MacConkey Sorbitol Agar is based on the formulation described by Rappaport and Henigh. It is selective and differential media for the detection of sorbitol-nonfermenting Escherichia coli serotype O157:H7 associated with hemorrhagic colitis. E. coli serotype O157:H7 is a human pathogen associated with hemorrhagic colitis that results from the action of a Shiga-like toxin (SLT). On standard lactose-containing MacConkey Agar, this strain is indistinguishable from other lactose-fermenting E. coli. Unlike most E. coli strains, E. coli O157:H7 ferments sorbitol slowly or not at all. Therefore, MacConkey Agar with Sorbitol permits recognition of E. coli O157:H7 in stool cultures. The addition of cefixime and tellurite is a more selective and differential medium designed to inhibit Proteus mirabilis, non-O157 E. coli strains, and other sorbitol-nonfermenting strains that need to be screened during the attempted isolation of E. coli O157:H7.
Composition of Sorbitol MacConkey Agar
| Ingredients | Gm/L |
| Peptone | 17.0g |
| Proteose peptone | 3.0g |
| D-Sorbitol | 10.0g |
| Sodium Chloride | 5.0g |
| Bile Salts Mixture | 1.5g |
| Neutral Red | 0.03g |
| Crystal Violet | 0.001g |
| Agar | 13.5g |
Final pH 7.1 +/- 0.2 at 25ºC
Preparation of Sorbitol MacConkey Agar
- Suspend 50.03 g of the powder in 1000ml distilled water. Mix thoroughly.
- Heat with frequent agitation and boil for 1 minute to completely dissolve the powder.
- Autoclave at 121°C for 15 minutes.
- Cool to 45-50°C and pour into sterile Petri plates.
Principle of Sorbitol MacConkey Agar
MacConkey Sorbitol Agar is only slightly selective, since the concentration of bile salts, which inhibits gram-positive microorganisms, is low in comparison with other enteric plating media. Peptone and proteose peptone supply necessary nutrients like nitrogenous and carbonaceous compounds, long-chain amino acids, minerals, vitamins, and trace ingredients for the growth of organisms. Crystal violet present in the medium inhibits the growth of gram-positive bacteria, especially enterococci and staphylococci. Sodium chloride maintains osmotic equilibrium. Neutral red is an indicator. D-Sorbitol is a fermentable carbohydrate. Differentiation of enteric microorganisms is achieved by the combination of sorbitol and the neutral red indicator. The growth of E.coli O157:H7 on MacConkey Agar with Sorbitol shows colorless colonies and most of the fecal flora ferment sorbitol and appear pink. Colorless or pink to red colonies are produced depending upon the ability of the isolate to ferment the carbohydrate sorbitol.
Result and Interpretation on Sorbitol MacConkey Agar
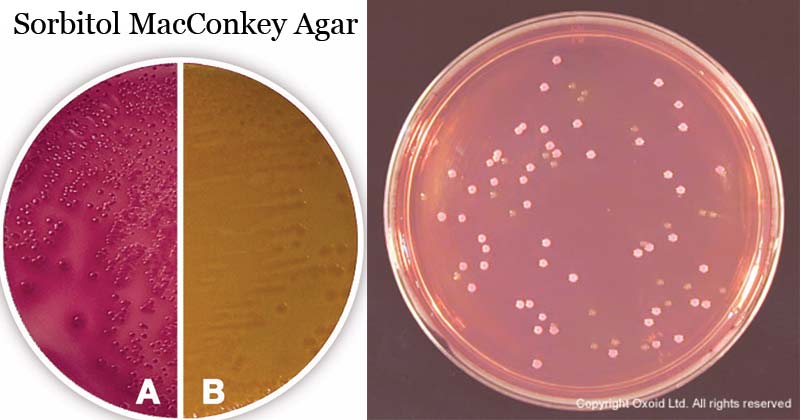
Figure A: Escherichia coli ATCC 25922 and Figure B: Escherichia coli O157:H7.
Source: Carl Roth and Oxoid
- After 18-24 hours of incubation, the plates should show isolated colonies in streaked areas and confluent growth in areas of heavy inoculation.
Sorbitol fermenters: pink to red colonies, some surrounded by zones of precipitated bile
Sorbitol non-fermenters: colorless colonies.
Uses of Sorbitol MacConkey Agar
- It is used for isolation and detection of enteropathogenic Escherichia coli O157: H7 the primary serovar associated with hemorrhagic colitis (HC) and hemolytic uremic syndrome (HUS).
- It aids in the differentiation of the E. coli O157: H7 from other strains, especially lactose fermenters E. coli.
Limitations of Sorbitol MacConkey Agar
- It has been reported that some Enterobacteriaceae and Pseudomonas aeruginosa are inhibited on MacConkey Agar when incubated in a CO2-enriched atmosphere.
- Prolonged incubation of the culture may result in colonies of E. coli serotype O157:H7 losing their characteristic colorless appearance and also the color of sorbitol-positive colonies can fade, making them hard to distinguish from sorbitol-negative colonies.
- Prolonged incubation can result in fading of pink-colored sorbitol-positive colonies making interpretation more difficult. Upon prolonged incubation, some strains of E. coli O157:H7 can ferment sorbitol and produce pink-colored colonies.
- There are additional species of facultatively anaerobic gram-negative rods that do not ferment sorbitol.
- Although most non-O157:H7 coli ferment sorbitol, about 6% of the isolates do not. These atypical strains along with other sorbitol non-fermenting bacteria such as Morganella and Hafnia appear identical to O157:H7 colonies and therefore confirmatory tests may be necessary.
- There exist sorbitol negative strains of serotypes other than O157:H7 which may or may not produce toxins and clinical symptoms. MacConkey Agar with Sorbitol does not differentiate between toxin-producing and nonproducing strains of E. coli O157.
- MacConkey Sorbitol Agar, however, should not be solely used to detect pathogenic E.coli O157: H7 strains as some nontoxic strains will also not ferment sorbitol. Gram staining, biochemical tests, and serological procedures should be performed to confirm findings.
References
- Himedia
- Dalynn Biologicals
- Hardy Diagnostics
- Thermo Fisher Scientific Inc.
- Becton, Dickinson and Company
- Conda lab
- Tille P.M (2014)Bailey and Scott’s diagnostic microbiology, Thirteen edition, Mosby, Inc., an affiliate of Elsevier Inc., 3251 Riverport Lane, St. Louis, Missouri 63043
- Ronald M. Atlas and James W. Snyder (2014). Handbook of media for clinical and public health microbiology. CRC Press. Taylor & Francis Group, LLC. Page no.287.

What is the maximum limit of detection of Mackonkey?
Thank you