- DNA replication is the process by which an organism duplicates its DNA into another copy that is passed on to daughter cells.
- Replication occurs before a cell divides to ensure that both cells receive an exact copy of the parent’s genetic material.
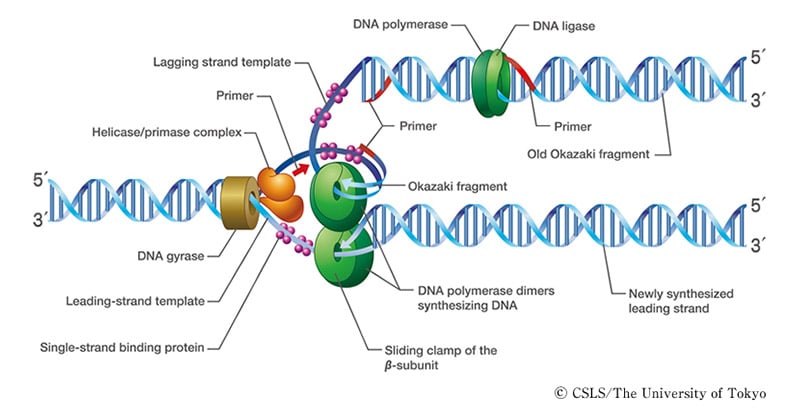
Features of Eukaryotic DNA Replication
- Replication is bi-directional and originates at multiple origins of replication (Ori C) in eukaryotes.
- DNA replication uses a semi-conservative method that results in a double-stranded DNA with one parental strand and a new daughter strand.
- It occurs only in the S phase and at many chromosomal origins.
- Takes place in the cell nucleus.
- Synthesis occurs only in the 5′to 3′direction.
- Individual strands of DNA are manufactured in different directions, producing a leading and a lagging strand.
- Lagging strands are created by the production of small DNA fragments called Okazaki fragments that are eventually joined together.
- Eukaryotic cells possess five types of polymerases involved in the replication process.
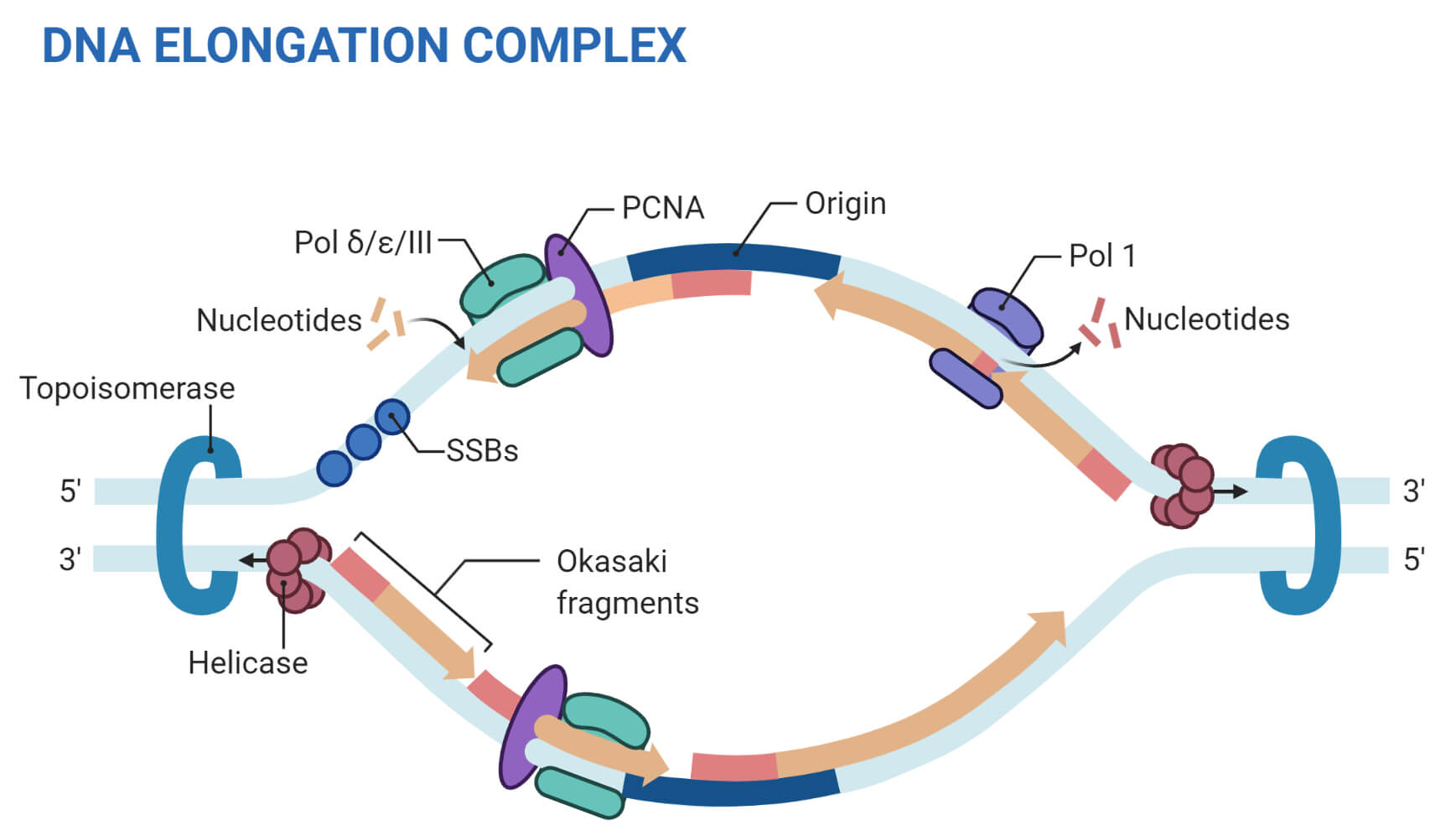
THE ENZYMES OF DNA REPLICATION
- Helicases: Unwind the DNA helix at the start of replication.
- SSB proteins: Bind to the single strands of unwound DNA to prevent reformation of the DNA helix during replication.
DNA Polymerases:
- Eukaryotic cells contain five different DNA polymerases; α, β, γ, δ and ε.
- DNA polymerases α and δ replicate chromosomal DNA, DNA polymerases β and ε repair DNA, and DNA polymerase γ replicates mitochondrial DNA.
- DNA polymerase α and δ synthesize the lagging strand, via Okazaki fragments.
- The RNA primers are synthesized by DNA polymerase α which carries a primase subunit.
- DNA polymerase ε synthesizes the leading strand.
- Telomerase, a DNA polymerase that contains an integral RNA that acts as its own primer, is used to replicate DNA at the ends of chromosomes (telomeres).
- DNA topoisomerase I: Relaxes the DNA helix during replication through creation of a nick in one of the DNA strands.
- DNA topoisomerase II: Relieves the strain on the DNA helix during replication by forming supercoils in the helix through the creation of nicks in both strands of DNA.
- DNA ligase: Forms a 3′-5′phosphodiester bond between adjacent fragments of DNA.
Process of Eukaryotic DNA Replication
- Replication of each linear DNA molecule in a chromosome starts at many origins, one every 30–300 kb of DNA depending on the species and tissue, and proceeds bi-directionally from each origin.
- At each origin, a replication bubble forms consisting of two replication forks moving in opposite directions. The DNA replicated under the control of a single origin is called a replicon. DNA synthesis proceeds until replication bubbles merge together.
- At the origin, enzymes unwind the double helix making its components accessible for replication.
- The helix is unwound by helicase to form a pair of replication forks.
- The unwound helix is stabilized by SSB proteins and DNA topoisomerases.
- The RNA primers required are made by DNA polymerase α which carries a primase subunit.
- DNA polymerase α initiates synthesis of the lagging strand, making first the RNA primer and then extending it with a short region of DNA.
- DNA polymerase δ then synthesizes the rest of the Okazaki fragment.
- The leading strand is synthesized by DNA polymerase ε.
- The leading strand is synthesized continuously in the 5′to 3′ direction while the lagging strand is synthesized discontinuously in the 5′to 3′ direction through the formation of Okazaki fragments.
- At the completion of synthesis, DNA ligase seals the breaks between the Okazaki fragments as well as around the primers to form continuous strands.
DNA Proofreading
In eukaryotes only the polymerases that deal with the elongation (delta and epsilon) have proofreading ability (3’ → 5’ exonuclease activity).
If an error is detected, the erroneous base is removed via 3′to 5′exonuclease activity replaced with the correct base.
Excision repair:
Removes pyrimidine dimers formed by UV rays or other mutated bases and replaces them.
Significance of Eukaryotic DNA Replication
- DNA replication is a fundamental genetic process that is essential for cell growth and division.
- DNA replication involve the generation of a new molecule of nucleic acid, DNA, crucial for life.
- DNA replication is important for properly regulating the growth and division of cells.
- It conserves the entire genome for the next generation.
References
- David Hames and Nigel Hooper (2005). Biochemistry. Third ed. Taylor & Francis Group: New York.
- Bailey, W. R., Scott, E. G., Finegold, S. M., & Baron, E. J. (1986). Bailey and Scott’s Diagnostic microbiology. St. Louis: Mosby.
- Madigan, M. T., Martinko, J. M., Bender, K. S., Buckley, D. H., & Stahl, D. A. (2015). Brock biology of microorganisms (Fourteenth edition.). Boston: Pearson.
- https://en.wikipedia.org/wiki/Proofreading_(biology)
- https://sciencing.com/comparing-contrasting-dna-replication-prokaryotes-eukaryotes-13739.html

Question…
SSB proteins are present in prokaryotes such as Escherichia coli.
Are proteins homologous to SSBs in eukaryotic cells called RPAs, or not?
Yes, the proteins homologous to SSBs (Single-Stranded DNA-Binding Proteins) in prokaryotic cells, such as Escherichia coli, are called RPAs (Replication Protein A) in eukaryotic cells.
DNA polymerase ε synthesizes the leading strand, not polymerase δ as written above.
Dear Chean,
Thanks for the correction. The page has been updated. 🙂
Very informative, i like to learn from it.