The ribosome word is derived – ‘ribo’ from ribonucleic acid and ‘somes’ from the Greek word ‘soma’ which means ‘body’.
Ribosomes are tiny spheroidal dense particles (of 150 to 200 A0 diameters) that are primarily found in most prokaryotic and eukaryotic.
- They are sites of protein synthesis.
- They are structures containing approximately equal amounts of RNA and proteins and serve as a scaffold for the ordered interaction of the numerous molecules involved in protein synthesis.
- The ribosomes occur in cells, both prokaryotic and eukaryotic cells.
- In prokaryotic cells, the ribosomes often occur freely in the cytoplasm.
- In eukaryotic cells, the ribosomes either occur freely in the cytoplasm or remain attached to the outer surface of the membrane of the endoplasmic reticulum.
- The location of the ribosomes in a cell determines what kind of protein it makes.
- If the ribosomes are floating freely throughout the cell, it will make proteins that will be utilized within the cell itself.
- When ribosomes are attached to the endoplasmic reticulum, it is referred to as rough endoplasmic reticulum or rough ER.
- Proteins made on the rough ER are used for usage inside the cell or outside the cell.
- The number of ribosomes in a cell depends on the activity of the cell.
- On average in a mammalian cell, there can be about 10 million ribosomes.
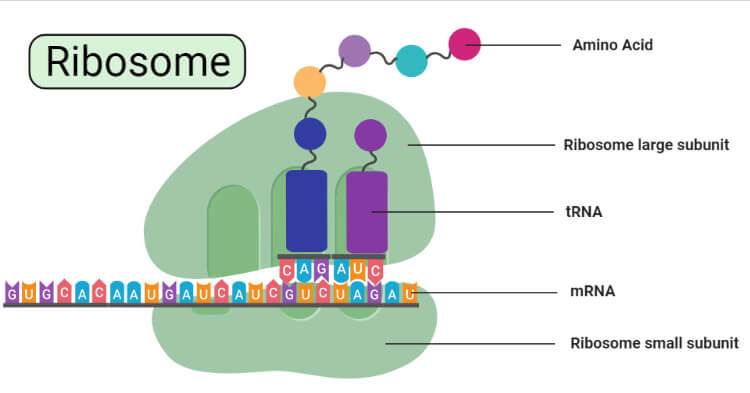
Structure of Ribosomes
- A ribosome is made from complexes of RNAs and proteins and is, therefore, a ribonucleoprotein.
- Around 37 to 62% of RNA is comprised of RNA and the rest is proteins.
- Each ribosome is divided into two subunits:
- A smaller subunit which binds to a larger subunit and the mRNA pattern, and
- A larger subunit which binds to the tRNA, the amino acids, and the smaller subunit.
- Prokaryotes have 70S ribosomes respectively subunits comprising the little subunit of 30S and the bigger subunit of 50S.
- Their small subunit has a 16S RNA subunit (consisting of 1540 nucleotides) bound to 21 proteins.
- The large subunit is composed of a 5S RNA subunit (120 nucleotides), a 23S RNA subunit (2900 nucleotides) and 31 proteins.
- Eukaryotes have 80S ribosomes respectively comprising of little (40S) and substantial (60S) subunits.
- The smaller 40S ribosomal subunit is prolate ellipsoid in shape and consists of one molecule of 18S ribosomal RNA (or rRNA) and 30 proteins (named as S1, S2, S3, and so on).
- The larger 60S ribosomal subunit is round in shape and contains a channel through which growing polypeptide chain makes its exit.
- It consists of three types of rRNA molecules, i.e., 28S rRNA, 5.8 rRNA and 5S rRNA, and 40 proteins (named as L1, L2, L3 and so on).
- The differences between the ribosomes of bacterial and eukaryotic are used to create antibiotics that can destroy bacterial infection without harming human cells.
- The ribosomes seen in the chloroplasts of mitochondria of eukaryotes are comprised of big and little subunits composed of proteins inside a 70S particle.
- The ribosomes share a core structure that is similar to all ribosomes despite differences in its size.
- The two subunits fit together and work as one to translate the mRNA into a polypeptide chain during protein synthesis.
- Because they are formed from two subunits of non-equal size, they are slightly longer in the axis than in diameter.
- During protein synthesis, when multiple ribosomes are attached to the same mRNA strand, this structure is known as polysome.
- The existence of ribosomes is temporary, after the synthesis of polypeptide the two sub-units separate and are reused or broken up.
Types of Ribosomes
Based on the size and the sedimentation coefficient (S), ribosomes are of two types:
- 70S ribosome
- 80S ribosome
70S ribosome
- They are smaller in size.
- Sedimentation coefficient: 70S
- Molecular weight: 2.7× 106 daltons.
- They are found in:
- prokaryotic cells of the blue-green algae and bacteria.
- mitochondria and chloroplasts of eukaryotic cells.
80S ribosome
- Sedimentation coefficient: 80S
- Molecular weight: 40 × 106 daltons.
- They are found in the eukaryotic cells i.e. in plants and animals.
- The ribosomes present in mitochondria and chloroplasts are smaller than 80S cytoplasmic ribosomes.
- In the 80S ribosome of yeast, 79r-protein are present where only 12 r-protein are found to be specific.
Chemical Composition of Ribosomes
- Ribosomal RNAs
- Ribosomal proteins
- Metallic ions
Ribosomal RNAs
- 70S ribosomes consist of three types of rRNA:
- 23S rRNA
- 16S rRNA
- 5S rRNA
- In the 50S ribosomal subunit (larger subunit), 23S and 5S rRNA are present.
- In the 30S ribosomal subunit, the 16S rRNA is present.
- In the 80S ribosomes four types of rRNA are present:
- 28S rRNA
- 18S rRNA
- 5S rRNA
- 5.8 rRNA
- In the larger 60S ribosomal subunit, 28S, 5S, and 5.8S rRNAs are present.
- There is the presence of 18S rRNA in the 40S ribosomal subunit ( Smaller)
Ribosomal proteins
- Bacteria are composed of different ribosomal proteins.
- It was found that E. coli consists of 55 ribosomal proteins.
- For example; Core proteins (CP), Split proteins (SP)
Metallic ions
- divalent metallic ions: Mg++, Ca++ and Mn++
Functions of Ribosomes
- The ribosome is a complex molecular machine, found within all living cells, that serves as the site of biological protein synthesis (translation).
- Ribosomes link amino acids together in the order specified by messenger RNA (mRNA) molecules.
- Ribosomes act as catalysts in two extremely important biological processes called peptidyl transfer and peptidyl hydrolysis.
- The nascent polypeptide chain is protected from the activity of protein digestive enzymes.
How does the ribosomal movement take place in translation?
- In addition to a binding site for an mRNA molecule, it consists of other 3 binding sites. for tRNA molecules;
- A site
- P site
- E site
- Amino acid needs to be added to a growing peptide chain.
- The charged tRNA whose base pairs with the complementary codon on the mRNA molecule enters the A site.
- Then in the new forming polypeptide chain, an amino acid is added which is held by the tRNA in the adjacent P site.
- Then the large ribosomal subunit moves forward to the E site.
- This process or the cycle is repeated.
- Each time an amino acid is added to the polypeptide chain, where the new protein grows from its amino to its carboxyl end.
- Finally, the stop codon will be encountered in the mRNA.
- The termination release factors like the RF1 and RF2 recognize the stop codons.
- Then the peptidyl-tRNA bond is hydrolyzed.
- Finally, the newly formed polypeptide is released from the ribosome.
- By the study of the structure and its biochemical characteristics, antibacterial agents are developed in such a way they can inhibit this protein synthesis process.
- Examples of such antibiotics are:
- Aminoglycosides
- Chloramphenicol
- Fusidic acids
- Lincosamides
- Macrolides
- Oxazolidinone
- Streptogramins
- Tetracyclines
References
- Verma, P. S., & Agrawal, V. K. (2006). Cell Biology, Genetics, Molecular Biology, Evolution & Ecology (1 ed.). S .Chand and company Ltd.
- Alberts, B. (2004). Essential cell biology. New York, NY: Garland Science Pub.
- Kar,D.K. and halder,S. (2015). Cell biology genetics and molecular biology.kolkata, New central book agency
- Wilson, D. N. (2014). Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nature Reviews Microbiology, 12(1), 35–48. https://doi.org/10.1038/nrmicro3155
- Wilson, D. N., & Cate, J. H. D. (2012). The structure and function of the eukaryotic ribosome. Cold Spring Harbor Perspectives in Biology, 4(5), 5. https://doi.org/10.1101/cshperspect.a011536
- https://biology.tutorvista.com/animal-and-plant-cells/ribosomes.html
- https://alevelbiology.co.uk/notes/ribosomes-structure-and-functions/
- https://biologywise.com/ribosomes-function
- https://biologydictionary.net/ribosome/

awesome 👌
YOUR MATERIAL IS SUPURUB
You didn’t really give me a picture of ribosomes.