Anchored PCR is a modified PCR technique used to amplify and characterize unknown DNA sequences adjacent to a known DNA region by using one sequence-specific primer and one non-specific anchor primer.
In traditional PCR, amplification is performed using two specific primers that are complementary to a known gene sequence. These primers bind to the region of interest, and amplification proceeds accordingly. Characterizing an unknown DNA sequence adjacent to a known sequence is not possible using traditional methods. It also requires the target molecules to be separated from other molecules. Examples include characterizing cDNA, sequences around insertional sequences, and rearrangements of antigen receptor genes.
Anchored PCR modifies the traditional PCR method to solve this problem. In anchored PCR, two primers are used: one specific primer and one non-specific, called the anchor primer. This method is complex. It requires consideration of several factors, such as whether the target is single or double-stranded, the amount and heterogeneity of the target, and the quality and length of the known sequence.
Objectives of Anchored PCR
The objectives of this method are:
- To characterize an unknown sequence of DNA contiguous to a known sequence.
- To amplify and clone sequences with unknown 5′ or 3′ extremities.
- To utilize two primers, known and unknown primers, to flank the ends of the cDNA
- To amplify full-length mRNA with only a small amount of sequence information available.
- To separate the required DNA from a complex mixture of unwanted substances.
Requirements of Anchored PCR
- Template RNA
- oligo(dT)20 primer
- Sequence-specific primers
- Reverse transcriptase
- Reverse transcriptase buffer
- dNTP’S
- Taq DNA polymerase
- TE Buffer
- Mg2+
- Thermocycler
Principle of Anchored PCR
In anchored PCR, amplification of the sample can be performed with limited knowledge of its sequence. Prior knowledge of the sequences for primer designing is eliminated with this method. Here, a single specific primer is designed to amplify the sample, while the other primer, known as the anchor primer, is non-specific.
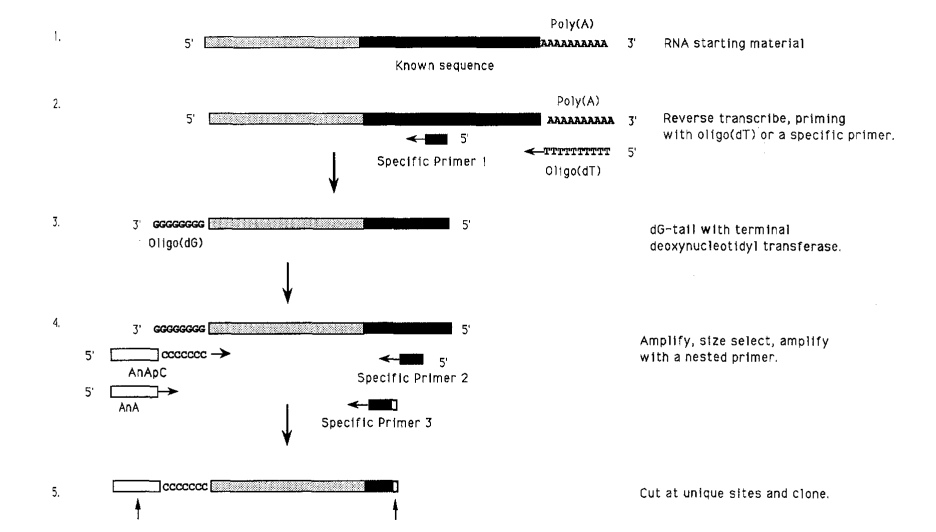
Here, amplification occurs in two rounds: the first uses a single specific primer, and the second uses a second sequence-specific primer. In terms of cDNA, amplification can be done from either the 5’ or 3’-end. For the 3’ end, an oligo(dT) primer, which is complementary to the pol(A) tail of the mature mRNA, is used, whereas this primer is used for the enzymatically synthesized homopolymer tail added to the cDNA (after the first strand synthesis) to the 5’ end upstream of the known sequence. The resulting product is then further sequenced for characterization.
Steps / Protocol of Anchored PCR
The overall steps involved in this method include:
- Primer designing
- RNA Preparation
- First-Strand Synthesis
- Tailing
- Amplification
- Reamplification with a Nested Primer
- Analysis of the Products
Primer designing
Two primers should be designed in this method: the specific and the Anchor primer.
- Anchor primer
When designing the anchor primer, one should ensure that:
-It has high annealing temperatures. At low annealing temperatures, there is a chance of the anchor primer to time incorrectly.
-It should be rich in GC content, which provides higher annealing temperatures.
-It should have a restriction site.
-When adding restriction sites and other functional sequences, it should be ensured that it has no homology (similarity) with themselves, repetitive DNA, or the specific primers.
- Sequence-specific primer
Two sequence-specific primers are used, one for the original amplification and the second for the re-amplification. The second primer used for the re-amplification can lie immediately adjacent to the first specific primer or can partially overlap at the 3′ end. To design the sequence-specific primer, the guidelines are:
-The length is dependent on the GC content, annealing temperature, and complexity of the DNA (i.e., the lower the DNA complexity of the original mixture, the less stringent the annealing conditions need to be, and thus one can use shorter primers). AT-rich sequences are not preferred because of their higher annealing temperatures.
-Primers that share 3’ homology, even a few bases, with other sequences should be avoided. Additionally, primers with internal secondary structures or direct repeats should also be avoided.
-The melting temperatures of the primers should be close so that the annealing temperatures can be similar to their melting temperature (Tm).
-Restriction sites can be incorporated at the 5’ end of the primer if the goal is to clone the products.
-Degeneracies at some bases of the primer ensure that several related sequences can be amplified with a single primer. However, such primer sets can reduce the concentration of individual primers, which can, in turn, affect the annealing time.
RNA Preparation
The quality of the RNA used determines the sensitivity of the reaction. RNA is obtained from cells using various methods, such as the acid-phenol extraction method, guanidium isothiocyanate-cesium trifluoroacetate (CsTFA) method, etc. While extracting, RNase-free reagents and water should be used. The poly(A)+ RNA is incubated for 2 min at 65°C, which melts secondary structures, removing hairpins and loops that may interfere with cDNA synthesis.
First-Strand Synthesis
The primer used for this can be either the specific primer or the anchor primer (oligo(dT)); the former yields higher yields by reducing background. The cDNA is synthesized using the reverse transcriptase. After synthesis, the primer should be removed by gel filtration because it can be tailed in the next step and interfere with amplification.
Tailing
Following the cDNA synthesis, homopolymer tails are added to the cDNA utilizing the terminal deoxynucleotidyl transferase. Tailing is done for amplification upstream (5’) and is enzymatically attached to the 3’ end. dGTP is added as the tail, since it is self-limiting, meaning the tailing reaction stops at approximately 20 nucleotides in length. When it comes to other nucleotides, the tailing reaction can be much longer, providing varying potential priming sites for the non-specific primer and resulting in products of variable lengths. The template RNA is then destroyed using NaOH or heat. After tailing, the primer is removed.
Amplification
This involves three steps:
- Denaturation: 94°C for 3 min in the first cycle.
- Annealing: 45°C for 1 min. As mentioned before, the annealing temperature for the primers used should be high to prevent non-specific priming. To further prevent any non-specific priming and amplification, the reaction mixture is kept in ice-cold conditions until the amplification procedure, or the polymerase is added only after all components are added. This prevents the synthesis of any undesirable products.
- Extension: It is done at 72°C for 1 min.
This is done for the first cycle. For the subsequent 5 cycles, 94°C for 1 min, 45°C for 1 min, and 72°C for 1 min were followed. This is then followed by 94°C for 1 min, 60°C for 1 min, and 72°C for 1 min for a further 25 cycles. The first few cycles are performed at a temperature that ensures priming, but subsequent rounds can be performed at higher temperatures. The templates for the next round of amplification are selected by gel electrophoresis to increase reaction specificity.
Reamplification with a Nested Primer
A nested specific primer is utilized to increase the specificity of the reaction. This primer can be any sequence that is 3′ of the primer that was used in a previous round of PCR and thus won’t be present in any sequences that were illegitimately primed during the first round of amplification. For anchored PCR, the DNA is size-selected: it is extracted using chloroform, ethanol-precipitated, resuspended in TE buffer, and then run on an agarose gel. The bands are selected, the gel is melted, and the gel is used for the next round of amplification. Care should be taken to select samples and amplify them in the absence of ethidium bromide, especially for biologically important products.
Analysis of the Products
The amplified product can be analyzed by sequencing directly, or it can be cloned into a bacterial vector first. The former allows for hiding the RT (Reverse transriptase) and Taq polymerase errors; however, it also hides the heterogeneity of the product.
Types of Anchored PCR
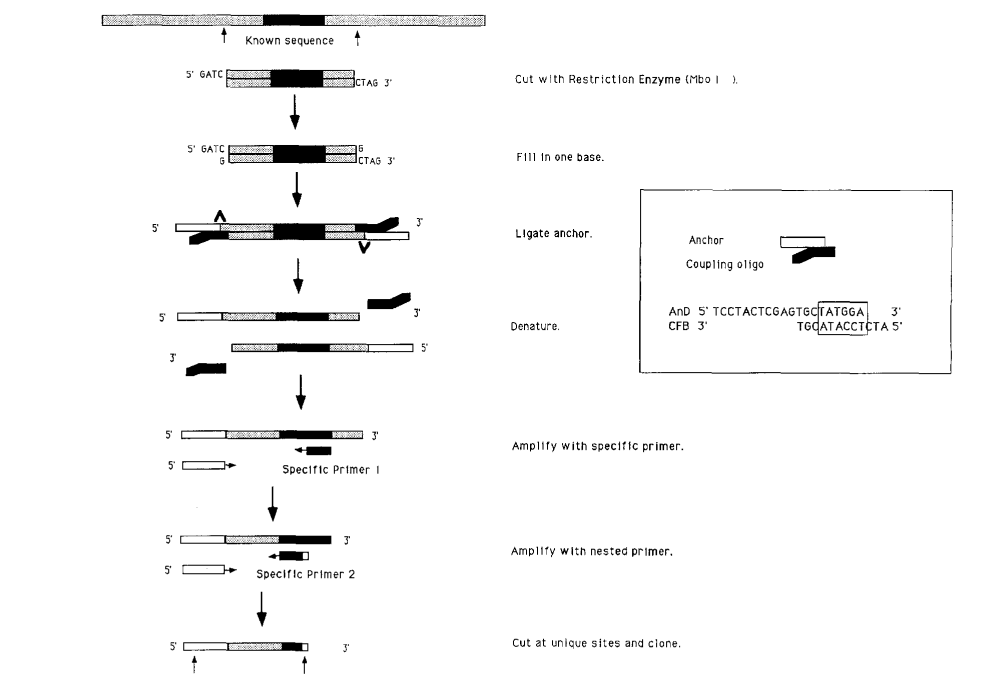
Ligation-mediated anchored PCR
One variation of anchored PCR, ligation-mediated anchored PCR, is used when the objective is to characterize a sequence adjacent to a known segment of double-stranded genomic DNA. The anchor primers are ligated to restriction endonuclease-generated fragments.
Anchored Multiplex PCR
This modified method is mainly used for studying the gene rearrangement testing of the ALK (encoding anaplastic lymphoma receptor tyrosine kinase), RET (encoding ret proto-oncogene), and ROS1 (encoding ROS proto-oncogene 1) genes for targeted lung cancer therapy.
Examples of Anchored PCR
DriverMap™ AIR DNA (Cellecta, Inc.)
- Anchor PCR is used for amplifying target CDR3 DNA fragments.
- The master mix contains PCR Buffer (5X), dNTP Mix, Anchor Primer Mix (10X), Water, and DNA Polymerase.
- The recommended temperature and time for the amplification are given below.
| Temperature | Time | Cycles |
| 98°C | 1 min | 1 |
| 98°C | 20 sec | 22-25 |
| 65°C | 10 sec | |
| 72°C | 30 sec | |
| 72°C | 30 sec | 1 |
| 4°C | ∞ | 1 |
ArcherTM Anchored Multiplex PCR (Integrated DNA technologies)
- Provides flexible assay content.
- Use multiplex PCR in addition to anchor primers.
Applications of Anchored PCR
Some of the application os this method include:
- It is used for detecting gene rearrangements in cancer.
- Utilized in cancer genomics for high-throughput identification of gene fusions, deletions, and breakpoints. It is widely used to detect mutations in cancers such as ROS1 and ALK fusions.
- It can be used to map unknown 5’ or 3’ ends of complementary DNA, which would otherwise have to be determined through cDNA libraries.
- Very useful in studying variable T-cell receptor (TCR) repertoires
- It has found its application in Next Generation Sequencing studies.
Advantages of Anchored PCR
- Amplification of unknown sequences with a limited amount of knowledge is possible.
- Anchored Multiplex PCR can be used to detect gene fusions and rearrangements.
- Highly sensitive method
- Amplification can be performed with low amounts of starting material.
- The re-amplification procedure prevents nonspecific amplification by using nested primers.
Limitations of Anchored PCR
The limitations of this method include:
- It is a highly complex process with multiple amplification steps.
- Chances of contamination are high due to the size-specific choosing using gel electrophoresis after the initial amplification process.
- Variable-sized products may result from long homopolymer tailing.
- A T-rich primer cannot be utilized.
- Primers utilized should have a high annealing temperature; otherwise, non-specific amplification might occur.
Conclusion
Anchored Polymerase Chain Reaction is a modification of the traditional PCR method utilized for the characterization of unknown sequences. This method is used to characterize sequences contiguous to known sequences. This can be done with two primers: a sequence-specific primer, which primes the known DNA sequence, and a non-specific primer, also known as the anchored primer.
This primer is used to amplify cDNA sequences. This is a highly complex yet sensitive method useful for characterization of unknown sequences, Next Generation Sequencing studies, cancer research, etc.
References
- https://manuals.cellecta.com/drivermap-air-tcr-bcr-profiling-kit-human-dna-v2/v1/en/topic/cdna-synthesis
- https://sg.idtdna.com/page/products/next-generation-sequencing/anchored-multiplex-pcr
- https://bitesizebio.com/25410/anchored-multiplex-pcr-for-next-generation-sequencing/
- Okano, K., Uematsu, C., Matsunaga, H., & Kambara, H. (1998). Characteristics of selective polymerase chain reaction (PCR) using two‐base anchored primers and improvement of its specificity. Electrophoresis, 19(18), 3071-3078.
- Ferradini, L., Roman-Roman, S., Azogui, O., Geneve, C., Viel, S., & Hercend, T. (1993). The use of anchored polymerase chain reaction for the study of large numbers of human T-cell receptor transcripts. Molecular immunology, 30(13), 1143-1150.
- Zheng, Z., Liebers, M., Zhelyazkova, B., Cao, Y., Panditi, D., Lynch, K. D., … & Le, L. P. (2014). Anchored multiplex PCR for targeted next-generation sequencing. Nature medicine, 20(12), 1479-1484.
- Loh, E. (1991). Anchored PCR: amplification with single-sided specificity. Methods, 2(1), 11-19.
- Dorit, R. L., & Ohara, O. (1992). cDNA amplification using one‐sided (anchored) PCR. Current protocols in molecular biology, 17(1), 15-6.
