Firstly, we should consider the fact that DNA replication happens in a semiconservative way, meaning that each strand acts as a template for a new strand synthesis, and thus make a double-stranded DNA. One strand is a parent strand and the other is newly synthesized. DNA polymerase is the enzyme that is responsible for synthesizing this new DNA strand. They add nucleotides on the growing new strand, and this addition happens in a manner complementarily to the parent strand.
- DNA polymerase catalyzes the formation of phosphodiester bonds by adding deoxynucleoside triphosphates (dNTPs) onto the growing chain of a new strand
- In order for DNA polymerase to build and elongate the growing chain, it must have four types of dNTPs in the surroundings. These four dNTPs are deoxyadenosine triphosphate, deoxyguanosine triphosphate, deoxythymidine triphosphate, and deoxycytidine triphosphate.
- Or simply DNA polymerase is enzymes that create DNA molecules by adding nucleotides.
- DNA polymerase adds nucleotides in the 3’ end of newly formed DNA and elongates from 5’ to 3’. However, they cannot add nucleotide on their own and must need a preexisting free 3’ end to initiate the addition, which is provided by the primer.
- These primers are usually composed of RNA and DNA bases.
- Every time a cell divides, DNA polymerase is needed to help replicate the cell’s DNA. In doing so, a copy of parent DNA can be passed to each of the daughter cells, this way genetic information can be passed from generation to generation.
DNA Polymerase. Created with BioRender.com
Properties of DNA Polymerase
- DNA polymerase proofread. They check their work and cleave out unwanted or wrong nucleotides form the chain.
- DNA polymerase can only initiate the addition of nucleotide if there is a preexisting 3’ end.
- Therefore, DNA polymerase needs a preexisting short stretch of a nucleotide called primer, to provide that 3’ end.
- On the basis of type of cell, DNA polymerase has various types and differ in structure, function, and rate of polymerization, and processivity.
- In a prokaryotic cell, there are three DNA polymerases, namely from DNA polymerase Ι to ΙΙΙ. Out of which DNA polymerase ΙΙΙ is the main enzyme for replication in E.coli.
- In a eukaryotic cell, there are five main DNA polymerase: DNA Polymerase α, ε, δ, γ, and β.
- Since in eukaryotic cells, in addition to nuclear DNA, there is also the presence of mitochondrial DNA, which accounts for the fact that the replication of DNA happens in two different cellular compartments. Hence precise DNA polymerase accounts for a specific function.
Structure of DNA Polymerase
- We will only be discussing the structure of one of the most important DNA polymerases in eukaryotic cells i.e., family B. However, the structures of other DNA polymerases are mostly homologous and only differ in the direction of orientation of their subdomains.
- DNA polymerase resembles a right hand and consists of five subdomains: N terminus, exonucleases, palm, finger, and thumb. These domains are arranged in a disclike structure with a central cavity.
- The N terminus is composed of two nonadjacent segments of amino acids. However, its function is not clearly understood. These N terminus subdomains are also found in RNA binding proteins.
- The exonucleases subdomains lie between the thumb and N terminus. When an incorrect nucleotide is added to the 3’ end of the primer, the polymerase helps to switch the primer terminus towards the exonuclease subdomain, facilitating the cleavage of the 3’ terminal nucleotide residue. This subdomain consists of five stranded β-sheets surrounded by a distribution of α helices, the active site of the exonuclease subdomain is centered around a conserved motif. The carboxylate groups of these residues are coordinated to a magnesium ion that is essential for catalyzing the cleaving of mis-incorporated nucleotide base.
- The palm subdomain consists of the catalytic core responsible for polymerase activity. This subdomain catalyzes the nucleotidyl phosphoryl transfer reaction by serving as ligands for metals ions A and B.
- The finger and thumb subdomains resemble a pocket that has two regions: insertion site and postinsertion site. The incoming nucleotides bind to the insertion site and the new base pair resides in the postinsertion site.
Mechanism of DNA Polymerase
- When dNTPs are added to the growing chain, there is the formation of phosphodiester bonds and the release of pyrophosphates molecules, this reaction is catalyzed by two metal ions namely: metal ions A and B, both of which are Magnesium ions.
- The metal ion A is responsible for bringing the 3’OH group of the primer to close proximity with a phosphate atom of incoming dNTP.
- Due to which the 3’ free hydroxyl group of primer now nucleophilically attacks a phosphate atom of the triphosphate group of incoming dNTPs and forms a phosphodiester bond.
- As a bond is formed, there is a significant charge developed on the oxygen that was preciously bonded with phosphate atom, and to stabilize this charge metal ion B plays a vital role. Additionally, the metal ion B also assists in the departure of the pyrophosphate groups.
- Hence, the working mechanism of DNA polymerase is commonly referred to as two metal ion mechanisms.
Functions of DNA Polymerase
- The ultimate function of DNA polymerase is to synthesize new DNA by the process of replication. Due to which the transfer of genetic information from one generation to another is possible.
- It also contributes to repairing DNA, which rectifies any errors in the DNA.
- It is a proofreader, as it removes incorrect pairs of nucleotides with its exonuclease activity. This function is very important as it maintains the normal functionality of the body.
- These enzymes typically work in a pairwise manner, each enzyme replicates one of the two strands of double-helix DNA. One is the leading strand and another is the lagging strand named on the basis of their relative speed of synthesis.
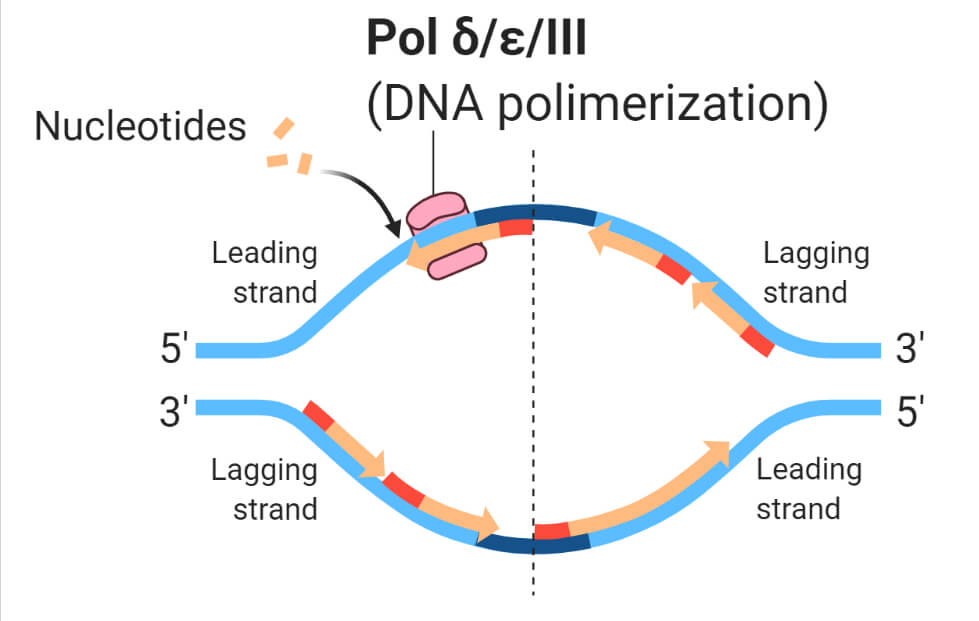
Types and Functions of DNA Polymerase in eukaryotic cell
DNA Polymerase α
- is a repair polymerase, with 3’ to 5’ exonucleases activities.
- has polymerase activity 5’ to 3’.
- also acts as a primase to synthesis primer.
DNA Polymerase β
- is a repair polymerase.
DNA Polymerase γ
- shows polymerase activity 5’ to 3’.
- exonucleases activity 3’ to 5’.
- involved in Mitochondrial DNA replication.
DNA Polymerase δ
- shows 3’ to 5’ exonuclease activity
- shows 5’ to 3’ polymerase activity.
- involved in lagging strand synthesis.
DNA Polymerase ε
- shows 3’ to 5’ and 5’ to 3’ exonucleases activities.
- it not only repair but also synthesize the leading strand efficiently in a 5’ to 3’ direction.
- prime enzyme for DNA replication.
Functions of DNA Polymerase in prokaryotic cell
DNA Polymerase Ι
- is a repair polymerase with both 3’ to 5’ and 5’ to 3’ exonuclease activity.
- polymerase activity in 5’ to 3’ direction.
- this enzyme is involved in the processing of Okazaki fragments generated by lagging strand synthesis.
DNA Polymerase ΙΙ
- has 3’ to 5’ exonuclease activity that participates in DNA repair.
- show polymerase activity from 5’ to 3’ direction.
DNA Polymerase ΙΙΙ
- the primary enzyme involved in DNA replication.
- has 5’ to 3’ polymerase activity.
- has 3’ to 5’ exonuclease activity.
References
- Xia, S., & Konigsberg, W. H. (2014). RB69 DNA polymerase structure, kinetics, and fidelity. Biochemistry, 53(17), 2752–2767. https://doi.org/10.1021/bi4014215
- Beard, W. A., & Wilson, S. H. (2014). Structure and mechanism of DNA polymerase β. Biochemistry, 53(17), 2768–2780. https://doi.org/10.1021/bi500139h
- Steitz T.A (1999) DNA Polymerases: Structural Diversity and Common Mechanisms. Journal of Biological Chemistry. Volume 274, Issue 25, P17395-17398 https://doi.org/10.1074/jbc.274.25.17395
- https://www.prospecbio.com/dna_polymerase
- Garcia-Diaz, M., & Bebenek, K. (2007). Multiple functions of DNA polymerases. Critical reviews in plant sciences, 26(2), 105–122. https://doi.org/10.1080/07352680701252817
- https://www.news-medical.net/life-sciences/DNA-Polymerase-Function.aspx

Thanks for this incredible information. however where I can copy your actual article resource ??