Isomerase enzymes form or assist in forming isomers of any biological components. They assist in the rearrangement process of different biomolecules during the formation or breakage of bonds. Topoisomerase is a type of isomerase enzyme.
What is Topoisomerase?
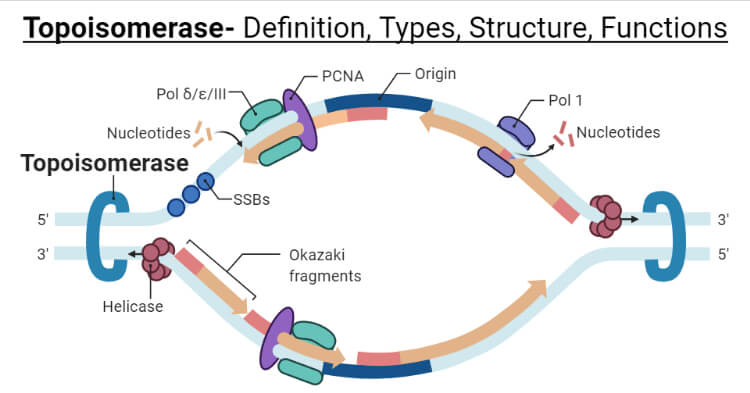
Topoisomerase is an essential enzyme that aids in the DNA replication process, segregation of chromosomes, transcription, and also in recombination.
- It was first found by J.C. Wang in the 1970s while working on Escherichia coli. It was the type I topoisomerase.
- As the name suggests, it helps in changing the DNA topology. It can increase or decrease the extent of the unwinding of DNA.
- It can also be called DNA topoisomerase as it only acts on DNA strands.
- It doesn’t work on RNA.
- It breaks the phosphodiester bond that is present in the backbone of DNA strands. The bonds are formed again as the enzyme leaves.
Some Important Terms
Twist (Tw): It is the total count of helical turns of the strands of DNA.
Writhe (Wr): It is the total count of turns of the double helix of DNA crossing on itself that indicates the supercoils of DNA.
Linking Number: It is the total number or addition of twists and writhes in DNA.
Linking No. = Wr+Tw
Topoisomerase Types
There are two types of topoisomerases:
- Type I Topoisomerase
- Type II Topoisomerase
Type I Topoisomerase
Type I Topoisomerase Definition
Type I topoisomerase is a type of topoisomerase that cuts on a single strand of DNA. It is not an ATP-dependent enzyme(exception: Reverse Gyrase).
It mainly changes the linking number by plus one.
Note: Odd types of topoisomerases come under type I and even types under type II.
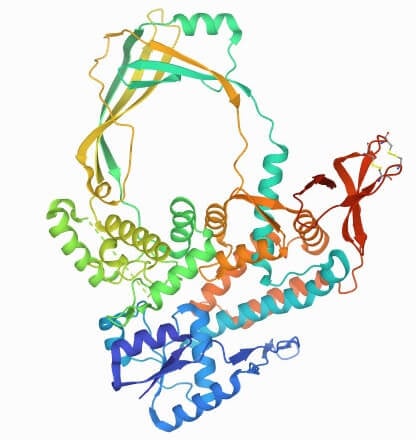
Type I Topoisomerase Structure
There is the presence of multiple varying domains in the type IA. It can be from I to IV. Toprim domain is contained in domain I. HTH (Helix-Turn-Helix) is present in domains III and IV. The tyrosine residues are present in the HTH of domain III. It appears like a lock with all three domains present at bottom of the topoisomerase structure.
Type IB contains active site (tyrosine) bind with C-terminal domain, N-terminal domain, capping, and catalytic lobe.
Type I Topoisomerase Types
It is of three basic types:
Type IA topoisomerases
It binds to the 5′ Carbon end of the DNA.
This type of topoisomerase show homology to topoisomerase I of E. coli.
It is of further three types:
- Topo IA: It is found in eubacteria.
- Topo III: It is found in eubacteria and eukaryotes.
- Reverse Gyrase: It is found in archaebacteria and eubacteria as well. It is the only type of type I topoisomerase that is ATP-dependent.
(Here topo indicates topoisomerase)
Type IB topoisomerases
It binds to the 3′ Carbon end of the DNA. It forms nick in one strand. This type of topoisomerase show homology to topoisomerase I of humans.
Type IC topoisomerases
It contains one type of topoisomerase i.e. topoisomerase V. It binds to the 3′ Carbon end of the DNA. It is found in archaebacterial. It shows the controlled mechanism of rotation.
Type I Topoisomerase Mechanism of action
It generally occurs in the following events occurring together at the same time.
- Cutting a single strand of DNA: Active site of the topoisomerase contains an amino acid tyrosine. The disruption of phosphodiester bond and formation of intermediate with phospho-tyrinosyl linkage favors the breaking of a DNA strand. The bond formation and cleavage mechanism in detail are the same as in the case of type II topoisomerase. Tyrosine may attack 3’or 5’carbon end.
- Passing of strand: After the cleavage, the uncut DNA strand passes through the break. In this step, the enzyme changes from closed conformation to open conformation favoring the passing of strand. No ATP is utilized in this conformational change in the case of type I.
- Religation: The phosphate linked with tyrosine is again attacked by the OH of the ribose group of the strand which was separated before and it results in the removal of intermediate linkage of tyrosine and rejoining of the cleaved strand. The enzyme returns to its initial stage(closed conformation) and is recovered for the next cycle.
Type I Topoisomerase Functions
- They are involved in the removal of supercoils of DNA in biological processes such as replication and transcription.
- Help in relaxing DNA.
- They help in breaking strands during recombination.
- They are also involved in the condensation of the chromosome.
- During mitosis, the DNA strands need to be free from interwinding which is done by topoisomerase I.
Type II Topoisomerase
Type II Topoisomerase Definition
Type II topoisomerase is a type of topoisomerase that cuts on both strands of DNA at once. It is an ATP-dependent enzyme. It changes the linking number by two.
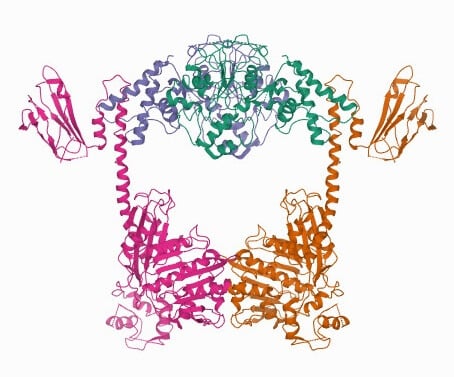
Type II Topoisomerase Structure
Topoisomerase IIA in eukaryotes consists of two same monomers (A-A) whereas in prokaryotes they are formed heterotetramers (A2B2).
Topoisomerase IIB is formed of heterotetramers only.
Topoisomerase II consists of four domains which include:
- ATPase domain at N-terminal
- A variable C-terminal domain
- Domain for binding of DNA located centrally
- A conserved domain of about a hundred amino acids i.e. toprim domain.
Type II Topoisomerase Types
It is of two basic types:
Type IIA topoisomerases
It is found in viruses and all cellular organisms. It is of three types:
- Topo II: It is found in eukaryotes.
- Topo IV: It is found in bacteria. It differs from Gyrase. It is not involved in DNA wrapping while Gyrase is involved in DNA wrapping and promoting negative supercoils.
- Gyrase: It is found in bacteria and some eukaryotes. It introduces negative supercoiling decreasing the linking number by two.
Type IIB topoisomerases
It includes Topo VI which can be found in archaea and some plants.
Type II Topoisomerase Mechanism of action
It occurs as follows with ATP hydrolysis.
- Cleaving of DNA chain: The enzyme contains tyrosine residues. They form covalent bonds with the DNA strands and break the DNA chain. The lone pair of electrons of O-atom present in the tyrosine acts as a nucleophile and attacks on the Phosphorus in phosphate of DNA. It causes the shifting of a bond from phosphate to one of the O-atom attached to the ribose sugar forming a hydroxyl group. Hence the covalently bonded tyrosine attached with phosphorus breaks the phosphate-sugar backbone which cleaves the chain. This linking is termed 5′-phospho-tyrinosyl protein-DNA linkage.
A duplex is broken by the action of the enzyme on both strands at once.
- Crossing of the intact strand through the gap: In this case, another whole duplex strand passes through the gap over the broken duplex. In this the conformational change in enzyme requires ATP.
- Religation: It is done by the attack of 3′-OH of the sugar of separated strand on phosphate group which has formed an intermediate linkage with tyrosine. It repels the bond with tyrosine and reforms the broken bond to join again. It occurs on both strands of duplex together ligating them. The enzymes regain their conformation and continue the cycle.
Type II Topoisomerase Functions
- It increases the disentanglement of the chromosome.
- It does not aid in the supercoiling of DNA but is involved in their relaxation.
- DNA gyrase promotes the negative supercoils of DNA.
- One of the most important functions is that it brings the change of two units in the linking number of loops in DNA.
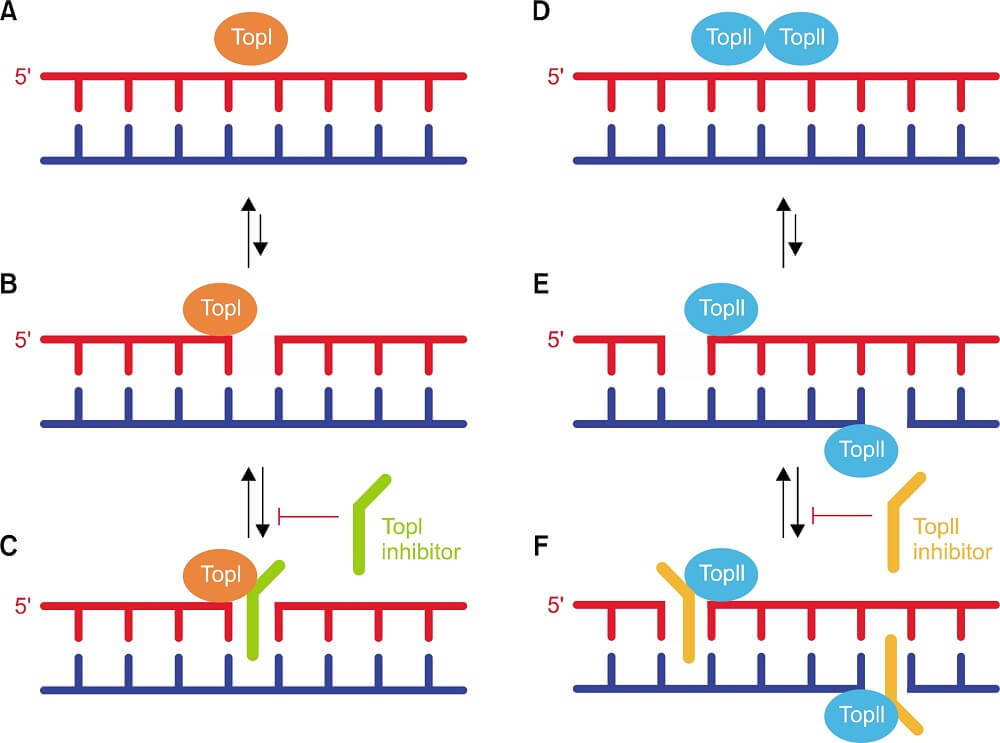
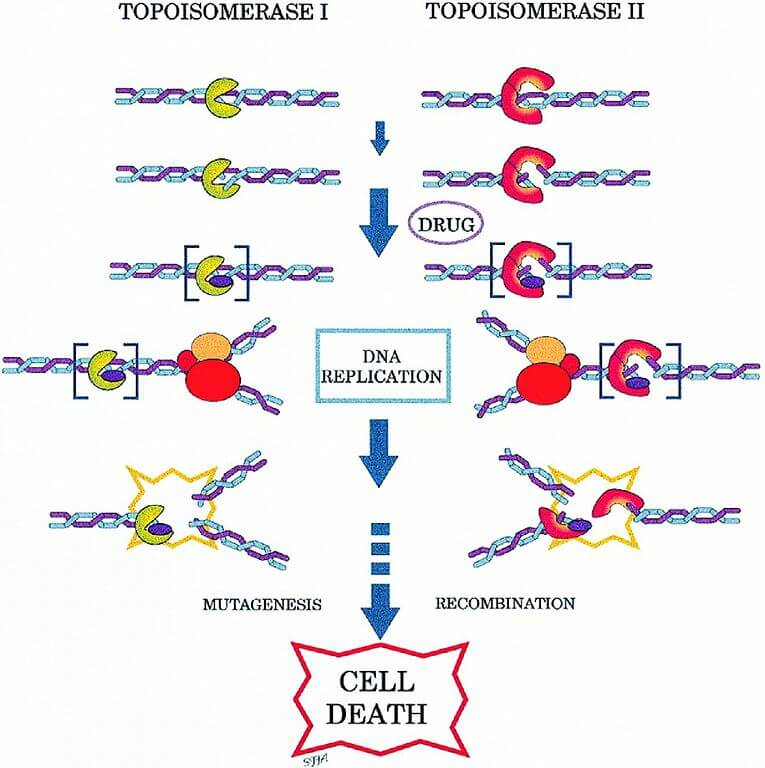
Topoisomerase Inhibition
Some chemical components can suppress the action of topoisomerase and are called topoisomerase inhibitors.
They can interfere with the ligation step of DNA which creates broken strands in the cell causing the death of the cell by apoptosis.
The topoisomerase inhibition principle is used for the development of drugs for bacterial infection. It includes antibiotics such as novobiocin, coumermycin of the class coumarins which interfere in ATP binding in type II topoisomerases in bacteria leading to its death. It also includes the quinolone class of antibiotics which prevent the religation of nicked DNA strands in the last step of the topoisomerase working mechanism.
Chemotherapeutic agents applied for the treatment of cancer can lead to inhibition of topoisomerase in humans. They can stabilize the intermediate formed by the linkage of tyrosine of topoisomerase and phosphate of DNA.
Clinical Significance of Topoisomerase
Many medications work by interfering with type II topoisomerases in bacteria. These medications include broad-spectrum antibiotics such as fluoroquinolones. They can make the topoisomerase damage the DNA.
Cancer cell topoisomerases are targeted by some chemotherapy medications such as Irinotecan and topotecan for type I and teniposide and etoposide for type II which are also called topoisomerase inhibitors.
In an autoimmune disorder Scleroderma, the Anti-topoisomerase antibodies(also called anti-scl-70 antibodies) can be seen against the topoisomerase I antigen.
Topoisomerase vs Helicase
| Topoisomerase | Helicase |
| It is involved in the prevention of supercoiling of DNA i.e. decreases tension on the unwound strands. | It is involved in the unwinding of DNA strands. |
| It works on DNA only. | It works acts on DNA and RNA. |
| It attacks the phosphodiester bond in the backbone of DNA. | It attacks the Hydrogen bonds between the double strands. |
| Its two types are: Type I Topoisomerase Type II Topoisomerase | Its two types are: RNA helicase DNA helicase |
Topoisomerase vs Gyrase
| Topoisomerase | Gyrase |
| It includes different types of enzymes including Gyrase. | Gyrase is a type of topoisomerase. |
| It is a large class of enzymes. | It is a type within the sub-class of Type II topoisomerase. |
| It is present in both prokaryotes and eukaryotes. | It is mostly present in prokaryotes and only in some eukaryotes. |
| It maintains the topology of DNA by the combined functions of different types of enzymes. It includes both negative and positive supercoiling of DNA. | Its specific function is to introduce negative supercoiling in DNA strands rather than to remove them. |
| Topoisomerases may or may not be ATP-dependent. | Gyrase is an ATP-dependent type of topoisomerase. |
References
- Nitiss, J. L., Soans, E., Rogojina, A., Seth, A., & Mishina, M. (2012). Topoisomerase assays. Current protocols in pharmacology, Chapter 3, Unit3.3–3.3.. https://doi.org/10.1002/0471141755.ph0303s57
- Levine, C., Hiasa, H., & Marians, K. J. (1998). DNA gyrase and topoisomerase IV: biochemical activities, physiological roles during chromosome replication, and drug sensitivities. Biochimica et Biophysica Acta, 1400(1-3), 29–43. https://doi.org/10.1016/s0167-4781(98)00126-2
- Wang JC (June 2002). Cellular roles of DNA topoisomerases: a molecular perspective. Nat. Rev. Mol. Cell Biol. 3 (6): 430–40. doi:10.1038/nrm831
- Sharma A; Hanai R; Mondragón A (August 1994). Crystal structure of the amino-terminal fragment of vaccinia virus DNA topoisomerase I at 1.6 A resolution. Structure. 2 (8): 767–77. doi:10.1016/s0969-2126(94)00077-8
- Cretaio, Erica & Pattarello, Luca & Fontebasso, Yari & Benedetti, Piero & Losasso, Carmen. (2007). Human DNA topoisomerase IB: Structure and functions. The Italian journal of biochemistry. 56. 91-102.
- Shinji Kuroda, … Toshiyoshi Fujiwara, Gene Therapy of Cancer (Third Edition), 2014.
- Seo YH. Dual Inhibitors Against Topoisomerases and Histone Deacetylases. J Cancer Prev 2015;20:85-91. https://doi.org/10.15430/JCP.2015.20.2.85

this is very helpful information. thank you so much
Very helpful
Este es una información muy completa
This is really too helpful .thank u so so much .