Amplified Fragment Length Polymorphism PCR (AFLP-PCR) is a DNA fingerprinting technique that selectively amplifies restriction fragments generated from digested genomic DNA to detect genetic polymorphisms and create unique genomic fingerprints.
Amplified Fragment Length Polymorphism (AFLP) PCR is an amplification method. It uses selective amplification of restriction fragments, which are a subset of digested DNA, to create and compare unique genome fingerprints. These fragments come from a total double digest of genomic DNA.
AFLP-PCR can be used for DNA fingerprinting of any sample, regardless of complexity or origin. This method does not require prior information about the gene of interest, making it superior to other DNA fingerprinting methods. Its high reproducibility and sensitivity enable the detection of small sequence variations.
Additionally, it requires only small sample amounts, as little as 0.05 μg. AFLP can be used with DNA from humans, animals, plants, and microbes, making it a potential universal DNA fingerprinting system. Applications include monitoring trait inheritance in breeding, diagnosing genetic diseases, pedigree analysis, forensic typing, parentage analysis, screening for genetic markers, and microbial typing.
Objectives of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
The objectives of this method include:
- To amplify sequences of any origin and complexity.
- To provide high reproducibility of results
- To be able to detect even small genome sequence variation without prior sequence knowledge.
- To evaluate genetic variation within a species or among closely related species.
Requirements of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
The materials required for this method include:
- Primers: Oligonucleotide primers, typically 20–40 nucleotides long, are short single-stranded DNA fragments used to initiate amplification. Here, two primers are used.
- DNA template
- Restriction endonuclease
- Adaptors
- Ligase enzyme
- DNA ligase buffer
- Reaction buffer
- Mg2+
- DNA polymerase
- Thermocycler
Principle of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
AFLP -PCR is an amplification method where selective amplification of DNA fragments obtained from treatment with restriction endonucleases is performed to study the genetic polymorphisms in the gene of interest. Genetic polymorphisms can be detected by the presence or absence of fragments after restriction digestion and amplification of genomic DNA. This method consists of 4 steps: DNA digestion, ligation, amplification, and gel analysis.
The genomic DNA of interest is first digested using two restriction enzymes.
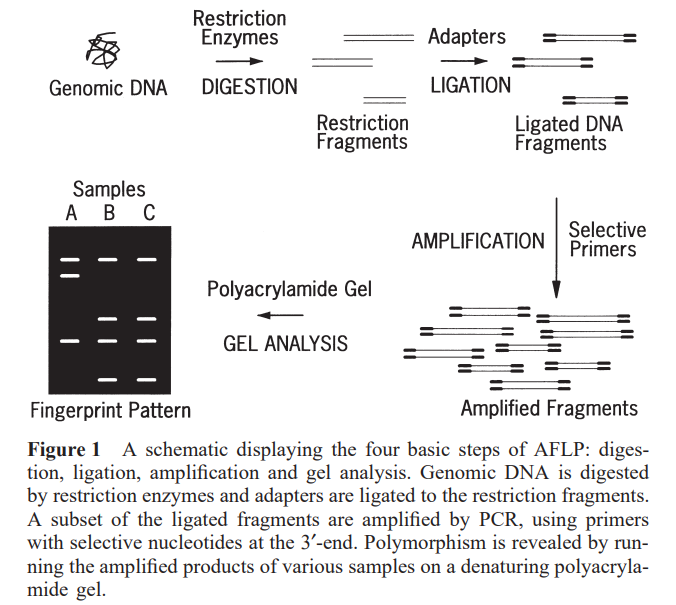
Most commonly, a rare cutter like EcoRI (6-bp restriction site) and a frequent cutter like MseI (4-bp restriction site) are widely used. The fragments obtained are then ligated with adaptors. These adapters contain two sequences: a longer sequence and a restriction-specific sequence homologous to either the 5’ or 3’ end of the restricted Genomic DNA. The ligation is done with a ligase such as the T4 ligase. Sequence-specific primers complementary to both the adaptor and the restriction site, with additional selective nucleotides at their 3’-ends, are then used to amplify the ligated DNA fragments.
The selective primers ensure that only fragments with complementary nucleotides extending beyond the restriction site are amplified under stringent annealing conditions, thereby reducing the complexity of the mixture. The polymorphisms can then be examined using polyacrylamide gel electrophoresis by comparing the patterns generated for each sample to determine the presence or absence of a particular DNA fragment.
Polyorphisms occur in 3 cases:
- Mutation in the restriction site
- Mutation in regions complementary to primer extensions
- Mutation in areas adjacent to the restriction site, or a deletion/insertion within the amplified region
Steps of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
The entire process of AFLP-PCR consists of four steps:
- DNA digestion
- Ligation
- Amplification, and
- Gel analysis
DNA Digestion
Here, the genomic DNA is digested into fragments using restriction enzymes. Two different enzymes are used for this purpose: a rare-cutting enzyme with 6- to 8-base recognition and a frequent-cutting enzyme with 4-base recognition. Such a high degree of specificity allows for the production of reproducible DNA fragments. The type of enzymes to be used depends on the genomic DNA and its methylation status. The use of two different enzymes is because:
- The number of fragments generated by amplification can be manipulated to produce fingerprint patterns of desired complexity.
- A large number of distinct fingerprints can be generated from the fragments using only a few primers.
Some examples of rare-cutting enzymes include EcoRI, AseI, HindIII, ApaI, and PstI. MseI and TaqI are commonly used as frequent cutters. Since the DNA of Eukaryotes is AT-rich, MseI, with a recognition sequence of TTAA, is the most preferred enzyme. The frequent cutters help generate small fragments of the desired size, within the range of 100–1000 base pairs. Three different types of fragments are produced, these include:
- Fragments cut with a rare-cutting enzyme on both ends.
- Fragments with a frequent-cutting enzyme on both ends.
- Fragments with both enzymes acting on each end.
Ligation
Since this method does not require any prior sequence knowledge, double-stranded nucleotide adaptors, approximately 10-30 base pairs long, are required. These adaptors are complementary to the sticky ends of the corresponding restriction sites and are ligated to these sites using T4 DNA ligase. The adjacent restriction half-site and the adaptor sequence serve as primer-binding sites for PCR amplification. To prevent any chance of restoration of the original restriction site, a change is induced in the recognition site of the adaptor sequence. This allows the restriction and ligation processes to be performed together in a single tube.
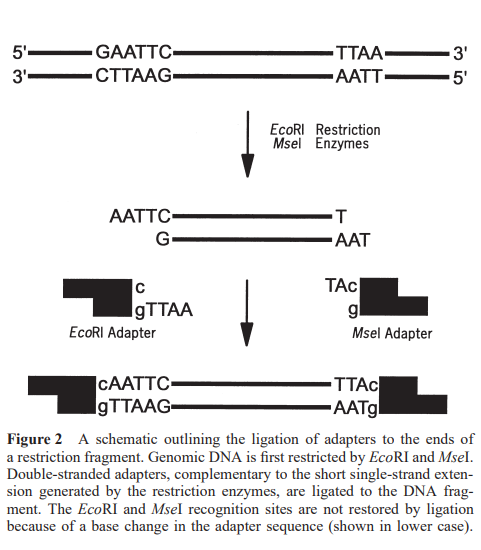
Additionally, adaptor-to-adaptor ligation is prevented with the use of non-phosphorylated adaptors. The complexity of the DNA mixture can be reduced by using biotin-labeled adaptors complementary to the rare-cutting restriction half-site, e.g.,g biotin-labeled EcoRI adapters) before amplification.
Use of such adaptors helps in the separation of fragments of at least one rare-cutting restriction site from the fragments having two frequent-cutting restriction sites (e.g., EcoRI-EcoRI and EcoRI-MseI fragments are separated from MseI-MseI fragments). The fragments obtained from these enzymes are selected using streptavidin-coated magnetic beads.
Selective amplification
For the selective amplification of the fragments, two specific primers are designed:
- One complementary for the adaptor and to the adjacent rare-cutter restriction site sequence with one to three additional selective nucleotides at the 3′-end.
- Whereas the second primer is complementary to the adaptor and frequent-cutter recognition site sequence with an additional selective one- to three-base extension.
Only a restricted number of fragments with complementary nucleotides that extend beyond the restriction site will get amplified under stringent annealing conditions. The nucleotide extension present on the 3’-end has two purposes:
- To allow a variety of test restriction fragments to be amplified.
- To increase the chance of detecting polymorphisms that may be present beyond the restriction site.
The primers complementary to the rare-cutting restriction site and the adaptor have a higher annealing temperature than the frequent-cutter primers.
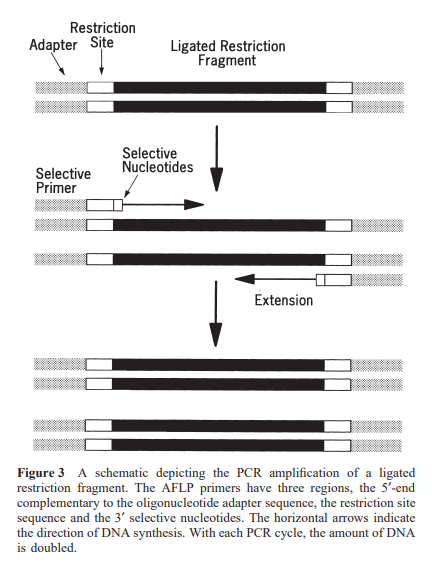
For a complex genome (108 -109 bp), a two-step amplification method requiring three selective nucleotides on one or both primers is needed. A pre-amplification step (first amplification), utilizing primers with single or no selective nucleotide, reduces the overall complexity of the mixture. The products from this step are then diluted and used as templates for the second amplification.
Gel analysis
The PCR products, before electrophoresis, are subjected to denaturation by heating at 90-95 °C for 3–5 min. Of the two strands, only one strand is labeled for analysis, typically the 5′-end with (γ32P or – 33P) ATP using T4 polynucleotide kinase. This is to prevent the occurrence of any double bands due to unequal mobility of the two strands.
Analysis of amplified fragments can be performed through two methods:
- Labeling with radioisotopes or a fluorescent dye: The strands labeled this way can be examined using autoradiography. The resulting bands may be visualized by eye or by high-resolution densitometric scanning for computer-assisted analysis, e.g., Gel Compar 3.1 software (Applied Maths, Kortrijk, Belgium).
- Silver staining: In this method, labeled primers are not required; both strands are visualized. This method has similar sensitivity and resolution to the labeling methods, with better resolution than labeling with 32P, especially in regions containing more than 300 bp. Recovering products from dehydrated gels via rehydration and direct transfer to a PCR tube for amplification is an added advantage of this method.
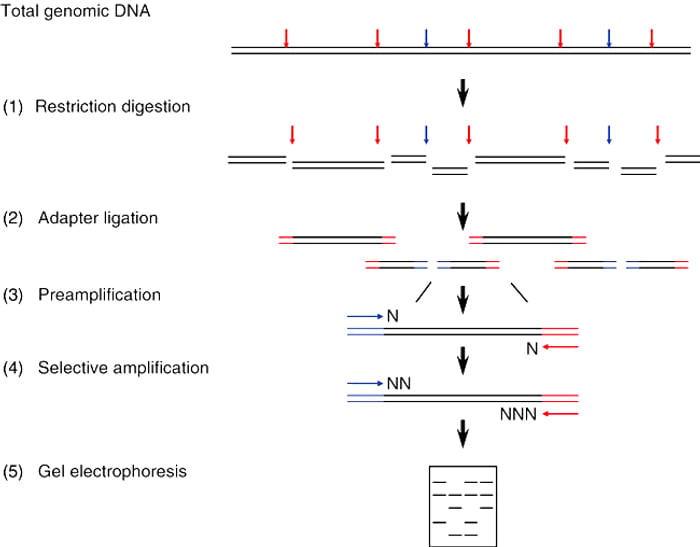
Types of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
Some of the modifications of this method include:
- Selectively amplified microsatellite polymorphic loci (SAMPL): It is a microsatellite-based modification of AFLP and can detect higher levels of genetic variation within genotypes. This method can detect high levels of polymorphisms due to its association with the hypervariable microsatellite region.
- Microsatellite-amplified fragment length polymorphism (M-AFLP): It allows for the detection of intravarietal genetic differences. It is a highly efficient method for generating a large number of polymorphic bands.
- Sequence-specific amplified polymorphism (SSAP): It is the first retrotransposon-based barcoding method based on AFLP. The primers are designed for the LTR region and can also be used for the RT internal sequence.
- Amplification of insertion mutagenized sites (AIMS): It is based on the reduction of band complexity through specific PCR amplification of insertion mutagenized sites.
- Methylation-sensitive amplified polymorphism (MSAP): Here, cleavage of the genomic DNA using methylation-sensitive restriction enzymes such as HpaII, MspI, etc., and followed by the usual steps of AFLP is performed.
- Resistance gene analog-anchored amplified fragment length polymorphism (AFLP-RGA): Degenerate RGA primers in combination with selective AFLP primers are used in the second round of amplification.
- Three-endonuclease AFLP (TE-AFLP): This method reduced the number of amplified fragments through both primer extension and selective ligation. Three endonucleases and two primers are used, which reduces competition during PCR, eliminating the need for two-step amplification.
- Secondary digest AFLP (SDAFLP): It is a variation of the MSAP technique, utilizing a restriction enzyme site-specific single primer to amplify the digested template DNA, followed by digestion with a methylation-sensitive enzyme.
- Miniature Inverted-repeat Transposable Elements (MITEs): These elements are found in maize and can be used as molecular markers.
- RNA Fingerprinting Using cDNA-AFLP: Complementary to northern blot analysis for studying gene expression, this method uses the RNA fingerprinting technique with the AFLP protocol on a cDNA template.
- Nonradioactive differential display (DD) AFLP: This method is used for monitoring differentially expressed genes.
Examples (Instruments) of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
AFLPTM Plant Mapping Kit (PE Applied Biosystems)
- Two modules are available based on genome size: Small Plant Genome Kit for 50–500 megabases and Regular Plant Genome Kit for 500–5000 megabases.
- 16 selective primers are available, which include eight MseI primers and eight EcoRI primers.
Pro TechEx-DNA Finger Printing Teaching Kit (Using AFLP Technique) (PROGENE)
- 5 reactions are available per kit
- Used to detect Single Nucleotide Polymorphisms
Applications of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
Some of the applications of this method, other than DNA and RNA fingerprinting, include:
- Genetic Diversity Studies Using AFLP Markers: Diversity studies based on AFLP markers are popular due to the elimination of the requirement for prior sequence information, as well as their multi-locus and genome-wide nature.
- QTL Mapping: AFLP markers are used to construct linkage maps for QTL analysis of agronomic traits, including disease resistance and salt tolerance.
- Characterization of mammalian genotypes: The AFLP technique can be used to study genotypes within the same strain and generate a dendrogram to assess relationships. It was used to study rat inbred strains.
- Epidemiological typing of bacteria: AFLP analysis, in addition to ribotyping, biotyping, cell envelope protein electrophoretic typing, and antibiogram typing, can be used to compare and study bacterial strains.
Advantages of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
The advantages of this method are:
- Since detection of AFLP fragments does not depend on hybridization, partial digestion, or faint patterns, and due to the stringent annealing temperature, the AFLP analysis method is reproducible and robust.
- A large amount os molecular markers can be generated without any prior knowledge of the genome.
- A small amount of DNA can be used as a starting material.
- Can be used for samples with any origin and varying complexity.
- The markers generated are independent in nature since 90% of these detect point mutations in mutation sites.
Limitations of Amplified Fragment Length Polymorphism PCR (AFLP-PCR)
- Since several steps, such as the pre-amplificationstep is involved in this method, it is a time-consuming and cumbersome process.
- Single analysis using radioisotopes and silver staining is required; it is an expensive process.
- Additionally, the adaptors and restriction enzymes required for this method are also expensive.
- Distinguishing between heterozygous and homozygous individuals is not possible, thereby limiting its use in population genetics analysis, genetic mapping, and marker-assisted selection.
Conclusion
PCR has beenwidely usedfort eh amplificationofgenef interest for various purposes. AFLP-PCR is a type of PCR method that is utilized for the detection of polymorphism within genomic samples using restriction endonucleases. It has found wide application in studying the genetic diversity of plants, bacteria, eukaryotes, fungi, and other organisms. This is due to the use of molecular markers, which are more accurate compared to phenotypic markers.
It is a useful method when prior information about a genomic sample is not available, and low levels of the sample are available. The many advantages of the technique, primarily its reproducibility and number of loci detected per reaction, will enhance its overall application.
References
- Blears, M. J., De Grandis, S. A., Lee, H., & Trevors, J. T. (1998). Amplified fragment length polymorphism (AFLP): a review of the procedure and its applications. Journal of Industrial microbiology and Biotechnology, 21(3), 99-114.
- Sheeja, T. E., Kumar, I. P. V., Giridhari, A., Minoo, D., Rajesh, M. K., & Babu, K. N. (2020). Amplified fragment length polymorphism: applications and recent developments. Molecular Plant Taxonomy: Methods and Protocols, 187-218.
- Savelkoul, P. H. M., Aarts, H. J. M., De Haas, J., Dijkshoorn, L., Duim, B., Otsen, M., … & Lenstra, J. A. (1999). Amplified-fragment length polymorphism analysis: the state of an art. Journal of clinical microbiology, 37(10), 3083-3091.
- Paun, O., & Schönswetter, P. (2012). Amplified fragment length polymorphism: an invaluable fingerprinting technique for genomic, transcriptomic, and epigenetic studies. In Plant DNA Fingerprinting and Barcoding: Methods and Protocols (pp. 75-87). Totowa, NJ: Humana Press.
- https://archive.nptel.ac.in/content/storage2/courses/102103013/module6/lec5/6.html
- https://www.indiamart.com/proddetail/pro-techex-dna-finger-printing-teaching-kit-using-aflp-technique-9551658597.html
