PCR involves three main steps: Denaturation, Annealing, and Extension. To make the method more efficient and specific, variations of the method have been developed, one of which is allele-specific PCR, used to amplify single-nucleotide polymorphisms.
Allele-Specific PCR (ASPCR) is a PCR method used to study mutations and polymorphisms by targeting specific alleles. This non-radioactive method can distinguish between alleles differing by one or more nucleotides based on whether amplification occurs during PCR.
In ASPCR, two sets of primers are used: one primer is specific to the mutant allele and will not efficiently bind or extend on the normal allele, while the other primer is specific to the normal allele and is refractory to the mutant sequence.
After amplification, the resulting PCR products are analyzed using agarose gel electrophoresis to identify mutations in the gene of interest. This method is widely used to diagnose genetic and infectious diseases, such as sickle cell anemia.
Objectives of Allele-Specific PCR (ASPCR)
The objectives of this method are:
- To determine the presence of a mutant allele.
- To create a mutant-specific primer
- To amplify the specific mutant allele
- To prevent any non-specific extension of nucleotides.
- For the detection of single-nucleotide polymorphism.
Requirements of Allele-Specific PCR (ASPCR)
The component required for ASPCR is similar to that of a traditional PCR; however, allele-specific primers for detecting the mutant allele are designed. The requirements are as follows:
- Primers: Oligonucleotide primers, typically 20–40 nucleotides long, are short single-stranded DNA fragments used to initiate amplification. In ASPCR, primers are specifically designed so that their 3’ end matches either the normal or the mutant allele exactly. A single-nucleotide difference at the 3’ end ensures that the primer will bind only to and amplify its specific target allele. Mismatched primers are intentionally designed to match only one allele and prevent amplification of the non-target allele.
- dNTP’s
- Reaction buffer
- Mg2+
- DNA polymerase
- Thermocycler
Principle of Allele Specific PCR (ASPCR)
The basis of the PCR method depends on the primer’s specific design. An allele has two forms: the normal (wild-type) form and the mutant form. When utilizing a primer specific for the normal allele, it cannot extend the mutant form, or the extension will be slow. The 3’ OH end of the primer binds with the sequence-specific area of the DNA template. This can be leveraged to create a primer specific to the gene to be amplified. By modifying the 3’ end of the primer, it can either form a match pair and amplify the gene or form a mismatch, thus preventing amplification of the gene. This primer can be designed specifically for the mutant to be amplified, thereby forming a matched pair only with the mutant, not with the wild-type allele.
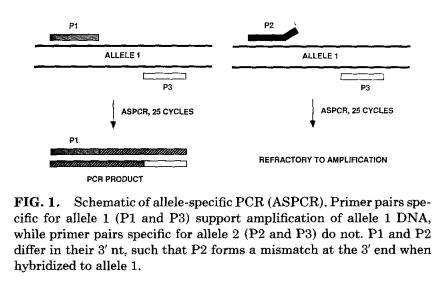
The 3’ end of the nucleotide of the primer is complementary to the gene of interest or the template DNA. This can form the required base pairing, which may be a mismatch or a matched pair depending on the site-specific sequence it binds to. The matched pairing occurs with the wild-type allele, whereas the mismatch occurs with the mutant allele. The foundation of ASPCR is built on the concept of mismatch. Hence, a mismatch pairing occurs, and no extension or only slow extension occurs, which can be analyzed to determine the presence of a mutant in the template DNA. Here are two primers: one for the wild-type allele (allele 1, P1) and one for the mutant allele (allele 2, P2). When P1 base pairs with allele 1, extension occurs, but with the mutant allele, P1 will be refractory to extension.
Similarly with P2, the same phenomenon occurs. Additionally, a third primer, referred to as the distal primer, is designed to be complementary to a sequence common to both alleles. The position of P3 relative to P1 and P2 yields a PCR product size that is easily identifiable by agarose gel electrophoresis. The annealing temperature varies with the type of mismatch induced.

Steps / Protocol of Allele Specific PCR (ASPCR)
The entire process of ASPCR includes:
- Designing of primer
- Amplification
- Agarose gel electrophoresis
- Interpretation of the result
Designing of primer
When designing the primer, along with the normal criteria such as
- Have less GC Content.
- Less hairpin structure.
- 20-30 nucleotides in length.
The primer design should also include a certain mismatch at the 3’-OH end. The mismatch produced should be strong to prevent any chances of false amplification at the 3’-OH end. Some of the mismatched pairs are mentioned below:
- Strong mismatch: G/A, C/T, T/T
- Medium mismatch: A/A, G/G, C/C,
- Weak mismatch: C/A, G/T
For example, for a wild-type allele containing G at the 5’ end, the P1 primer should contain C at the 3’ end, whereas the P2 primer should have T in place of C. Weak mismatches increase the chance of strong base pairing and thus increase the chances of amplification resulting in improper results.
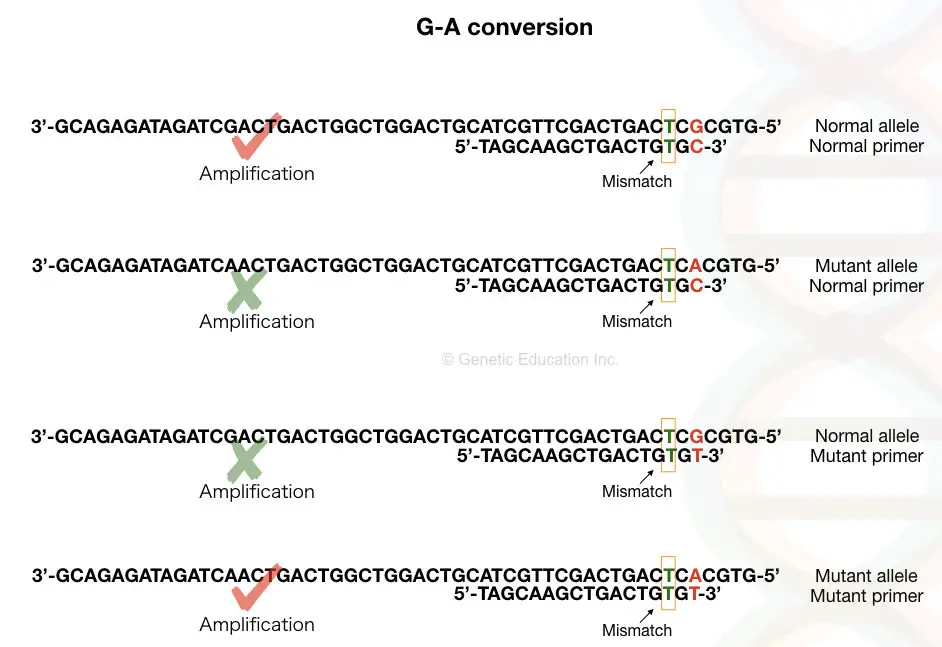
Amplification
This follows the same pattern as the traditional PCR method. It has three steps:
- Denaturation: The thermocycler is set to 95 °C for 30 seconds to denature the template DNA. This is the standard temperature and time for most templates, but for certain GC-rich templates, a longer denaturation of 2–4 minutes at 95°C is required to produce amplicons.
- Annealing: The annealing temperature varies for the mismatch primer utilized. A higher temperature is preferred. This prevents binding, which may result in nonspecific amplification, as seen when using a lower annealing temperature. The temperature is lowered, and the specifically designed primer anneals to the template DNA.
- Extension: Here, the temperature is increased to 65-75°C, and DNA polymerase begins adding the deoxynucleotides. This step typically lasts 15-60 seconds, extending to minutes/kb. Extension of sequence-specific primers with nucleotides complementary to the DNA template occurs in this step.
The number of cycles is set to a lower number, ideally to 22-25 cycles, to maintain the specificity of the reaction. An excessive number of ASPCR cycles can result in the deterioration of the specificity of the reaction in two ways:
- An increase in the production of fragments formed by incorrect priming with a 3′ mismatched nucleotide may arise
- Any false fragments arising from nonspecific priming may get synthesized along with the expected fragment.
With increased cycles, the likelihood of nonspecific binding and primer-dimer formation increases, leading to redundant results.
The reaction mixture used involves the following components in certain compositions.
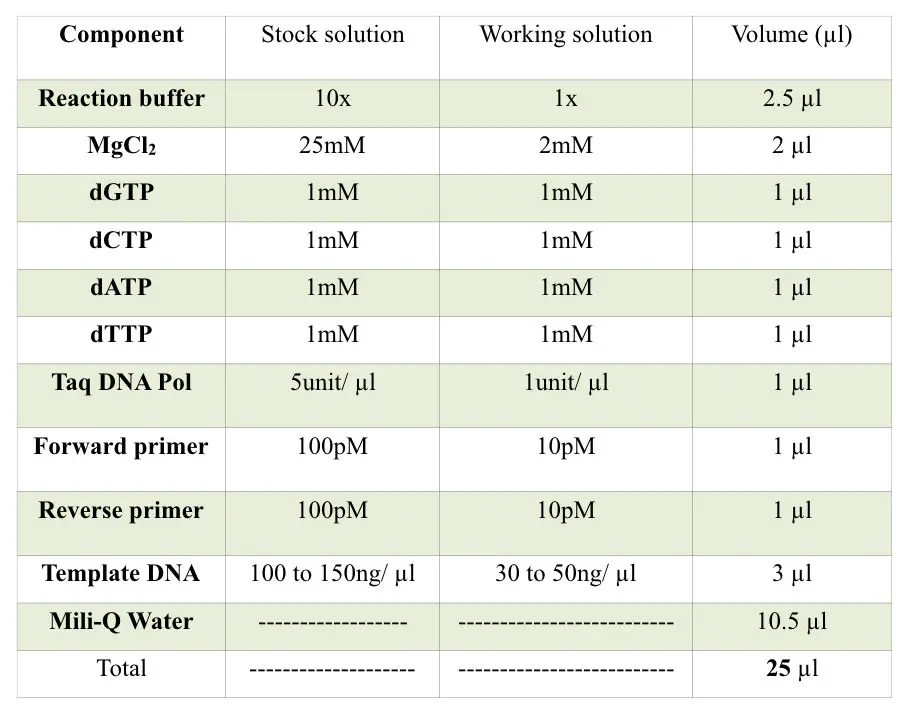
In addition to the reaction mixture, internal controls, negative controls, and positive controls should be included to monitor the results.
Agarose Gel Electrophoresis
1.5% agarose is utilized for the electrophoresis reaction. Into the wells, the product along with the loading buffer and the load gel is added. One lane is left for a suitable-sized DNA marker. After running, the gel was stained with Ethidium bromide and viewed under a UV transilluminator to interpret the results.
Interpretation of Result
In every electrophoretic reaction, an internal control must be present to ensure that the process is error-free. For interpretation, assuming there are three different samples: homozygous normal, homozygous dominant/mutant, and heterozygous carrier. The normal and the mutant homozygous samples each show a single band, whereas the heterozygous carrier shows both the normal and mutant allele bands; two bands are seen. This result is observed because the normal primer cannot extend the mutant allele, and vice versa. Thus, single bands are observed for both samples and the heterozygous carrier, as extension with both primers is possible due to the presence of both alleles.
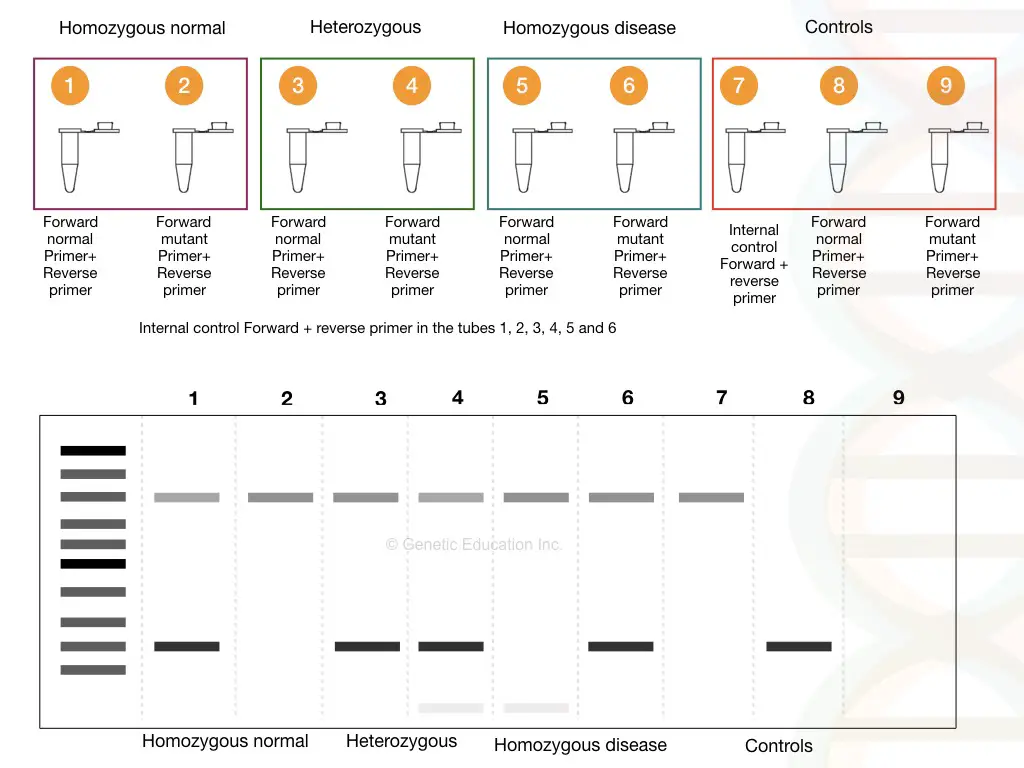
Types of Allele-Specific PCR (ASPCR)
Some variations of Allele-specific PCR have been developed by modifying the existing method. This includes:
Tetra Primer Allele-Specific PCR (T-ARMS-PCR)
Four different primers are used in this method to detect various mutations, hence the name. The primer volume used provides valuable diagnostic information. It uses forward and reverse primers for the wild-type allele; similarly, the same method is used for the mutant allele.
Multiplex-ARMS PCR
Two separate multiplex reactions are used to detect more than two SNPs. It is a complex process and carries an increased risk of false-positive results. However, it is a faster and cheaper process compared to the tetra ARMS PCR.
Quantitative ARMS-PCR (qARMS-PCR)
This method is a mix of qPCR and Allele-specific PCR, in which the amplified alleles can be quantified. Thus, the exact amount of the normal type or mutant allele present in the sample can be quantified.
Examples of Allele-Specific PCR (ASPCR)
Type-it Fast SNP Probe PCR Kit (800) (QIAGEN)
- The kit is based on the specific HotStarTaq Plus DNA Polymerase and a new buffer system. Both systems help in reliable and accurate allelic discrimination.
- The mix included in the kit enables highly specific and accurate probe binding.
- It can even be used for GC-rich loci, which are often hard to detect, or in cases of low starting template amounts.
- Well separation of allelic clusters is seen with this kit.
- For SNP genotyping, strong fluorescent signals are provided by the TaqMan Probe.
PCR Allele Competitive Extension Genotyping (Integrated DNA Technologies)
- It is an allele-specific PCR kit focused on fluorescent detection and competitive allele-specific PCR genotyping technology.
- It is a good fit for biallelic discrimination of single-nucleotide polymorphisms and indels (Insertions and Deletions) at specific loci.
- Beneficial forresearchers that requires toperform high-volume experiments with large sample sets, e.g., plant breeding projects.
- If the sample genotype is homozygous, one or two fluorescent signals will be generated, whereas both the sinalswill be generated in case of a heterozygous genotype.
Devyser CFTR Core kit (Dvsyr)
- Used for genotyping normal and mutant alleles at 33 loci of the CFTR gene using purified human genomic DNA.
- It can detect 36 of the most common mutations in populations of European origin through allele-specific detection of wild-type and variant alleles.
- The assay can also be used to detect polythymidine variants within intron 9 (IVS8) of the CFTR gene.
- This kit is based on multiplex allele-specific PCR amplification to detect both the wild-type and mutant alleles in the CFTR gene.
- It generates fluorescently labeled fragments, which are analyzed by capillary electrophoresis on a Genetic Analyzer instrument. These fragments are then identified based on size and fluorescent labels.
Applications of Allele-Specific PCR (ASPCR)
ASPCR finds its applications in various fields. Some of the notable applications include
- It has exclusive application in SNP detection, genotyping, and the detection of allelic variation.
- For the detection of mutant alleles, thereby helping in the early detection of diseases arising from mutations.
- Due to its mutational detection capabilities, it is used in diagnostics, such as for Beta-thalassemia and sickle cell anemia.
- It is also used to detect JAK2 mutations and variants of the Human Immunodeficiency Virus (HIV).
Advantages of Allele-Specific PCR (ASPCR)
The advantages of this technique include:
- It can clearly distinguish between the two allele variants.
- Single-nucleotide polymorphisms caused by base variation can be precisely detected.
- It can be utilized for genotyping.
- It is a fast, accurate, and reliable technique and thus is useful for diagnostic purposes.
- Precise distinguishing between homozygous and heterozygous alleles.
Limitations of Allele-Specific PCR (ASPCR)
Some of the limitations of this method include:
- It is a labor-intensive and complex technique due to primer design and the incorporation of mismatches.
- Detection of single-nucleotide polymorphisms is less at a time.
- Includes more primer sets, thus more expensive.
- Only the known SNPs can be detected. Unknown or new variation/mutation cannot be detected and analyzed.
- The chances of false-positive results are high.
- Temperature-sensitive method, thus a small alteration in the annealing temperature can affect the entire process.
- Cannot examine chromosomal alteration, larger mutations, and duplications.
Conclusion
Polymerase Chain Reaction is a gene amplification methodutlilsed fro the amplification and detection of a gene of interest from a sample. It is used in various diagnostic, medical, agricultural, and research fields to study nucleic acids.
Allele-specific Polymerase Chain Reaction, also known as Amplification-Refractory Mutation System (ARMS) PCR, is a variation of the conventional PCR method based on the principle that an allele has two variants: wild type and Mutant type.
By designing two sets of primers specific to each type and incorporating mismatches into them, the two forms can be distinguished. This method has been immensely useful in genotyping, detecting allelic variations, and detecting single-nucleotide polymorphisms.
References
- Imyanitov, E. N., Buslov, K. G., Suspitsin, E. N., Kuligina, E. S., Belogubova, E. V., Grigoriev, M. Y., … & Hanson, K. P. (2002). Improved reliability of allele-specific PCR. Biotechniques, 33(3), 484-490.
- Ugozzoli, L., & Wallace, R. B. (1991). Allele-specific polymerase chain reaction. Methods, 2(1), 42-48.
- https://geneticeducation.co.in/arms-or-allele-specific-pcr-principle-procedure-protocol-applications-and-limitations/#Variants_of_ARMS-PCR
- https://devyser.com/kits-and-reagents/devyser-cftr-core
- https://www.qiagen.com/us/products/discovery-and-translational-research/detection/ish-and-northern-blotting/snp-detection/type-it-fast-snp-probe-pcr-kit
- https://www.idtdna.com/pages/technology/qpcr-and-pcr/pcr-allele-competitive-extension-genotyping
