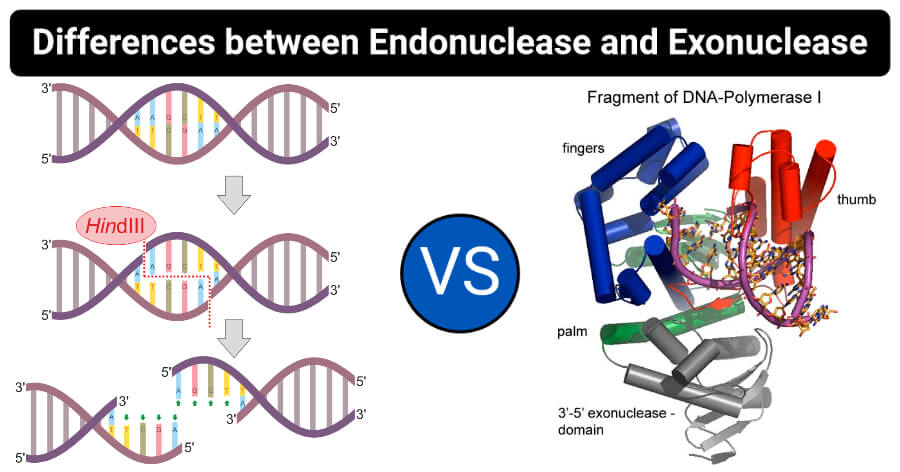
Image Source: Helixitta and Christopherrussell.
Endonuclease Definition
An endonuclease is a group of enzymes that cleave the phosphodiester bond present within a polynucleotide chain.
- Endonucleases are capable of breaking the bond from the middle of a chain.
- These enzymes are either specific or non-specific to the sequences being cleaved. The endonucleases that are specific to a particular sequence are termed restriction endonucleases.
- Restriction endonucleases are obtained from various bacteria and archaea, each of which is specific for different sites in the polynucleotide chain.
- The cleaved sequences form single-stranded ends, also called sticky ends, which are then hybridized by DNA ligase. The resulting DNA is called recombinant DNA, and the process is called recombination.
- The restriction endonucleases are one of the most important groups of endonucleases which are divided into three types based on their mechanism of action.
- Type I endonucleases are large multi-subunit complexes that cleave at random sites of about 1000 base pairs. Type II endonucleases are smaller subunits that cleave the sequences in the absence of ATP. Type III endonucleases are also complex that cleave the sequences of 25 base pairs.
- The endonuclease can cleave double-stranded DNA, single-stranded DNA, or even RNAs.
- These enzymes are vital for DNA repair as they allow the recognition and cleavage of damaged DNA precisely.
- This prevents unnecessary cleavage and further damage that might occur with other enzymes.
- Restriction endonucleases might suffer from a lag period before their action. Such a lag period might be due to the time required for the recognition of the specific sites.
- Because endonuclease cleaves a DNA segment in the middle, it results in oligonucleotides.
- Some endonuclease has a defensive function where they can prevent the entry of some pathogens.
Exonuclease Definition
Exonucleases are enzymes that cleave DNA sequences in a polynucleotide chain from either the 5’ or 3’ end one at a time.
- Exonuclease, like endonuclease, is a hydrolyzing enzyme that cleaves the phosphodiester bond between the nucleotides.
- One of the most important routes of RNA degradation in both archaea and eukaryotes is by the multi-protein exosome, which consists of multiple exoribonucleases.
- Besides, exonucleases are also found in the venoms of snakes and lizards. These toxins work by the cleavage of DNA coding for essential proteins within the body.
- Exonulceases are important during replication as one of these enzymes works together with RNA polymerase II degrade the newly formed RNA primer present on the new transcript which is then replaced by DNA nucleotides.
- Exonuclease activity is also exploited during editing and proofreading DNA for errors.
- The exonuclease in prokaryotes and eukaryotes are of three types; a decapping 5’ to 3’ exonuclease (Xrn1), an independent 5’to 3’ exonuclease and a polyA-specific 3’ to 5’ exonuclease.
- All of these exonucleases are involved in the formation, replication, and transcription of RNAs.
- Exonucleases, unlike endonucleases, do not have a lag period as they cleave the sequences from the ends, resulting in sticky ends.
- Similarly, exonucleases also cleave individual nucleosides from either of the ends instead of resulting in oligonucleotides.
- Exonuclease does not have defensive properties against the entry of pathogenic microbes.
Key Differences (Endonuclease vs Exonuclease)
Basis for Comparison |
Endonuclease |
Exonuclease |
| Definition | An endonuclease is a group of enzymes that cleave the phosphodiester bond present within a polynucleotide chain. | Exonucleases are enzymes that cleave DNA sequences in a polynucleotide chain from either the 5’ or 3’ end one at a time. |
| Cleavage | Endonucleases cleave the nucleotide sequence from the middle. | Exonulceases cleave a nucleotide sequence from the ends. |
| Lag period | Some endonucleases like restriction endonucleases have a lag period before their activity. | Exonuclease does not have a lag period before their activity. |
| Results in | Endonucleases cleave DNA sequences, resulting in oligonucleotides. | Exonulceases cleave DNA sequences, resulting in individual nucleotides or nucleosides. |
| Ends | Endonucleases might form either sticky or blunt ends. | Exonulceases form sticky ends. |
| Specificity | Specific endonucleases, also called restriction endonucleases, are available that cleave specific sites within a DNA sequence. | Exonuclease is usually non-specific. |
| Defensive properties | Endonucleases have defensive properties against the entry of pathogenic microorganisms. | Exonulceases do not have defensive properties. |
| Effect on circular DNA | Restriction endonuclease can cleave specific sites within a circular DNA. | Exonulceases have less activity towards circular DNA as compared to linear DNA. |
| Inhibition | Endonucleases cannot be inhibited phosphorothioate bonds unless the entire sequence has the bonds between all nucleotides. | Exonuclease can be inhibited by adding five phosphorothioate bonds in a row to a sequence. |
| Free ends | Free 3’ or 5’ ends are not necessary for the action of endonucleases. | The ends should be free for the action of exonucleases. |
| Examples | EcoRI, BamHI, Deoxyribonulcease I are some examples of endonucleases. | Snake venom, Exonuclease I, Xrn1 are some examples of exonucleases. |
Examples of endonucleases
EcoRI
- EcoRI is a restriction endonuclease that cleaves helical structures of DNA at specific sites to form fragments.
- This enzyme is isolated from the E. coli species and is a part of the restriction-modification system.
- EcoRI cleaves the G/AATTC where the ‘/’ indicates the phosphodiester bond that is specific for the cleavage by the restriction enzyme.
- It results in a four nucleotide sticky end with a 5’ overhang of AATT.
- EcoRI is a homodimer with a 31 kilodalton subunit of a globular domain with α/β architecture.
- EcoRI has been widely used for various molecular biology techniques including cloning, DNA screening, and error removal.
- Cleavage of DNA results in sticky ends which enhances the action of ligase enzyme and makes the ligation reaction more efficient.
- Non-specific cleavage can be done with EcoRI when the medium has low salt and enzyme concentration.
Examples of exonucleases
Xrn1
- Xrn1 is an exoribonuclease that is a 5’ to 3’ exonuclease found in humans.
- It is coded by the Xrn1 gene and hydrolyzes RNAs in the 5’ to 3’ direction.
- It is involved in the replication-dependent histone mRNA degradation by interacting directly with mRNA decapping protein.
- Xrn1 is also involved in many nuclear and cytoplasmic functions ranging from transcription, translation recombination, and meiosis in both humans and yeast cells.
- It also initiates transcriptional termination of DNA and RNA to prevent the overbuilding of thee nucleotide chains.
- Mutations in this enzyme are often associated with osteosarcoma, suggesting the possible role of this enzyme in bone formation and processing.
Video Lecture – Nucleases | Exonucleases | Endonucleases (Dhara Fatnani)
References and Sources
- 3% – https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/endonuclease
- 2% – https://pediaa.com/difference-between-endonuclease-and-exonuclease/
- 2% – https://en.wikipedia.org/wiki/Endonuclease
- 1% – https://www.neb.com/tools-and-resources/selection-charts/properties-of-exonucleases-and-nonspecific-endonucleases
- 1% – https://en.m.wikipedia.org/wiki/Endonuclease
- <1% – https://www.pnas.org/content/104/25/10358
- <1% – https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2632897/
- <1% – https://www.ncbi.nlm.nih.gov/gene/54464
- <1% – https://www.ncbi.nlm.nih.gov/books/NBK22528/
- <1% – https://studiousguy.com/restriction-enzymes-types-examples/
- <1% – https://quizlet.com/84140332/fom-chapter-10-flash-cards/
