There are more than one codon for one amino acid. This is called degeneracy of genetic code. To explain the possible cause of degeneracy of codons, in 1966, Francis Crick proposed “the Wobble hypothesis”.
According to The Wobble Hypothesis, only the first two bases of the codon have a precise pairing with the bases of the anticodon of tRNA, while the pairing between the third bases of codon and anticodon may Wobble (wobble means to sway or move unsteadily).
The phenomenon permits a single tRNA to recognize more than one codon. Therefore, although there are 61 codons for amino acids, the number of tRNA is far less (around 40) which is due to wobbling.
The Wobble Hypothesis Statement
The wobble hypothesis states that the base at 5′ end of the anticodon is not spatially confined as the other two bases allowing it to form hydrogen bonds with any of several bases located at the 3′ end of a codon.
This leads to the following conclusions:
- The first two bases of the codon make normal (canonical) H-bond pairs with the 2nd and 3rd bases of the anticodon.
- At the remaining position, less stringent rules apply and non-canonical pairing may occur. The wobble hypothesis thus proposes a more flexible set of base-pairing rules at the third position of the codon.
- The relaxed base-pairing requirement, or “wobble,” allows the anticodon of a single form of tRNA to pair with more than one triplet in mRNA.
- The rules: first base U can recognize A or G, first base G can recognize U or C, and first base I can recognize U, C or A.
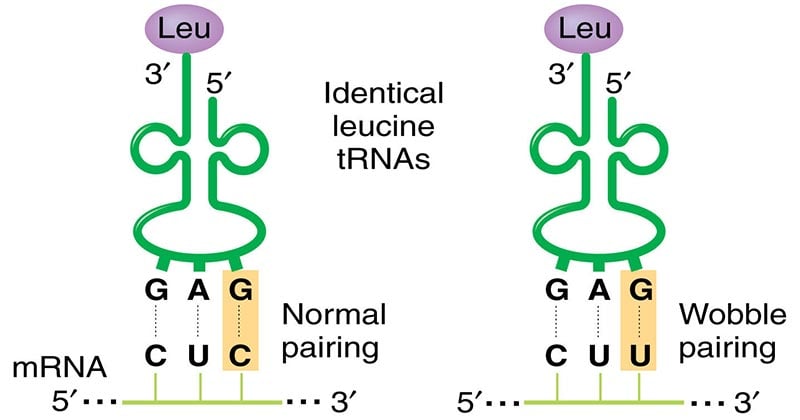
Crick’s hypothesis hence predicts that the initial two ribonucleotides of triplet codes are often more critical than the third member in attracting the correct tRNA.
Wobble base pairs
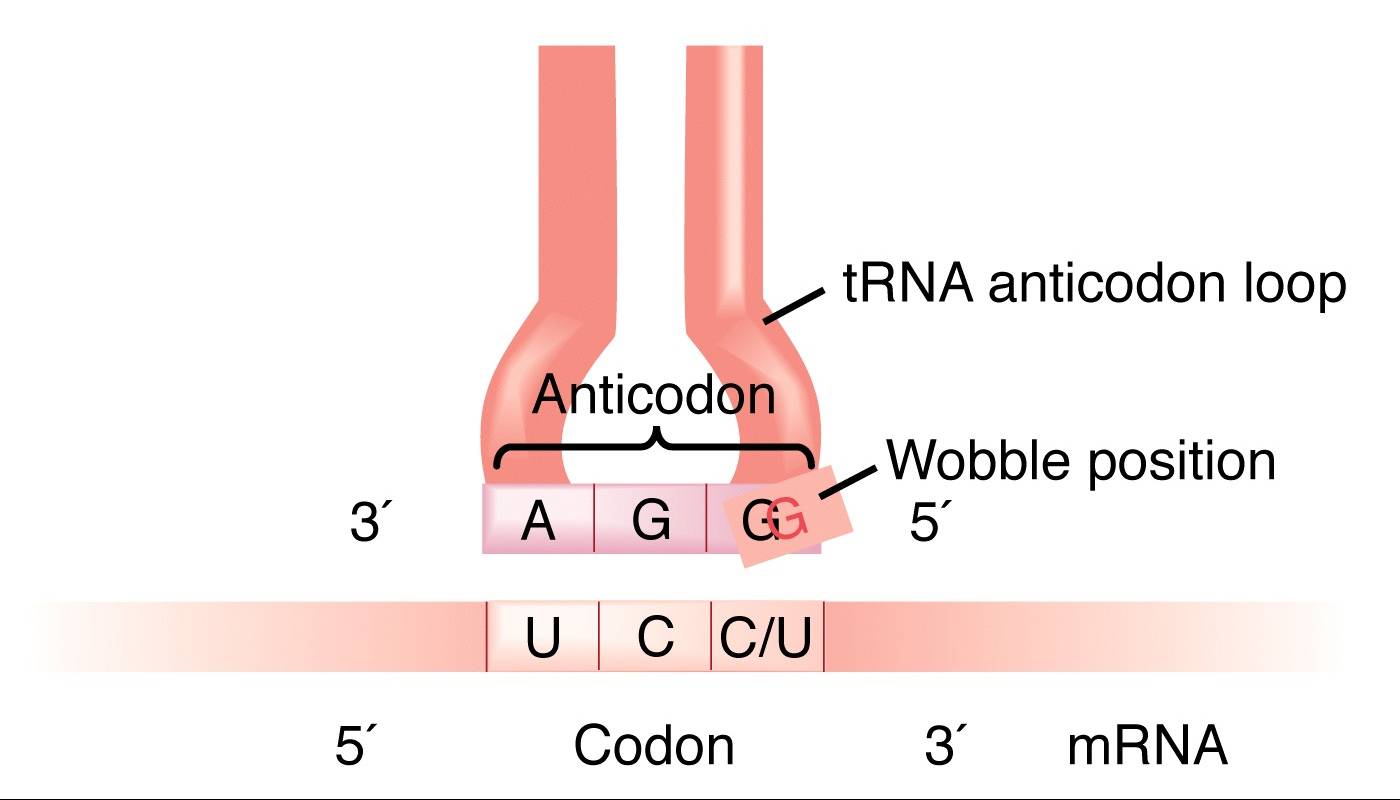
- A wobble base pair is a pairing between two nucleotides in RNA molecules that does not follow Watson-Crick base pair rules.
- The four main wobble base pairs are guanine-uracil (G-U), hypoxanthine-uracil (I-U), hypoxanthine-adenine (I-A), and hypoxanthine-cytosine (I-C).
- In order to maintain consistency of nucleic acid nomenclature, “I” is used for hypoxanthine because hypoxanthine is the nucleobase of inosine.
- Inosine displays the true qualities of wobble, in that if that is the first nucleotide in the anticodon then any of three bases in the original codon can be matched with the tRNA.
Significance of the Wobble Hypothesis
- Our bodies have a limited amount of tRNAs, and wobble allows for broad specificity.
- Wobble base pairs have been shown to facilitate many biological functions, most clearly proven in the bacterium Escherichia coli.
- The thermodynamic stability of a wobble base pair is comparable to that of a Watson-Crick base pair.
- Wobble base pairs are fundamental in RNA secondary structure and are critical for the proper translation of the genetic code.
- Wobbling allows faster dissociation of tRNA from mRNA and also protein synthesis.
- The existence of wobble minimizes the damage that can be caused by a misreading of the code; for example, if the Leu codon CUU were misread CUC or CUA or CUG during transcription of mRNA, the codon would still be translated as Leu during protein synthesis.
References
- Verma, P. S., & Agrawal, V. K. (2006). Cell Biology, Genetics, Molecular Biology, Evolution & Ecology (1 ed.). S .Chand and company Ltd.
- Klug, W. S., & Cummings, M. R. (2003). Concepts of genetics. Upper Saddle River, N.J: Prentice Hall.
- https://www.slideshare.net/sweetmerrymindfreak/genetic-code-53764823
- https://www.slideshare.net/ArchaDave/genetic-code-24735057
- https://www.scribd.com/document/344722502/Wobble-hyothesis-pdf
- https://en.wikipedia.org/wiki/Wobble_base_pair#Wobble_hypothesis

Am interested to explore more if cancerous cells may arise during translation and most specifically during this wobble translation
I need help to drow my method how i contact you my email is nado_53@hotmail.com
nice explaination