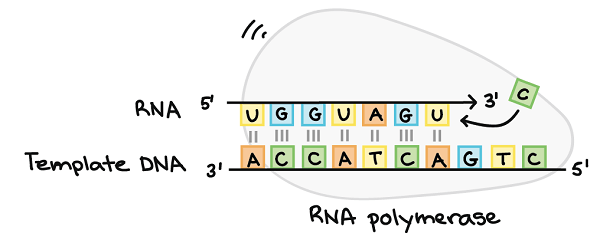
Figure: RNA Polymerase. Image Source: Khan Academy.
The major functions of RNA polymerase include:
- Formation of the initiator complex which aids in the unwinding process of the double-helical structure of DNA
- Synthesis and elongation of the RNA transcript by adding the nucleotide bases, Adenine (A), Cytosine (C), Guanine (G), and Uracil (U).
- It forms the termination sequences that stop and terminate transcription.
Know More: RNA polymerase- Definition, Types, and Functions
DNA Transcription Steps
Summary: Steps of Transcription
- 50 different protein transcription factors will bind to the promoter sites, on the 5′ side of the gene to be transcribed.
- The RNA polymerase binds to the transcription factor complex, allowing the double helix of DNA to open up.
- The RNA polymerase then reads one strand in the 3′ to 5; direction
- In eukaryotic cells, the nucleosome in the advancing RNA polymerase (Pol II), for protein-coding genes.
- The RNA polymerase and transcription factors complex replaces the nucleosome after DNA has been transcribed and Pol II has moved on.
- AS the RNA polymerase moves along the DNA strand, it assembles ribonucleotides into a strand of RNA, by utilization of triphosphate (ATP)
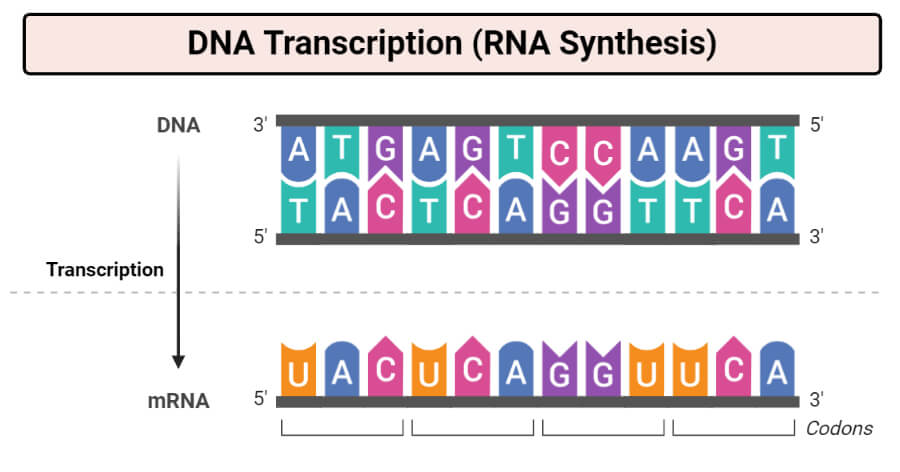
- Each ribonucleotide is inserted into the growing RNA strand by a base-pairing i.e each cytosine (C) is linked to guanine (G), while a Uracil(U) is linked to an Adenine (A).
- The RNA synthesis takes place in the 5′-3′ direction
- As each nucleoside triphosphate is added to the 3′ end of the growing strand, the two terminal phosphates are removed.
- When transcription comes to an end, the transcript is released from the polymerase, and the polymerase enzyme is also released from the DNA
DNA Transcription Process
- In prokaryotic cells, the entire mechanism of transcription is summarized in three stages: Initiation, Elongation, and Termination.
- At the end of the termination in prokaryotes, the mRNA formed is ready for translation.
- Unlike in eukaryotes, after termination, an immature mRNA is formed, and therefore, more processes are needed to form a mature mRNA which is then translated into proteins.
- Generally, the transcription process transcribes DNA into mRNA, the type of RNA that carries the information that is needed in the synthesis of proteins.
- In eukaryotes, there are two broad steps that take place in transcription;
- Pre-messenger RNA formation using an RNA polymerase enzyme
- Editing of pre-messenger RNA by splicing
- Pre-messenger RNA formation involves the initiation, elongation, and termination phases which end by forming mRNA.
- The mRNA then undergoes different stages of splicing to form mature mRNA.
Formation of pre-messenger RNA
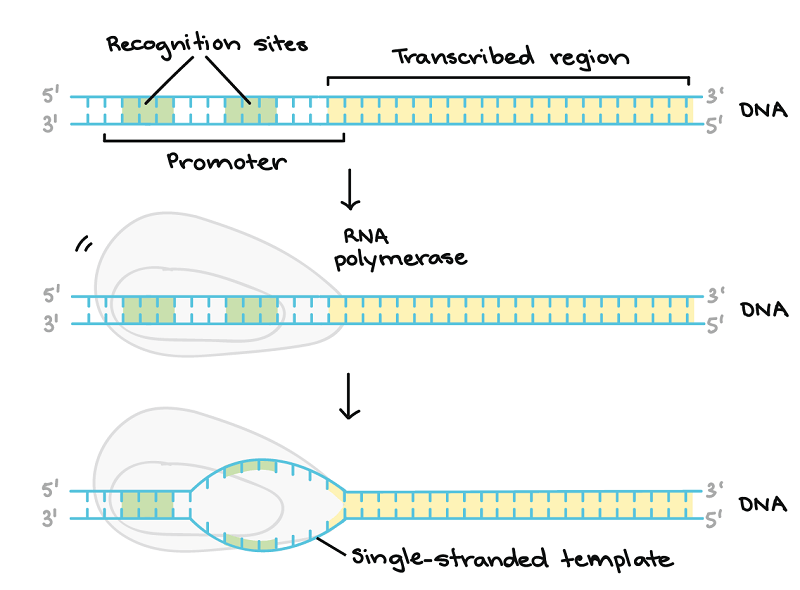
Figure: DNA Transcription Initiation. Image Source: Khan Academy.
- When transcription starts, DNA must unwind, aided by RNA polymerase, which catalyzes the process.
- In transcription only one of the DNA strands is transcribed, the strand that has the initiator sequence. this strand is known as the sense strand, while the complementary strand is known as the antisense strand.
- mRNA that is transcribed is normally a copy of the sense strand, however, it is the antisense strand that is transcribed.
- The ribonucleoside triphosphate (NTPs) aligns along the antisense DNA strand by base pairing, then the RNA polymerase joins the ribonucleotides together forming a pre-messenger RNA molecule, complementary to a region on the antisense strand.
- Transcription is completed when the RNA polymerase enzyme finds a triplet of bases read as a stop signal. AT this stage, the DNA molecule rewinds to reform the double helix.
- The formation of the messenger RNA (mRNA) is done in three stages: Initiation, elongation, and termination
Initiation
Promoter and initiation in prokaryotes
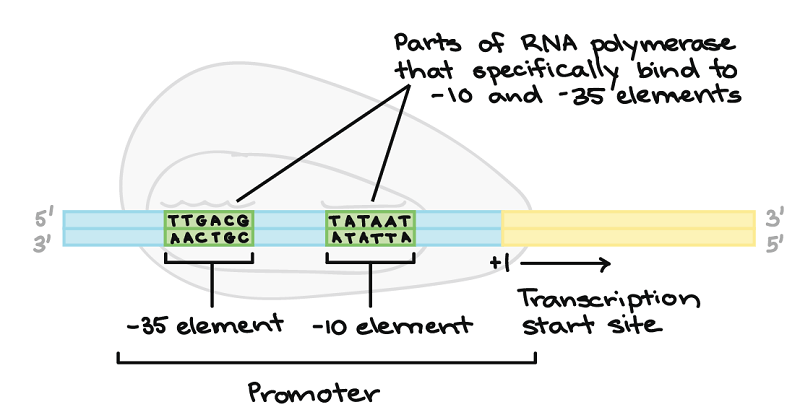
Figure: Promoter and initiation in prokaryotes. Image Source: Khan Academy.
- The initiation of transcription is signaled at a region known as a promoter.
- The promoter is the site for RNA polymerase binding, such that the promoter guides the polymerase where it should sit on the DNA in order to initiate transcription.
- RNA polymerase is the enzyme that catalyzes the mechanism of transcription.
- The RNA polymerase enzyme has a sigma (σ) factor, which is the dissociative unit, that allows the enzyme to recognize the promoter sequence (starting point of transcription), which is spaced between -35 and -10 regions.
- The promoter sequence is recognized by RNA polymerase’s holoenzyme subunit, by attaching to and moving along the DNA template molecule. This forms a closed promoter complex.
- A single DNA molecule may have multiple promoter sequences or closed promoter complexes.
- The promoter which is bound with transcription factors along with the RNA polymerase forms a complex.
- The transcription factors are regulatory proteins that control transcription rate.
- When the RNA polymerase is bound to the promoter sequence, it denaturalizes the DNA duplex locally, forming open promoter complex which becomes the unwound part of the double-stranded DNA, exposing the bases on each of the two DNA strands.
Promoter and initiation in Eukaryotes
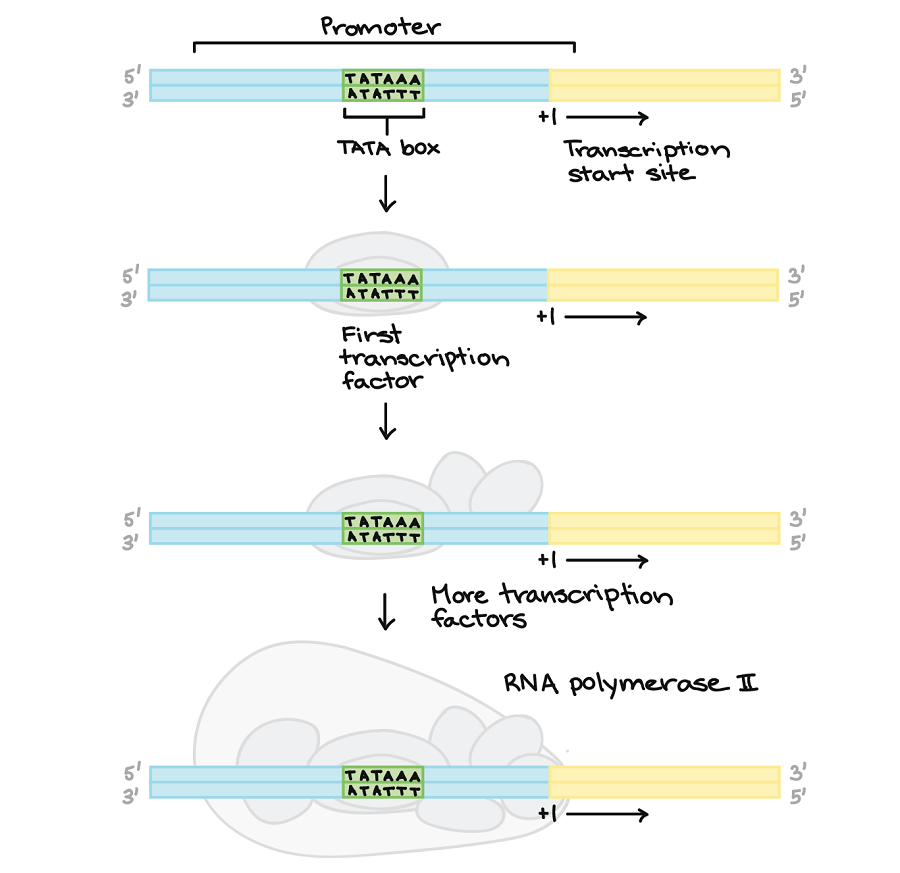
Figure: Promoter and initiation in Eukaryotes. Image Source: Khan Academy.
- In eukaryotes, the RNA polymerase does not directly attach to the promoter sequence like in prokaryotes.
- A helper promoter known as a basal (general) transcription factor binds to the promoter first, which helps the RNA polymerase attach to the DNA template.
- Eukaryotes have a promoter sequence called a TATA box, which is recognized by the transcription factors, which eventually allow the binding of the RNA polymerase.
- The TATA box has lots of As and Ts making it easy to pull the strands of DNA apart.
Elongation
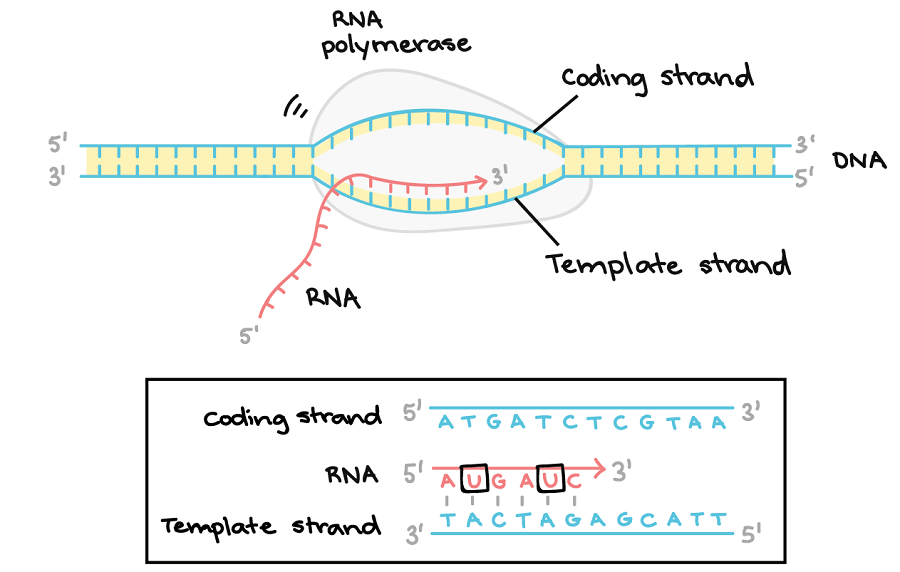
Figure: DNA Transcription Elongation. Image Source: Khan Academy.
- After initiating transcription, the sigma (σ) factor dissociates from the RNA polymerase.
- The template strand is read in the 3′ to 5′ direction, which means that RNA synthesis takes place in the 5′ to 3′ direction, with the nucleoside triphosphate (NTPs) acting as substrates for the enzyme.
- The other strand from the DNA template is known as the coding strand, because the base sequence of the new mRNA is identical to it, except for the replacement of thiamine with uracil base.
- The RNA polymerase catalyzes the formation of a phosphodiester bond between the adjacent ribonucleotides.
- The energy used by the RNA polymerase is derived from splitting the high-energy triphosphate into monophosphate, releasing the inorganic diphosphates (PPi).
- A transcription bubble is formed and it must be maintained sine transcription takes place on the double-stranded DNA template. The bubble moves along the DNA duplex during elongation.
- Stalling or pausing are common, which are later essential for transcription termination.
Termination
- This is the process of ending transcription, which happens when signaled by a stop sequence known as a terminator sequence.
- This happens when the RNA polymerase transcribes the terminator sequence.
- The RNA polymerase then releases the DNA temple which unwinds back to a double-helical structure.
Termination in bacteria
There are two termination methods in bacteria
Rho-dependent termination
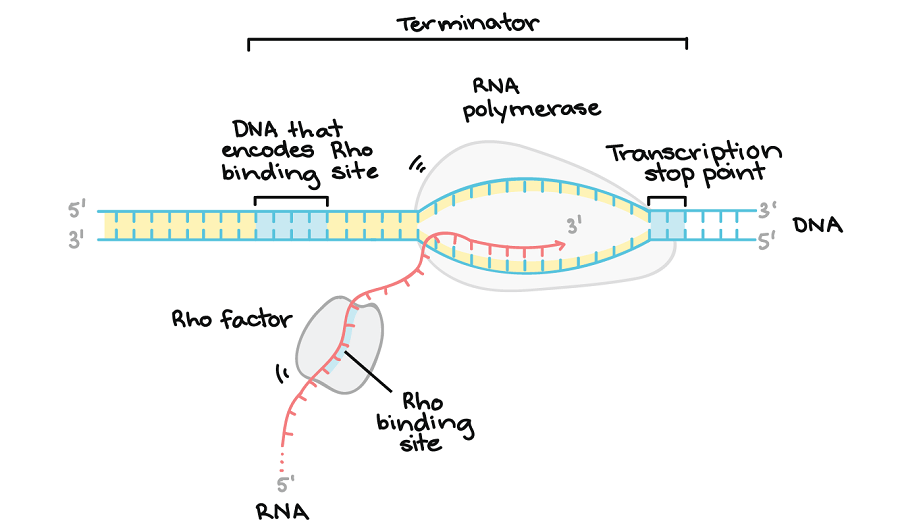
Figure: Rho-dependent termination. Image Source: Khan Academy.
This is the termination process where the RNA molecule contains a binding site for a protein known as the Rho factor, which binds to the DNA sequence. It starts to climb up the transcript towards the RNA polymerase ad reaches the transcription bubble. At the bubble, the Rho factor pulls the RNA transcript and the DNA template strand apart, releasing the RNA molecule and terminating the transcription process. A transcription stop point sequence that is found later in the DNA causes the RNA polymerase to stop and allow the Rho factor to catch up and terminate the process.
Rho-independent termination
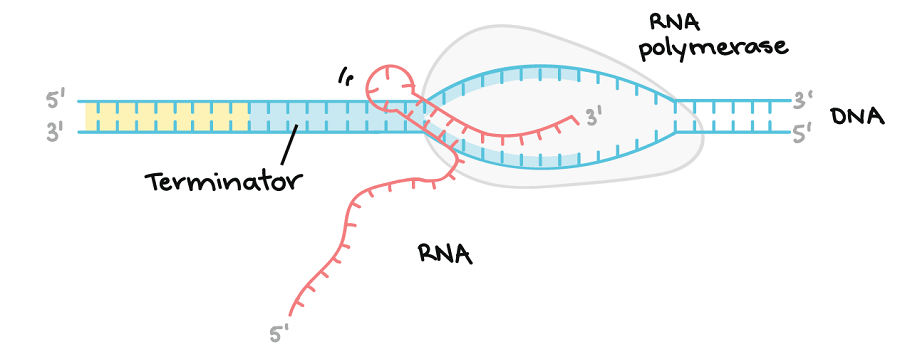
Figure: Rho-independent termination. Image Source: Khan Academy.
This process depends on a specific sequence found on the DNA template strand. During transcription, as the RNA polymerase approaches the endpoint of the gen that is transcribed, it reaches a region that is rich in Cytosine (C) and Guanine(G). The RNA that is transcribed from this region folds back on itself, and the complementary C and G bind together forming a stable hairpin that makes the RNA polymerase to stall. The hairpin is followed by a Uracil (U) in the RNA terminator which complementary to the DNA template Adenine (A). The U-A region forms a weak interaction with the DNA template and with the stalled RNA polymerase causes an instability allowing the enzyme to fall off and end from the new RNA transcript.
Video Lecture: DNA Transcription and mRNA processing (Video By Khan Academy)
Pre-translational mRNA processing
- In eukaryotes, the mRNA that has been transcribed is known as pre-mRNA, and therefore, it must undergo other processes for it to mature into mature mRNA.
- These are known as the pre-translational mRNA processes. They include:
5′ Capping
- This is the addition of methylated guanine cap to the 5′ end of mRNA
- The 5′ cap helps in the recognition of the mRNA molecule by the ribosomes, and to also protect the immature mRNA from the degradation of RNases.
Polyadenylation
- This is the addition of a poly (A) tail to the 3′ end of mRNA. The poly (A) tail is made up of several molecules of adenosine monophosphate, which stabilizes RNA, because of its natural instability.
Splicing
- This is the coding of one genetic sequence for different proteins, conserving the genetic material.
- The process involves:
- Removing of the non-coding sequences known as introns by spliceosome excision.
- Joining of the coding sequences known as exons by ligation.
- Splicing is sequence-dependent, therefore it occurs within the transcript.
- This allows many proteins to be made from a single pre-mRNA
- At the end of the splicing process, mature mRNA will have been made.
- Mature mRNA then becomes the messenger carrier which allows protein synthesis to occur.
- Mature mRNA has open Reading Frames (ORF), a region that gets translated into proteins. Translation on the ORFs is done in three blocks of three nucleotides known as codons.
- At the end of the 5′ and 3′ ends are untranslated regions (UTRs) which are not translated during protein synthesis.
DNA Transcription in Eukaryotes (Difference from prokaryotes)
Transcription in eukaryotes and prokaryotes have some similarities and differences.
Similarities
Some of the common similarities include;
- DNA is used as the template in both organisms
- RNA polymerase is the main enzyme that facilitates the entire mechanism in both organisms
- The RNA molecule is the end product in both organisms
- The chemical composition of the transcript is the same in both organisms
Differences
The major differences in the transcription between eukaryotes and prokaryotes include:
Promoter:
- Prokaryotes have three promoter elements i.e -10, -35 promoters, and upstream elements
- Eukaryotes have many different promoter elements i.e TATA box, initiator elements, downstream core promoter, CAAT box, and the CG box
RNA polymerase:
- Prokaryotes have one type of RNA polymerase that aids in the synthesis of the RNA strand.
- Eukaryotes have three types of RNA polymerases I, II, and III that aid in the RNA strand synthesis.
Initiation:
- The initiation complex of eukaryotes is made up of various transcription factors that dissociate on completion of the initiation process.
- While prokaryotes do not form an initiator complex
Transcription and translation concurrence
- Another major difference is that, in prokaryotes, transcription and translation occur simultaneously while in eukaryotes, transcriptions must be complete before the translation mechanism is initiated.
- The RNA in eukaryotes undergoes post-transcriptional modifications like capping, polyadenylation, and splicing to form a mature mRNA that proceeds to translation. These processes do not take place in prokaryotes
RNA genetics
- The mRNA in prokaryotes has many different genes on the single mRNA hence they are termed as polycistronic
- Eukaryotes have a single gene on one mRNA molecule hence termed as monocistronic.
Termination
- Termination in prokaryotes is aided by either Rho-dependent or Rho-independent factors, while in eukaryotes, transcription is terminated by the poly-adenylated (A) signal and the downstream terminator sequence.
Reverse Transcription
- This is the conversion of RNA to DNA, by which RNA acts as the template in the synthesis of a type of DNA known as the complementary DNA (cDNA).
- The central dogma defines the mechanisms that involve DNA synthesis (replication), RNA synthesis (transcription), protein synthesis (translation), and cDNA synthesis (reverse transcription). Therefore, DNA codes for RNA, RNA codes for proteins, and RNA can also code for DNA in the case of reverse transcription.
- RNA-encoded viruses undergo the reverse transcription mechanism where their genomic RNA is converted to DNA by the aid of reverse transcriptase enzyme.
- Reverse transcription is also known as retrotranscription or retrotras. It is an extremely erroneous process that can lead to mutations that can cause drug resistance.
- The conversion of RNA to DNA is normally desirably applied in the laboratory majorly as a diagnostic tool for most RNA viruses such as HIV, hepatitis, influenza, coronaviruses, etc
Transcription Inhibitors
Transcription inhibitors are elements that are used to inhibit the action and mechanism of the RNA polymerase enzyme, which hinders the process of transcription. Transcription inhibitors are used majorly to hinder bacterial mechanisms of transcription in disease-causing pathogens. Some of the most commonly used inhibitors include
- α-amanitin – This is an inhibitor extracted from yeast, that is selective for RNA polymerase II and RNA polymerase III.
- Rifampicin inhibits bacterial transcription by inhibiting DNA-dependent RNA polymerase by binding to the beta-subunit.
- 8-hydroxyquinoline is also an antifungal transcription inhibitor
- Others include actinomycin D, CDK9 inhibitors such as DRB, and flavopiridol, triptolide.
- An inhibitory mechanism is histone methylation that bars the action of transcription.
Reference and Sources
- 3% – https://www.slideshare.net/ammara12/transcription-81803017
- 1% – https://www.studocu.com/en-au/document/the-university-of-adelaide/biology-i-molecules-genes-and-cells/lecture-notes/term-2-lecture-notes/7577176/view
- 1% – https://www.ncbi.nlm.nih.gov/books/NBK21230/
- 1% – https://www.chem.uwec.edu/webpapers2006/sites/demlba/folder/ProvsEuk.html
- 1% – https://quizlet.com/25761894/dna-replication-flash-cards/
- 1% – https://bio.libretexts.org/Bookshelves/Introductory_and_General_Biology/Book%3A_Biology_(Kimball)/06%3A_Gene_Expression/6.02%3A_The_Transcription_of_DNA_into_RNA
- 1% – http://www.atdbio.com/content/14/Transcription-Translation-and-Replication
- <1% – https://www.thoughtco.com/protein-synthesis-translation-373400
- <1% – https://www.thoughtco.com/dna-transcription-373398
- <1% – https://www.researchgate.net/publication/327222318_Role_of_DNA_and_RNA_in_Protein_Synthesis
- <1% – https://www.researchgate.net/publication/262053270_mRNA_quality_control_at_the_5′end
- <1% – https://www.researchgate.net/publication/13600431_An_Integrated_Model_of_the_Transcription_Complex_in_Elongation_Termination_and_Editing
- <1% – https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6822018/
- <1% – https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2834840/
- <1% – https://www.nature.com/scitable/topicpage/ribosomes-transcription-and-translation-14120660/
- <1% – https://www.hindawi.com/journals/omcl/2015/536962/
- <1% – https://www.forbes.com/sites/quora/2017/10/18/why-do-genetic-mutations-occur-and-how-can-we-stop-them/
- <1% – https://www.differencebetween.com/difference-between-rna-polymerase-i-and-vs-ii-and-vs-iii/
- <1% – https://www.differencebetween.com/difference-between-prokaryotic-and-vs-eukaryotic-transcription/
- <1% – https://www.differencebetween.com/difference-between-gene-expression-in-prokaryotes-and-vs-eukaryotes/
- <1% – https://www.coursehero.com/file/pnj9f3h/a-Both-DNA-and-RNA-are-synthesized-in-a-5-to-3-direction-b-During-RNA-synthesis/
- <1% – https://www.chegg.com/homework-help/questions-and-answers/sigma-subunit-bacterial-rna-polymerase-recognizes-promoter-sites-dna-contains-catalytic-ac-q41121720
- <1% – https://www.biologydiscussion.com/rna/transcription/transcription-in-prokaryotes-and-eukaryotes-with-diagram/15546
- <1% – https://uk.answers.yahoo.com/question/index?qid=20120510125017AAHW4O4
- <1% – https://study.com/academy/lesson/inhibitors-of-dna-rna-synthesis-how-rifamycins-and-quinolone-kill-bacteria.html
- <1% – https://study.com/academy/answer/what-is-the-product-of-transcription-a-dna-b-genes-c-protein-d-rna-e-all-of-these-are-correct.html
- <1% – https://quizlet.com/87117135/biology-chapter-12-flash-cards/
- <1% – https://quizlet.com/166453550/chapter-7-bio-100-flash-cards/
- <1% – https://pediaa.com/difference-between-prokaryotic-and-eukaryotic-rna-polymerase/
- <1% – https://idoc.pub/documents/molecular-cell-biology-8th-edition-harvey-lodish2120wwwebook-dlcom-en5kw0090xno
- <1% – https://ibiologia.com/dna-transcription/
- <1% – https://en.wikipedia.org/wiki/Transcription_bubble
- <1% – https://en.wikipedia.org/wiki/Protein
- <1% – https://en.wikipedia.org/wiki/MRNA
- <1% – https://en.wikipedia.org/wiki/Eukaryotic_translation
- <1% – https://en.wikipedia.org/wiki/DNA-dependent_RNA_polymerases
- <1% – https://biology.stackexchange.com/questions/15082/what-does-5-and-3-mean-in-dna-and-rna-strands
- <1% – https://bio.libretexts.org/Bookshelves/Introductory_and_General_Biology/Book%3A_General_Biology_(Boundless)/15%3A_Genes_and_Proteins/15.3%3A_Eukaryotic_Transcription/15.3A%3A_Initiation_of_Transcription_in_Eukaryotes
- <1% – https://bio.libretexts.org/Bookshelves/Genetics/Book%3A_Working_with_Molecular_Genetics_(Hardison)/Unit_III%3A_The_Pathway_of_Gene_Expression/10%3A_Transcription%3A_RNA_polymerases
- <1% – https://answers.yahoo.com/question/index?qid=20130205132932AAxpXWJ
- <1% – http://www.gs.washington.edu/academics/courses/braun/55106/readings/hahn.pdf
- <1% – http://genesdev.cshlp.org/content/11/21/2755.full.html
- <1% – http://chemguide.co.uk/organicprops/aminoacids/dna3.html

ı think there is wrong info about end of the termination right under the DNA transcription process title and it is Unlike in eukaryotes, after termination, an immature mRNA is formed, and therefore, more processes are needed to form a mature mRNA which is then translated into proteins sentences. sorry about my English skill
Thank you, we will look into this.
I agree with you