Nucleosomes are the basic repeating subunit of chromatin, composed of the nucleosome core, linker DNA, and linker histone proteins (Cutter & Hayes, 2015). Nucleosomes have a fundamental role in the structure and function of chromosomes, which are essential for storing and expressing genetic information.
It consists of DNA wrapped around histone proteins, forming repeating units that contribute to chromatin compaction within the nucleus. This structural arrangement allows for the selective unfolding of the specific regions of a gene by facilitating gene expression while maintaining the integrity of the genome (Kornberg & Klug, 1981).
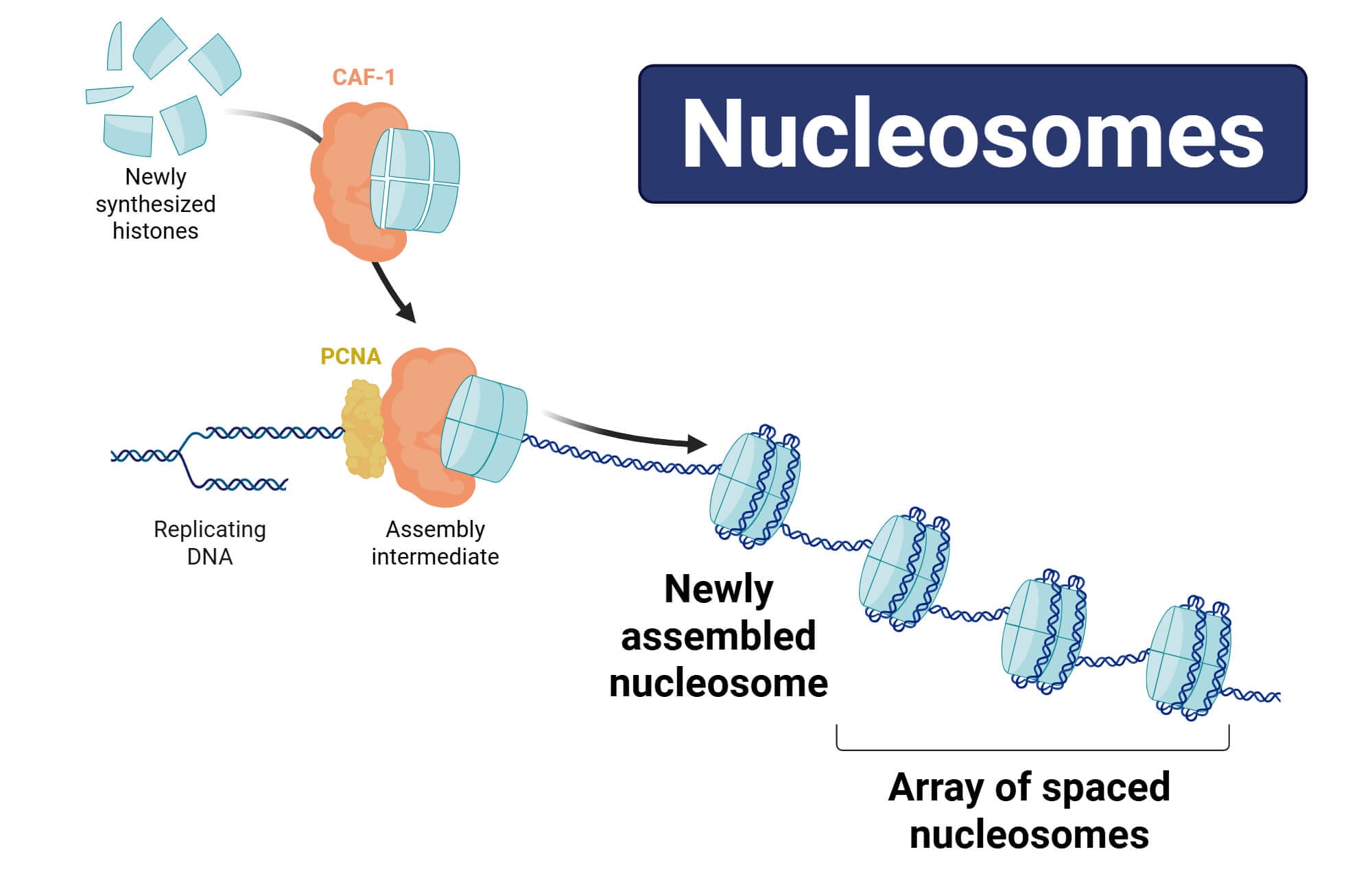
Research has shown that histone proteins play a crucial role in maintaining the structural stability of chromatin and regulating gene expression across different cell types. This diversity in the structure of chromosomes is linked with various histone combinations and modifications, thereby influencing DNA accessibility and utilization through cellular machinery. Here, nucleosomes provide valuable insights into the mechanism of gene regulation, chromatin dynamics, and overall complexity of cellular functions (Kornberg & Klug, 1981).
Components of Nucleosomes
The components of nucleosomes are,
- Nucleosome core
- Linker DNA
- Linker histone proteins (H2A, H2B, H3 and H4)
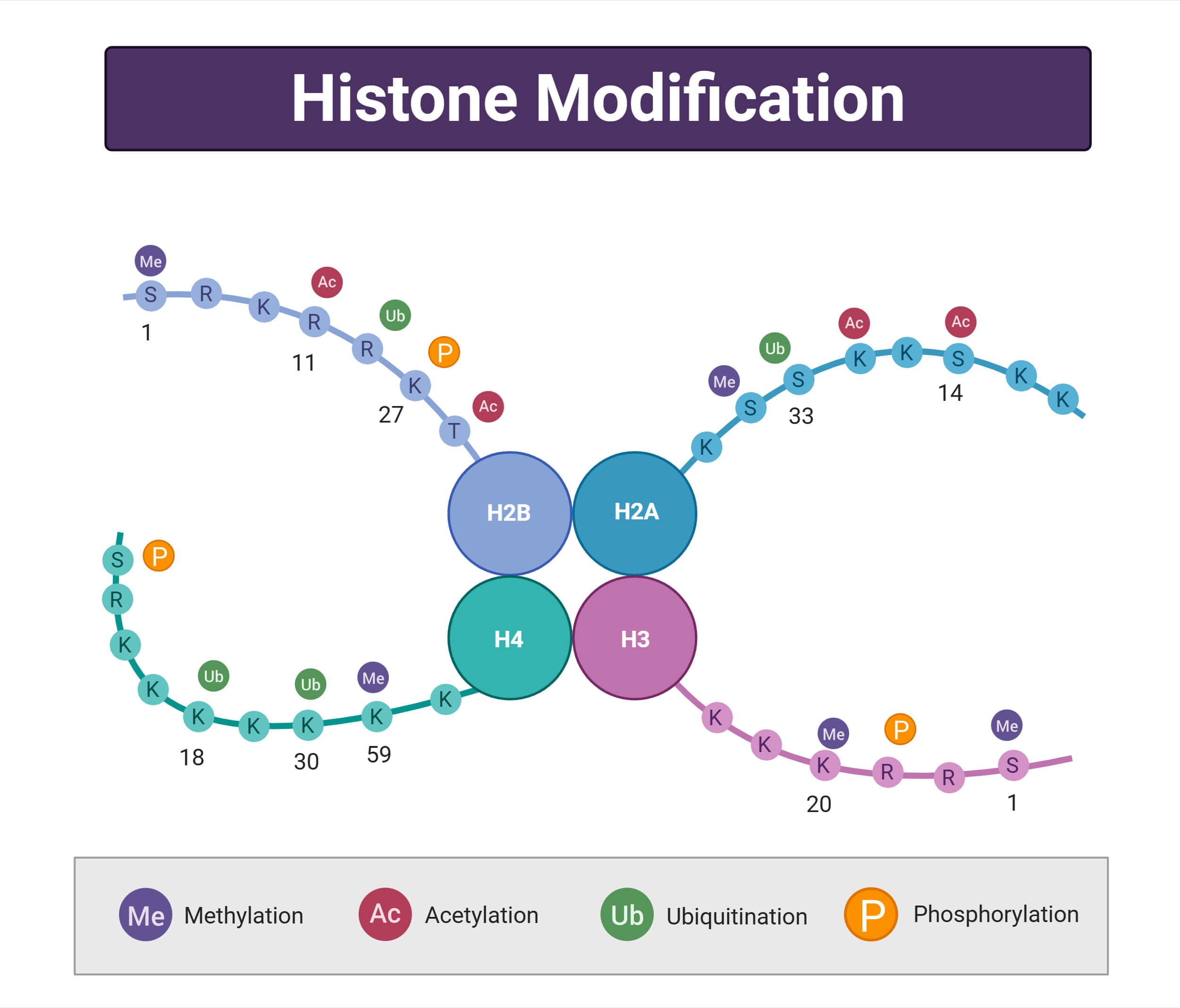
The core DNA in eukaryotes is associated with histone proteins. There are altogether 4 core histone proteins, with a molecular weight of approximately 11-15kDa. About 25% of the mass of core histone contains an unstructured N-terminal ‘tail’ domain in the absence of DNA or other macromolecules. These histone proteins form dimers that interact through 4 helix bundles in dimer-dimer interaction. This interaction has been described as a handshake interface that heterodimerizes H2A with H2B and H3 with H4. The H1 histone acts as a linker protein that is a primary component of nucleosomes and is included in all discussions related to the structure of nucleosomes (Cutter & Hayes, 2015).
Structure of a Nucleosome
Nucleosomes consist of 145-147 base pairs of DNA associated with octamers of histone proteins. These are made from the core histone proteins H2A, H2B, H3, and H4. These core histone proteins construct a spool-like structure with 1.75 left-handed superhelical turns around wrapped core DNA with a disc-shaped structure measuring 5.5nm in height and 11nm in diameter. The double helix of DNA with nucleosomes is affected by DNA sequences and other nuclear factors. It also consists of linker DNA that is connected to linker histone proteins. DNA in nucleosomes helps to protect nucleases and restricts the binding of trans-acting factors (Cutter & Hayes, 2015; McGinty & Tan, 2015).
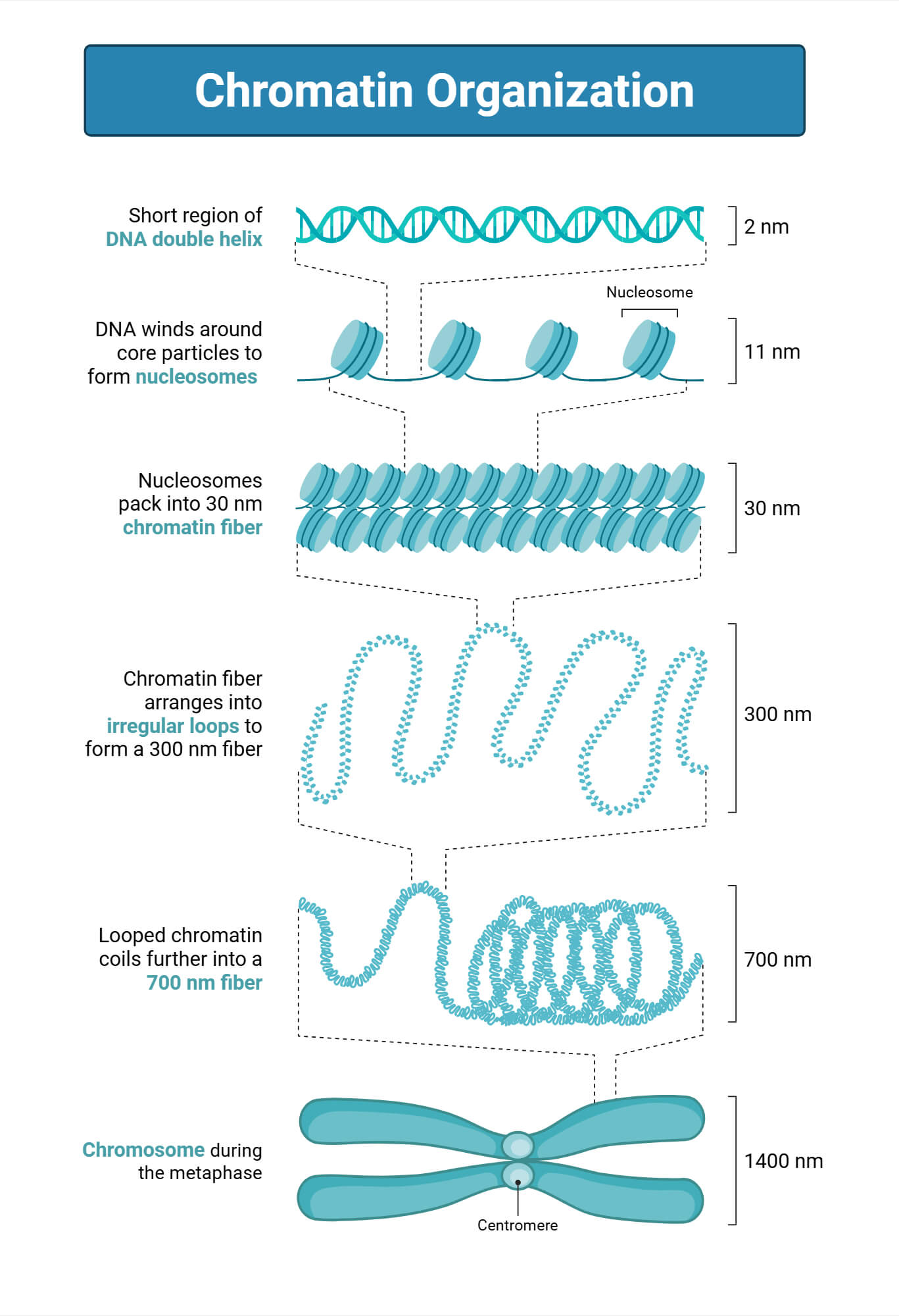
Functions of Nucleosome
The nucleosome has 3 primary functions based on its organization and regulation of genomic DNA, which are as follows:
- Genomic Concretion: It facilitates the first level of genomic compaction/concretion by organizing DNA segments of approximately 200 base pairs into a dense structure. This ensures extensive eukaryotic genome is confined within the cell nucleus.
- Signal hub for chromatin process: It acts as a signaling hub for chromatin and chromatin-associated enzymes with a diverse array of post-translational modifications (PTMs), regulating the recruitment of enzymes and influencing nucleosome stability.
- Assembling into high-order structure: It can self-assemble chromatin structures into higher-order formations. It enables the compaction of the genome during cell division.
These functions play a crucial role in regulating cellular processes and DNA repair (McGinty & Tan, 2015; Tan & Davey, 2011).
Nucleosome Positioning and Dynamics
The positioning and dynamics of nucleosomes are foundational to gene regulation, DNA accessibility, and gene expression. The dyad position of the nucleosome represents the center of DNA wrapped around histone proteins, influencing DNA transcription and protein regulation. These positions are analyzed through highly advanced sequencing techniques such as DNA sequencing, post-translational modifications (PTMs) of histone proteins, and chromatin remodeling. These factors help determine the stability of the nucleosome under various conditions (Teif & Clarkson, 2018).
Nucleosome dynamics play a significant role in DNA accessibility, as nucleosomes can reposition and change their structure via cellular signals. Therefore, nucleosomes serve as dynamic regulators in biological processes such as transcription, replication, and repair (Holdenrieder et al., 2008).
Nucleosomes in DNA Replication and Repair
Nucleosomes illustrate how structural constraints decrease with the alteration of nucleosome occupancy. This occurs in response to DNA damage. It has also been reported that the chromatin region gets damaged by UV light. The primary mechanisms for repairing double-strand breaks (DSBs) include non-homologous end joining (NHEJ) and homologous recombination (HR).
In NHEJ, free DNA ends are quickly ligated together, which can be error-free but may also lead to mutations if terminal bases are modified or removed. HR uses an intact template, typically from a sister chromatid, to accurately restore the original DNA sequencing, minimizing mutations. While mammalian cells predominantly utilize NHEJ, they also engage HR during DNA replication. Yeast cells similarly prefer HR during specific cell cycle phases but maintain a functional NHEJ pathway throughout the cycle.
Other repair pathways include base excision repair (BER) and nucleotide excision repair (NER), which address different types of DNA lesions, ensuring genomic stability and integrity (Hauer & Gasser, 2017).
Techniques for Studying Nucleosomes
Several techniques are commonly used in molecular biology and biochemistry to analyze the structure, composition, and dynamics. Here are some key methods for studying nucleosomes as per (Shi et al., 2023),
- Coarse-grained model: This model simplifies the atomistic dynamics of nucleosomes. It incorporates physical processes such as histone modifications, DNA sequence effects, and linker histone protein interactions.
- Free energy calculations: The 1-cylinder-per-nucleosome model reproduces a free energy profile related to chromatin interactions, including histone modifications and DNA interactions.
- Deep learning approach: This approach utilizes sequence features to represent nucleosome identification. It uses a deep learning algorithm to map a six-dimensional free energy surface based on inter-nucleosome distances.
- Integrated experimental-computational approach: This technique combines experimental data with computational simulations that predict nucleosome dynamics and the relationship between regulatory factors.
Nucleosomes and Gene Regulation
Nucleosomes play a crucial role in gene regulation by influencing the accessibility of DNA to transcriptional machinery and other regulatory proteins. The interplay between nucleosome positioning, histone modifications, and chromatin remodeling is essential for unraveling the complex mechanisms of gene regulation. Nucleosomes not only advance the knowledge of fundamental biology but also play a significant role in diseases linked to dysregulation of gene expression (Jiang & Pugh, 2009).
Clinical Significance of Nucleosomes
Nucleosomes have significant clinical relevance, particularly in oncology, where they serve as biomarkers in various diseases. They are even released into circulation during cell death. Elevated levels of circulating nucleosomes have been observed in patients with different types and stages of cancer. Although they are not specific for diagnosing cancer due to their elevation in benign conditions, they can provide valuable insights for therapeutic monitoring. A decrease in circulating nucleosomes indicates a positive response to chemotherapy; however, it may suggest disease progression. This makes nucleosomes a promising tool for assessing treatment efficacy and potentially guiding early therapeutic decisions in cancer management (Holdenrieder et al., 2008).
Recent Advances and Future Perspectives of Nucleosomes
Although recent advances have improved the understanding of nucleosome structure and the factors influencing DNA positioning, additional research is necessary to uncover the underlying mechanisms. The insights gained from examining various DNA sequencing methods, metal interactions, and medicinal agents enhance understanding of molecular recognition within nucleosomes and lay the groundwork for future research in nucleosome targeting and drug development (Tan & Davey, 2011).
Over the past 5 years, significant advancements have been made in nucleosome structure and function, particularly in the sequence-dependent nature of nucleosomal DNA. The recent advancement in co-crystallization and NMR techniques has identified critical interactions in the acidic patch arginine-anchor, which facilitate nucleosome recognition. Additionally, improvements in cryo-electron microscope technology have enabled detailed visualization of the 30 nm chromatin fiber and its interaction with chromatin enzymes. Despite these developments, there is less research on the complex roles of nucleosomes in chromatin dynamics.
Therefore, future research is essential to uncover new paradigms in nucleosome recognition to contribute to the functional organization of the eukaryotic genome (McGinty & Tan, 2015).
Conclusion
Nucleosomes play a crucial role in the structure, function, and gene regulation. They contribute to DNA compaction, gene expression, and chromatin dynamics. Although significant progress has been made in understanding nucleosome structure, function, and their involvement in DNA repair and replication, much research remains to fully elucidate the mechanisms behind nucleosome dynamics and their complex roles in chromatin remodeling.
Advances in experimental and computational techniques, including deep learning approaches, offer valuable tools for studying nucleosome behavior and its impact on gene regulation. The clinical significance of nucleosomes as biomarkers, particularly in oncology, opens new avenues for therapeutic monitoring and early disease detection.
Continued research is essential to uncover new paradigms in nucleosome recognition and their role in the functional organization of eukaryotes. Therefore, nucleosomes could become pivotal in personalized medicine, providing new targets for drug development and enhancing the precision of therapeutic strategies.
References
- Cutter, A. R., & Hayes, J. J. (2015). A brief review of nucleosome structure. FEBS Letters, 589(20), pp. 2914–2922. https://doi.org/10.1016/j.febslet.2015.05.016
- Hauer, M. H., & Gasser, S. M. (2017). Chromatin and nucleosome dynamics in DNA damage and repair. Genes & development, 31(22), 2204-2221. https://doi.org/10.1101/gad.307702
- Holdenrieder, S., Nagel, D., Schalhorn, A., Heinemann, V., Wilkowski, R., Von Pawel, J., Raith, H., Feldmann, K., Kremer, A. E., Müller, S., Geiger, S., Hamann, G. F., Seidel, D., & Stieber, P. (2008). Clinical relevance of circulating nucleosomes in cancer. Annals of the New York Academy of Sciences, 1137, 180–189. https://doi.org/10.1196/annals.1448.012
- Jiang, C., & Pugh, B. F. (2009). Nucleosome positioning and gene regulation: Advances through genomics. Nature Reviews Genetics, 10(3), pp. 161–172. https://doi.org/10.1038/nrg2522
- Kornberg, R. D., & Klug, A. (1981). The Nucleosome. Scientific American, 244(2), 52–65. https://doi.org/10.2307/24964283
- McGinty, R. K., & Tan, S. (2015). Nucleosome structure and function. In Chemical Reviews, 115(6), pp. 2255–2273. https://doi.org/10.1021/cr500373h
- Shi, D., Huang, Y., & Bai, C. (2023). Studies of the Mechanism of Nucleosome Dynamics: A Review on Multifactorial Regulation from Computational and Experimental Cases. Polymers, 15(7). https://doi.org/10.3390/polym15071763
- Tan, S., & Davey, C. A. (2011a). Nucleosome structural studies. Current Opinion in Structural Biology, 21(1), pp. 128–136. https://doi.org/10.1016/j.sbi.2010.11.006
- Tan, S., & Davey, C. A. (2011b). Nucleosome structural studies. Current Opinion in Structural Biology, 21(1), pp. 128–136. https://doi.org/10.1016/j.sbi.2010.11.006
- Teif, V. B., & Clarkson, C. T. (2018). Nucleosome positioning. Encyclopedia of Bioinformatics and Computational Biology: ABC of Bioinformatics, 1(3), pp. 308–31). https://doi.org/10.1016/B978-0-12-809633-8.20242-2
