Polymerase chain reaction (PCR) is a temperature-dependent nucleic acid amplification technique used to amplify the DNA or RNA in vitro enzymatically.
Developed by Kary Mullis and his associates at mid – 1980s, it is a very powerful and most important tool in modern biology – molecular biology and genetics. It combines the principle of nucleic acid hybridization with the principle of nucleic acid replication. Using this non-culture-based nucleic acid amplification technique, we can produce billions of copies of a single segment of DNA or RNA in a very short time.
Since its development, several modifications have been made, and now there are different types of PCR techniques available for different purposes. Reverse transcriptase PCR and Quantitative PCR (qPCR) are the most commonly used PCR types.
Reverse Transcription PCR (RT-PCR), is a type of PCR technique that enzymatically amplifies the RNA in vitro.
It is the only type of PCR that can amplify the RNA. It uses a reverse transcriptase enzyme in addition to the other basic components of the PCR.
First, the sample RNA is converted to complementary DNA (cDNA) in reverse transcription, catalyzed by the reverse transcriptase enzyme. These cDNA molecules are then used as a template for amplification in the PCR process.
RT-PCR is used to analyze the mRNA or micro RNA and study gene expression.
Objectives of Reverse Transcription PCR (RT-PCR)
- To amplify the specific segment of RNA, resulting in billions of copies of a single RNA segment.
- To diagnose certain infections, genes, and study gene expression.
Principle of Reverse Transcription PCR (RT-PCR)
RT-PCR combines the reverse transcription process with the conventional PCR process. The sample RNA is first converted to double-stranded DNA (complementary DNA) by reverse transcriptase enzyme in the reverse transcription process. The cDNA can then be thermally broken down into two single-stranded DNA templates. In these ssDNA templates, primers can anneal to their complementary sequences based on the nucleic acid hybridization principle. DNA polymerase then elongates the primer by sequentially adding the nucleotides to the 3’ end and generates a dsDNA following the principle of DNA replication. These three processes, denaturation, annealing, and elongation, are repeated in a cyclic manner regulating the reaction temperature and resulting in millions of copies of the cDNA.
Requirements (Enzymes) of Reverse Transcription PCR (RT-PCR)
1. Nucleic Acid Sample (Sample RNA)
RNA is the sample for RT-PCR, unlike other PCR techniques using DNA as their sample. Mostly mRNA is used as the sample. The RNA will be converted into cDNA before amplification.
2. Reverse Transcriptase Enzyme
It is an enzyme that catalyzes the formation of complementary DNA (cDNA) strands from the RNA strand. It is also called RNA-dependent DNA polymerase enzyme and is responsible for central-dogma reverse. It is the major component of RT-PCR as it converts sample RNA into cDNA for amplification.
3. DNA Polymerase Enzyme
DNA polymerases are enzymes that catalyze the synthesis of complementary DNA strands by assembling the nucleotides sequentially according to the template strand. Taq DNA polymerase, the DNA polymerase enzyme extracted from the bacterium Thermus aquaticus, is the most widely DNA polymerase as it is thermally stable and continues its activity after the repeated cycle of heating and cooling.
4. Primers (Oligo (dT) primers, random primers, and sequence-specific primers)
Three different types of primers are used in RT-PCR;
- Random Primers
These are the short single-stranded sequences of 6 to 8 nucleotides that bind at the complementary site of RNAs with or without poly(A) for cDNA synthesis using reverse transcriptase.
- Oligo (dT) Primers
They are oligonucleotides, mostly of 12 – 18 nucleotides, containing a segment of repeating deoxythymidine (dT) which binds at the polyA tail of mRNA.
- Sequence-specific Primers
These are the short single-stranded sequences of nucleotides that bind to the specific region of interest of the sample RNA. It is mostly used in one-step RT-PCR.
5. Deoxynucleotide Triphosphates
Deoxynucleotide triphosphates (dNTPs) are artificially synthesized nucleotides that act as building blocks for synthesizing cDNA and new cDNA strands during amplification. 4 different dNTPs are used; deoxyadenosine triphosphate (dATP), deoxyguanosine triphosphate (dGTP), deoxythymidine triphosphate (dTTP), and Deoxycytidine triphosphate (dCTP).
6. PCR Buffers and Other Chemicals
7. Thermocycler (PCR Machine)
Types of Reverse Transcription PCR (RT-PCR)
Based on whether the reverse transcription and the amplification steps occur either in a single reaction (or tube) or in two separate reactions (or tubes), RT- PCR can be classified into two types:
1. One-Step RT-PCR
It is a type of RT – PCR where the reverse transcription and the amplification reactions occur in a single tube. All the required components are added in a single tube. First, reverse transcription occurs, forming cDNA, which is then amplified in a PCR process.
Advantages of One–Step RT – PCR over Two–Step RT – PCR
- It has a simple and easy handling setup.
- It has higher accuracy and specificity.
- It has a lesser chance of contamination.
- It is a cheaper and faster method.
Disadvantages of One-Step RT-PCR over Two-Step RT-PCR
- It detects fewer templates per reaction mixture due to using multiple chemicals in a single reaction tube.
- Due to lower template detection, it requires a larger template for starting.
- It does not permit the storage and further analysis of the cDNA formed during the reaction.
- There is a higher chance of primer – dimer and non–specific binding.
- The chance of reaction failure is comparatively high.
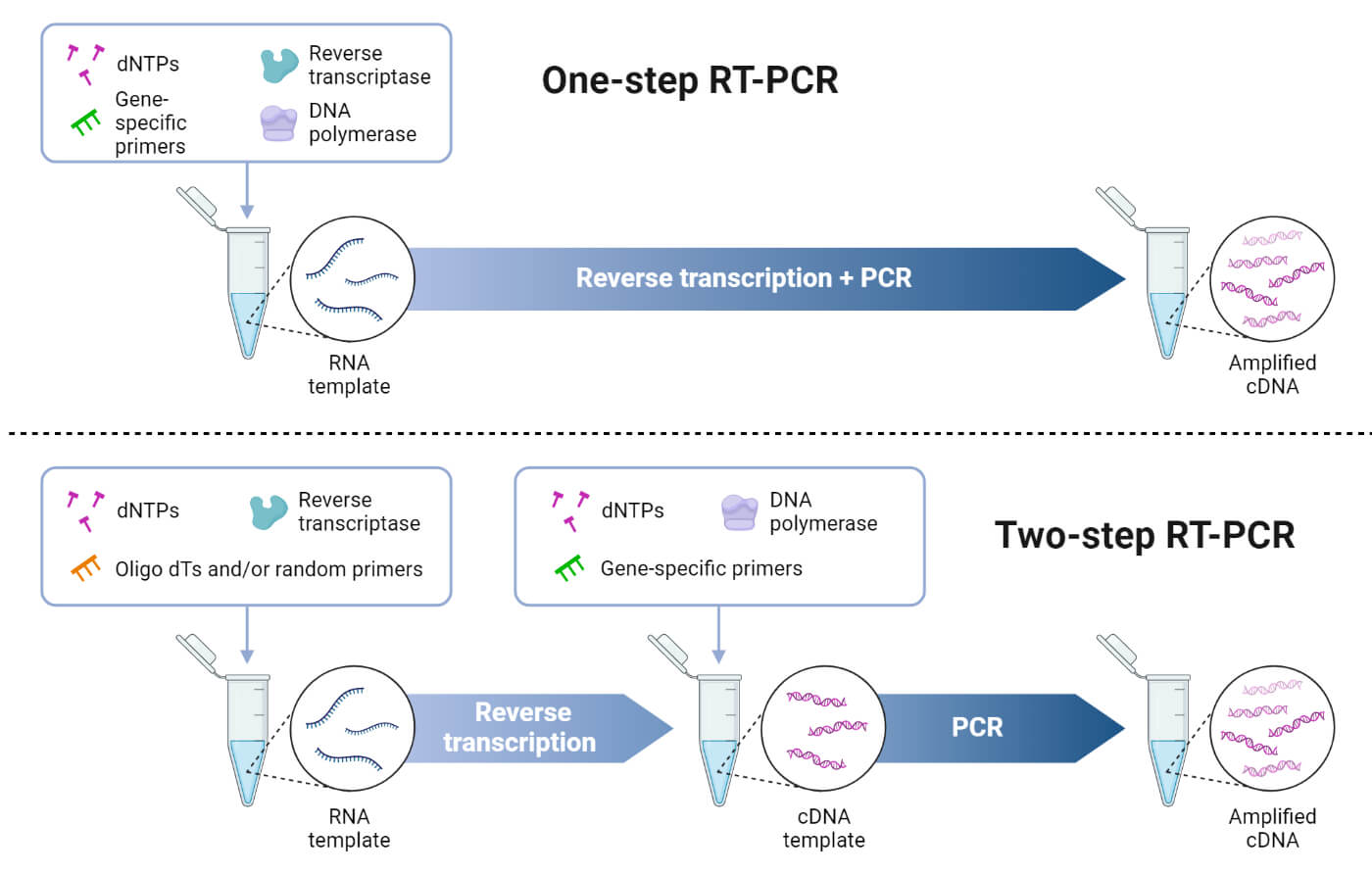
2. Two-Step RT-PCR
It is another type of RT – PCR where the reverse transcription and the amplification process occur in two separate tubes. In the first tube, a reverse transcription reaction takes place, yielding cDNA. These cDNAs are then transferred to another tube where the PCR mixture is added, and the cDNAs are amplified.
Advantages of Two-Step RT – PCR over One-Step RT – PCR
- It allows us to store cDNA formed by reverse transcription.
- It has higher efficiency, accuracy, and reliability and detects larger templates per reaction mixture.
- Comparatively, lower chance of reaction failure, non-specific binding, and primer–dimer bonding.
Disadvantages of Two–Step RT – PCR over One-Step RT – PCR
- There is a higher chance of contamination.
- It is a more complex and tedious process requiring more resources and a well-trained person.
Steps/Procedure of Reverse Transcription PCR (RT-PCR)
The core procedure can be broadly classified into two phases; reverse transcription and amplification. The procedure also varies on one-step and two-step RT – PCR. But, the general steps involved in both of the types are the same and can be summarized into four stages; Preparatory stage, reverse transcription, amplification, and product analysis stage, viz.:
1. Preparatory Stage
It is the initial stage where RNA extraction is done, and all the reaction mixture is prepared. First, all materials are arranged, safety measures are taken, the PCR reaction preparation area is cleaned, all the reagents are brought to working temperature, and the sample is extracted or brought from storage.
In the one-step RT-PCR, sample RNA, reverse transcriptase enzyme, RNase H, primers, DNA polymerase, dNTPs, buffers, and all other components are added in a specified and pre-calculated amount in a single reaction tube. The tube is then loaded into a thermocycler for further processing.
In the two-step, RT-PCR, sample RNA, reverse transcriptase, RNase H, primers, dNTPs, and other buffers and chemicals for reverse transcription are loaded in a tube. Then the tube is subjected to a specified temperature in a thermocycler where cDNAs are formed.
2. Reverse Transcription
It is the primary step where the RNA is converted into cDNA, which then undergoes amplification.
All the reaction mixture, including reverse transcriptase, RNase H, dNTPs mixture, primers, nuclease-free water, reverse transcription buffer, and other components in one-step RT-PCR and DNA polymerase and other amplification components in the two-step RT-PCR are added in a tube and subjected to a temperature of 40 – 50°C for 10 minutes to 30 minutes in a thermocycler. At this temperature, the primer will bind to the respective site of the RNA sample, and the reverse transcriptase enzyme will synthesize cDNA by adding the free dNTPs.
3. Amplification
This step is similar to the amplification process of other PCR techniques for DNA amplification. In a one-step RT-PCR, the same reaction mixture is subjected to an amplification process. At the same time, in the two-step RT-PCR, the cDNA is isolated and placed in another tube where DNA polymerase, primers, PCR buffer, dNTPs, and other chemicals are added. Then the tube is placed in a thermocycler for amplification.
The amplification step includes denaturation, annealing, and elongation occurring cyclically one after another for a certain number of cycles pre-programmed by the user.
4. Product Analysis Stage
It is the final step where the reaction mixture subjected to PCR is analyzed to confirm that desired amplification is achieved. The gel electrophoresis method is mostly used for product analysis. In real-time RT-PCR, there is no need for this additional step.
Applications of Reverse Transcription PCR (RT-PCR)
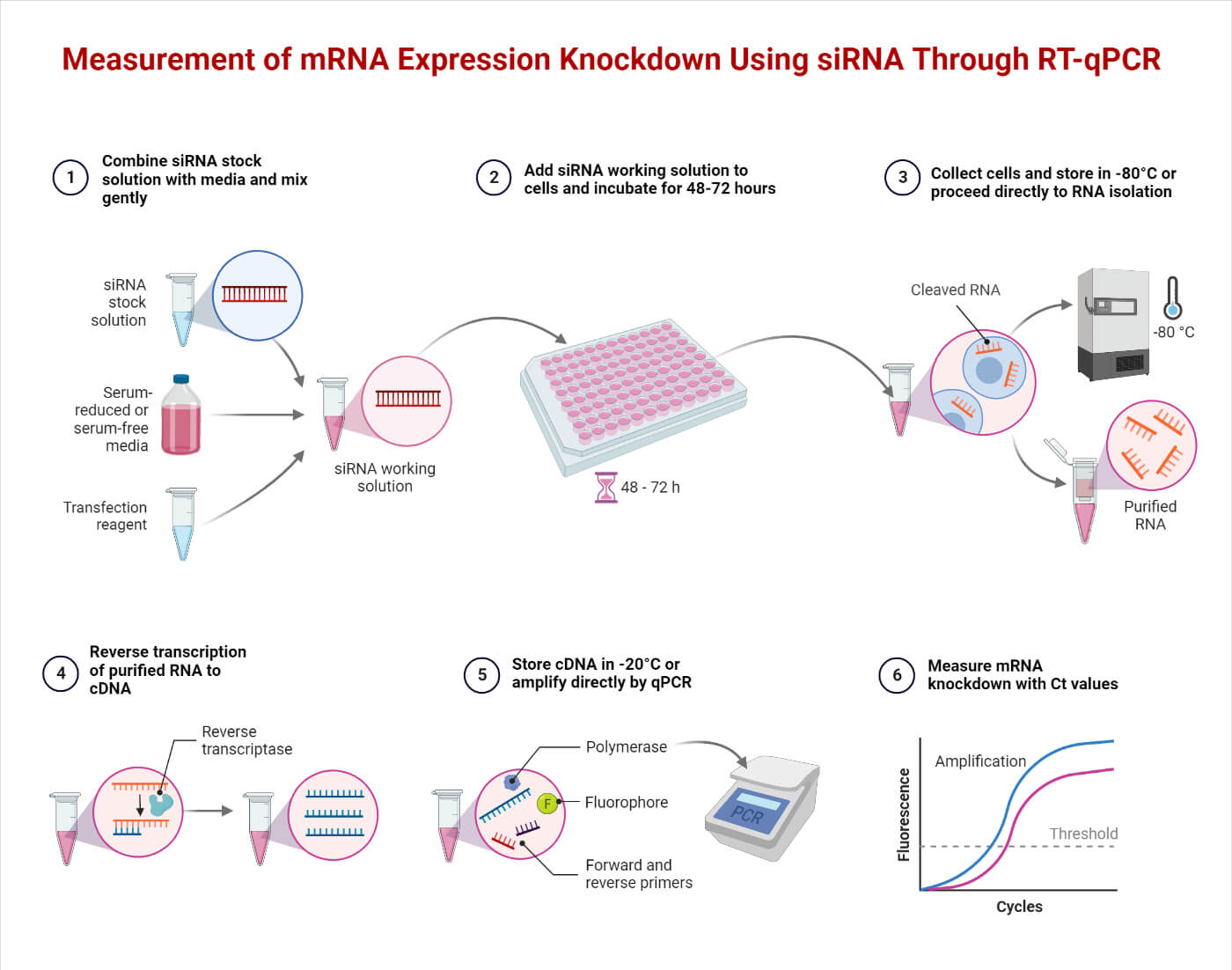
- Study Gene Expression
The Traditional Northern Blot technique requires a larger mRNA sample to analyze and study the gene expression. However, using RT-PCR, we can amplify the minute mRNA sample and study the sequence of nucleotides, thus analyzing the gene expression. It is used in studying and identifying multidrug-resistant genes and their expressions in pathogens.
- Identification of Unknown Species
RT-PCR is used to identify viruses like HIV, SARS viruses, dengue viruses, HCV, etc. Besides, other microorganisms and even higher organisms are identified by studying their rRNA and mRNA.
- Infectious Disease Diagnosis
Diagnosis of different types of viral infection, bacterial infection, fungal and parasite infection, cancer cell, and genetic diseases are done using the RT-PCR technique in clinical laboratories.
- Gene Insertion and Gene Therapy Study
RT-PCR is used to prepare cDNA from eukaryotic mRNA, which lacks introns and can be inserted into prokaryotes. RT-PCR is used in monitoring the result of gene insertion and gene therapy. These procedures are supposed to show particular gene expression and code for a particular protein, hence translating specific types of mRNA sequence. This specific mRNA sequence can be analyzed using RT-PCR.
- Study Mutation and Cancer Cells
RT-PCR can detect and quantify tissue-specific mutant alleles. It can also detect any undesired changes in the mRNA sequence and unique mRNAs, which are produced only by the different types of cancer cells in our body.
- Tools of Genetic Engineering and Viral Study
RT-PCR is used in genetic engineering for analyzing modified DNAs and their transcribed RNAs and amplifying target RNA.
Advantages of Reverse Transcription PCR (RT-PCR)
- It is a very rapid method for amplifying RNA and can enzymatically produce millions of copies of mRNA in a very short time.
- It is very simple to operate. The process is semi-automatic, operated, and regulated by a thermocycler without human involvement.
- It has very high specificity and sensitivity but is economical.
- It is a very accurate method for the identification of RNA viruses and infection by them. RNA viruses can be classified up to the level of strains. It has shortened the time for identifying RNA viruses and viral infections.
- It can detect a very minute amount of mRNA (about 5pg) compared to the traditional Northern Blot technique.
- Mutated genes and gene expression can be easily and promptly studied. This has made it possible to diagnose cancer in the early stage, study gene insertion, and monitor the result of gene therapy.
- It is both a qualitative and quantitative method; hence can be used to identify as well as quantify the sample RNA.
Limitations of Reverse Transcription PCR (RT-PCR)
- It can amplify RNAs only, especially mRNAs.
- Prior information regarding the sequence of the RNA is required for primer designing.
- It is a full temperature and enzyme-based system, so a slight change in the reaction temperature will decrease the efficiency of the enzyme. Hence, require a strict temperature regulation system.
- Slight contamination, having a similar primer binding site, can be amplified, giving a false positive or false negative result.
- The reaction can be highly influenced by a minute amount of organic or inorganic contaminant in the reaction mixture.
- The process is very tedious, requiring a complex reaction mixture and a skilled person to operate.
References
- Santos CF, Sakai VT, Machado MA, Schippers DN, Greene AS. Reverse transcription and Polymerase chain reaction: principles and applications in dentistry. J Appl Oral Sci. 2004 Mar;12(1):1-11. doi: 10.1590/s1678-77572004000100002. PMID: 21365144.
- Montarras, D., Pinset, C., Chelly, J., Kahn, A. (1994). RT-PCR and Gene Expression. In: Mullis, K.B., Ferré, F., Gibbs, R.A. (eds) The Polymerase Chain Reaction. Birkhäuser, Boston, MA. https://doi.org/10.1007/978-1-4612-0257-8_24
- Riedy MC, Timm EA Jr, Stewart CC. Quantitative RT-PCR for measuring gene expression. Biotechniques. 1995 Jan;18(1):70-4, 76. PMID: 7702857.
- Morier, Douglas. “reverse transcriptase”. Encyclopedia Britannica, 23 Nov. 2018, https://www.britannica.com/science/reverse-transcriptase. Accessed 31 August 2022.
- https://toptipbio.com/cdna-synthesis-primers/
- Reverse transcriptase (RT)-PCR: Principles, Applications • Microbe Online
- Reverse Transcription PCR: Principle, Procedure, Protocol, Advantages, Limitations, Applications (geneticeducation.co.in)
- Reverse Transcription System Technical Bulletin TB099 (promega.com)
- The Basics of Reverse Transcription PCR (RT-PCR) (excedr.com)
- PCR (cgiar.org)
- (PDF) The use of reverse transcriptase-polymerase chain reaction (RT-PCR) to investigate specific gene expression in multidrug-resistant cells | Lorraine O’Driscoll – Academia.edu

I love this article
THE BEST EVER