Transcription factors (TFs) are specialized proteins that bind to specific DNA sequences, such as promoters or enhancers, to regulate gene expression.
- It acts as a molecular switch to control the location, timing, and amount of gene expression.
- For the bacteria to adapt to a changing environment, TFs respond to external signals by switching on or off the activity of a specific promoter.
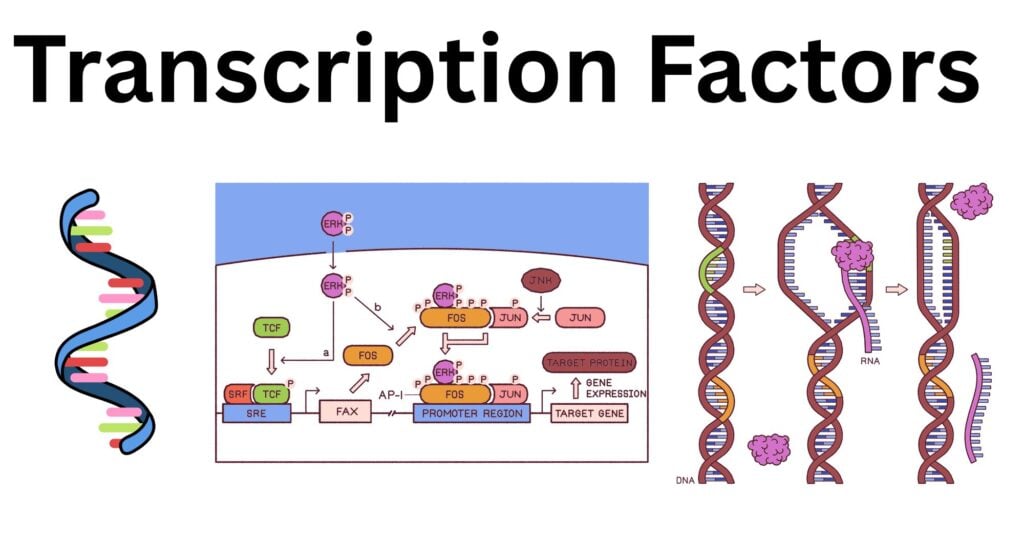
- In eukaryotes, TFs control developmental patterning, immune responses, and cell type specification.
- Depending on the environmental conditions or the cell type, a single TF may regulate different sets of genes.
- TF consists of an independently folding DNA-binding domain (DBD) that recognizes specific sequences or motifs, and one or more regulatory domains that modulate the activity of TF through ligand binding, protein-protein interactions, or covalent modification.
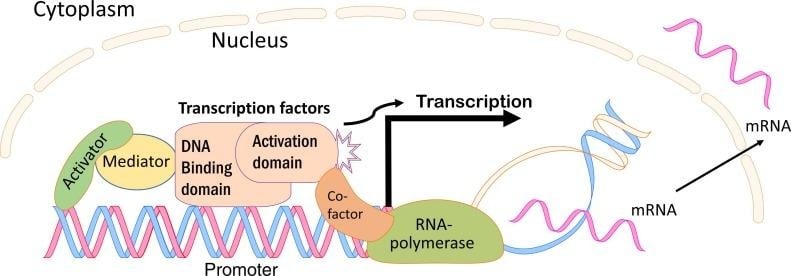
Structural Classification of Transcription Factors
Transcription factors are structurally classified based on the sequence homology and structural motifs of their DNA-binding Domains (DBDs).
Eukaryotic Structural Classification: –
- Approximately 93% of human TFs bind DNA in either monomer or homomultimer forms.
- Only about 3% of TFs contain more than one distinct type of DBD, such as the POU: Homeodomain configuration.
- TFs in eukaryotes (such as humans) are arranged into a six-level, detailed hierarchical system.
- The four abstract levels of TFs are superclass, class, family, and subfamily, which show the general topology and specific structural similarities of the DBD.
- The two physical levels of TFs are genus and species. The genus denotes the group of proteins encoded by the same gene, showing their common origin. Whereas the species denotes the individual protein isoforms or splice variants.
The ten major superclasses of eukaryotic TFs are: –
- Basic Domain: This includes families such as bZIP and bHLH.
- Zinc-coordinating Domain: This includes Nuclear Hormone Receptors and C2H2-zinc fingers.
- Helix-turn-helix (HTH) Domain: This includes the Homeodomain family and Forkhead.
- All-α-helical DBDs.
- α-helices exposed by β-structures.
- Immunoglobulin fold: This includes families like p53 and Rel.
- β-Hairpin exposed by an α/β-scaffold.
- β-Sheet binding to DNA.
- β-Barrel DNA-binding domain.
- Undefined DNA-binding domain: Proteins for which the structure is not yet solved.
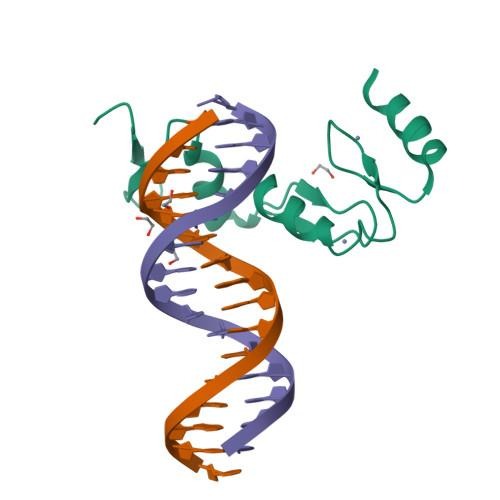
Prokaryotic Structural Classification: –
- Most bacterial TFs function as dimers.
- Members of the AraC Family carry two independent HTH motifs within a single domain to achieve higher specificity.
- In prokaryotes, the TFs are classified according to their domain architecture and signal response.
- The four primary functional categories of prokaryotic TFs are: –
- One-Component Systems (OCSs): Proteins combining a sensory domain and a DBD directly in a single polypeptide chain.
- Response Regulators (RRs): TFs that are regulated by separate histidine kinases via an N-terminal phospho-acceptor (receiver) domain. They are part of two-component systems.
- Sigma Factors (SFs): Initiation factors that direct RNA polymerase to specific promoters. They are divided into RpoD (housekeeping), RpoN (Sigma-54), and ECF (extra-cytoplasmic function).
- Transcriptional Regulators (TRs): A broad category for TFs that do not fit into OCS, RR, or SF architectures.
Key structural motifs:
- Helix-Turn-Helix (HTH): The most prevalent DBD motif in prokaryotes.
- Tri-helical HTH: The simplest version, seen in the Fis family.
- Tetra-helical HTH: Features a fourth C-terminal helix that includes AraC, LuxR, LacI, and TetR families.
- Winged HTH (wHTH): Features a C-terminal β-strand hairpin that contacts the minor groove. This is most commonly used by families like OmpR, MarR, and GntR.
- Ribbon-helix-helix (RHH): Also known as the MetJ/Arc domain that uses an N-terminal β-strand for DNA contact.
Mechanism of Action of the Transcription Factors
Prokaryotic Mechanism of Action
Repression Mechanisms:
- Steric Hindrance: Most repressors bind to DNA sites that overlap the promoter, thereby physically blocking RNA polymerase (RNAP) access.
- DNA Looping: Repressor dimers bound at distal sites interact with each other, looping the DNA to prevent RNAP engagement.
- RNAP Jamming: In some cases, RNAP is allowed to bind the promoter but is blocked by the repressor, preventing it from initiating transcription.
Activation Mechanisms:
- Recruitment: Activators bind upstream of the promoter and use an “Activating Region” to make direct protein-protein contact with RNAP, facilitating its recruitment to the DNA.
- Conformational Change: Some activators change the physical structure or spacing of promoter DNA so that essential elements are better positioned for RNAP binding.
- Anti-repression: Activators can function by displacing Nucleoid-Associated Proteins (NAPs) that are normally silencing or locking a promoter.
- Signalling Control: Activity is often toggled by environmental signals through one-component systems (direct ligand binding/sensing) or two-component systems (phosphorylation by a separate histidine kinase).
- Basal Apparatus: Specific sigma factors associate with the RNAP catalytic core to direct it to specific sets of promoters.
Eukaryotic Mechanism of Action
- Indirect Recruitment: Most eukaryotic TFs do not contact RNAP directly. Instead, they recruit large multi-subunit cofactor complexes.
- Coactivators: Proteins like the Mediator complex bridge the TF to RNA polymerase II to support the preinitiation complex.
- Chromatin Modifiers: TFs recruit enzymes to transition chromatin from a repressive to an accessible state by modifying histones via methylation or acetylation, or remodelling nucleosomes.
- Collaborative Logic: Due to low individual specificity, eukaryotic TFs must work in a group, utilizing cooperative binding or synergy to occupy target sites effectively.
- Nucleosome Management: TFs must compete with or displace nucleosomes to access DNA. Specialized pioneer factors can bind to closed chromatin and initiate nucleosome displacement.
- Context Dependency: The same TF may act as an activator or a repressor depending on the cell type, local sequence context, and available cofactors.
- Steric Displacement: Similar to bacteria, some eukaryotic TFs act through steric mechanisms, such as blocking other proteins from binding to the same site.
Promoters and Enhancers
Promoters
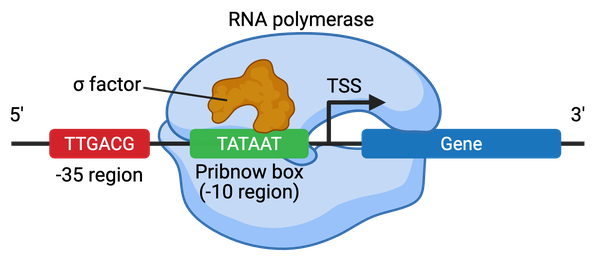
- Promoters are the region of the DNA, which is located upstream of, or at the transcription start site (TSS) of a gene, where general transcription factors and RNA polymerase bind to initiate transcription.
- In bacteria, promoters contain two key hexameric motifs centered at the -10 (TATAAT) and -35 (TTGACA) positions relative to the TSS.
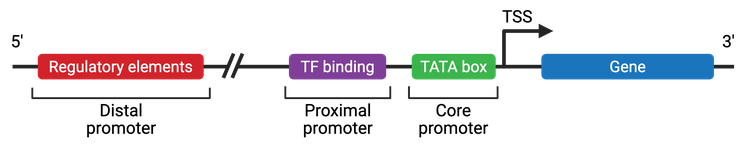
- In humans and plants, promoters include a variety of motifs such as the TATA box (around -30 bp), INR box, BRE, and GC-box.
- Across all the taxa, promoters share distinct structural traits that help RNAP find them.
- Promoters often have A-T rich sequences, which makes it easier for the two DNA strands to separate and form an open complex for transcription.
- They often contain sequence-dependent bent DNA elements that assist in wrapping the DNA around the RNA polymerase.
- In eukaryotes, upstream promoter regions are usually more rigid than the downstream coding sequences, which may help prevent accidental nucleosome formation.
A
B
Enhancers
- Enhancers are regulatory DNA sequences that bind transcription factors (TFs) to boost the rate of transcription of a gene.
- They interpret cellular and environmental signals to decide when and how much a gene should be expressed.
- Regardless of their orientation, they activate a promoter over long distances, often tens or hundreds of kilobases away.
- Multiple TFs are bound in combinations, known as the enhanceosome, to regulate gene expression precisely.
- Certain pioneer transcription factors can recognize enhancer sites even in compacted chromatin to initiate the gene activation process during development.
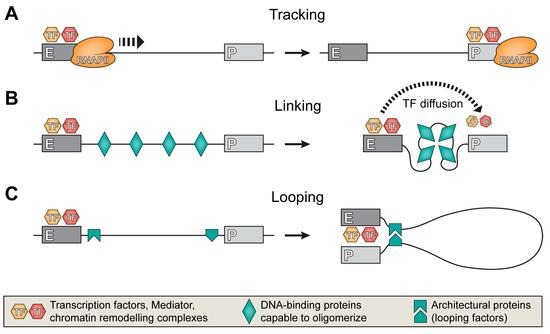
- Tracking, linking, and looping are the mechanisms for establishing enhancer-promoter communication.
- In the Tracking model, transcription factors and machinery, such as RNA polymerase II, are physically carried along the chromatin from the enhancer to the promoter via low-level intergenic transcription.
- In the Linking model, specialized proteins bind to the DNA segment between the enhancer and promoter. These proteins then condense or oligomerize, compressing the DNA to pull the two elements into physical proximity.
- In the Looping model, architectural proteins bind directly to both the enhancer and the promoter and interact with one another, causing the intervening DNA to loop out so the two elements can meet.
Key differences in the architecture of promoters and enhancers between Prokaryotic and Eukaryotic systems are as follows: –
Prokaryotic system
- Regulatory DNA is usually not strictly divided into promoter and enhancer regions. The entire block is often referred to as a promoter.
- Upstream Activating Sequences (UAS) are almost always located proximal to the core promoter, typically within 300 base pairs.
- Bacterial activators usually make direct physical contact with the RNA polymerase to recruit it to the DNA.
- Most bacterial promoters are regulated by only one or a few factors to adapt to immediate environmental shifts.
Eukaryotic system
- There is a clear distinction between the core promoter and distal enhancers.
- In humans, enhancers can be nearly 1,000,000 base pairs from their target gene.
- Eukaryotic activators recruit large coactivator complexes such as mediator and chromatin modifiers to make indirect contact with RNAP.
- The regulation relies on the 3D architecture of the genome by utilizing mechanisms like Topologically Associating Domains (TADs) and loop extrusion to manage interactions.
General vs. Specific Transcription Factors
Transcription factors (TFs) are broadly categorized into general and specific factors based on whether they are required for the transcription of all protein-coding genes or only for specific genes in response to stimuli.
General Transcription Factors
- These are the minimal set of proteins that are essential to initiate transcription.
- These are generally constitutive in nature.
- They target promoters universally.
Prokaryotic General TFs:
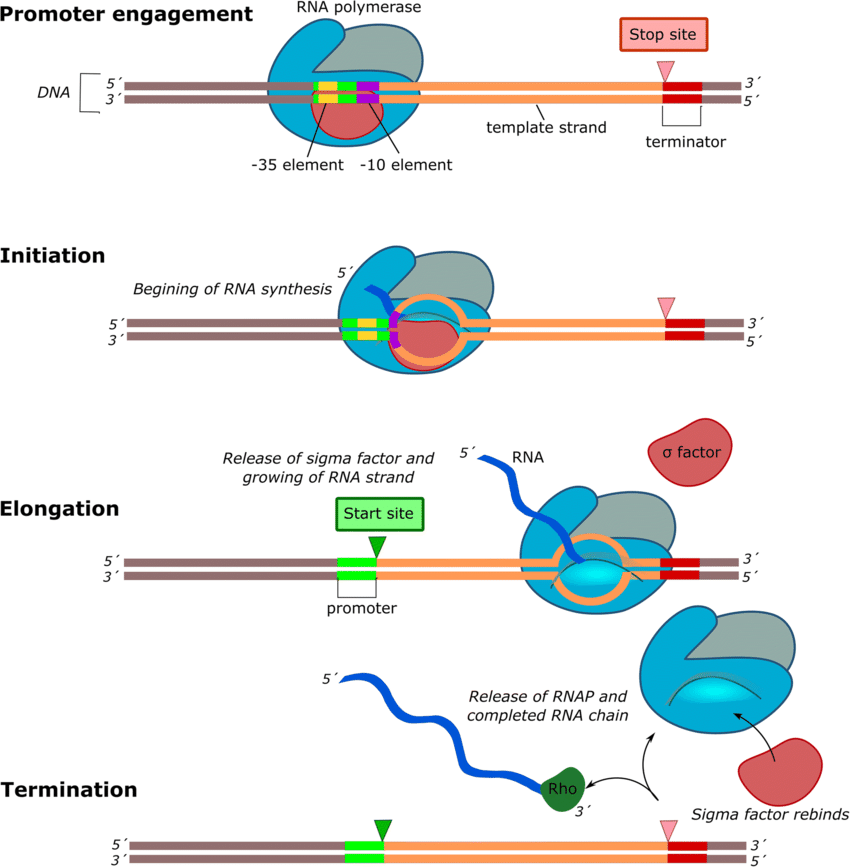
- The σ factor is the primary general TF in bacteria.
- To initiate transcription, the σ factor recognizes the promoter sequence and recruits RNA polymerase to the DNA.
- Most bacteria, such as E. coli, have one primary σ factor, i.e. σ70, for housekeeping genes.
- Several alternative σ factors direct the polymerase to specific sets of genes during development or stress.
Eukaryotic General TFs:
- They have a more complex set of basal TFs like TBP and TFIIB.
- To form the preinitiation complex, these TFs are needed, which help in positioning RNA polymerase at the Transcription Start Site (TSS).
- The eukaryotic TFIIB and the bacterial σ factor are considered to be evolutionary orthologs.
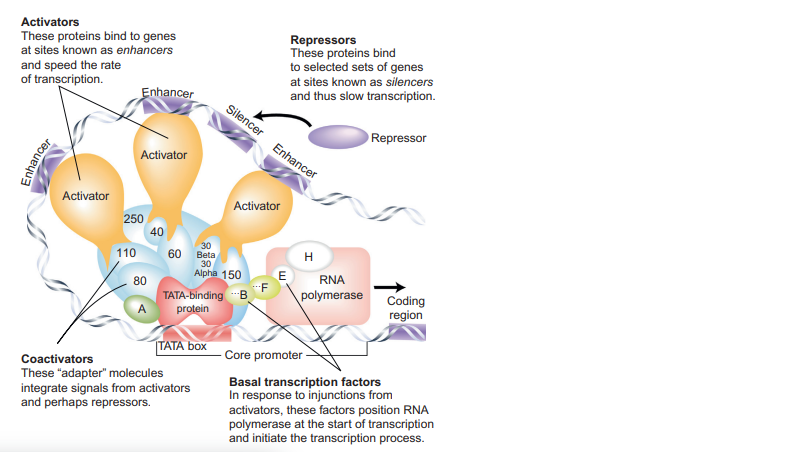
Specific Transcription Factors
- These TFs either activate or repress the individual gene or the set of genes by sensing environmental or internal signals.
- Activators positively regulate transcription by recruiting RNA polymerase or cofactors to a specific site.
- Repressors negatively regulate transcription by physically blocking the DNA-binding sites of other proteins.
- Specific TFs bind to regulatory sequences, such as enhancers in eukaryotes and operators in bacteria, to enable gene-specific transcription regulation.
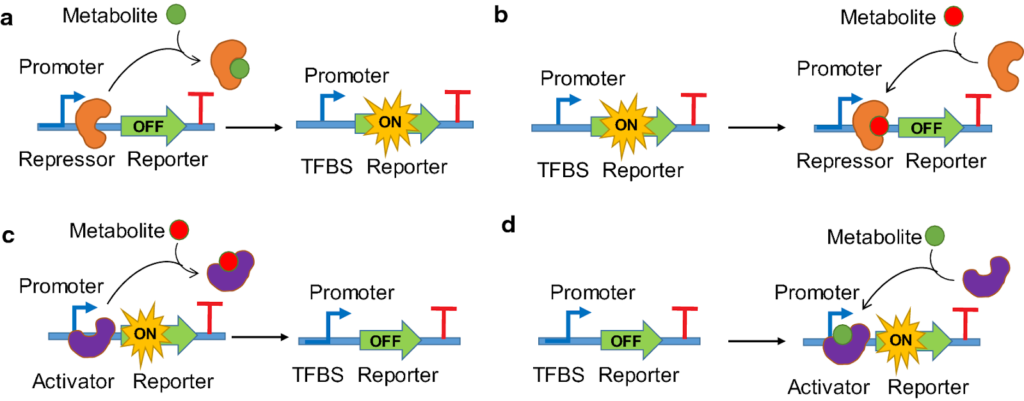
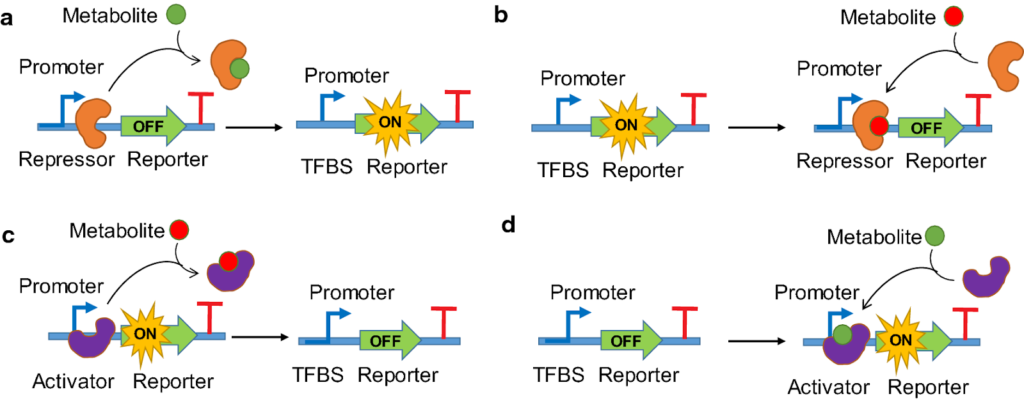
Prokaryotic Specific TFs:
- These TFs detect environmental changes directly by binding a metabolite or being phosphorylated by a sensor. Eg: – AraC activator, Lac repressor, etc.
- To bind DNA, most use a Helix-Turn-Helix (HTH) structure.
A
B
Eukaryotic Specific TFs:
- These TFs are highly diverse master regulators that control complex processes like developmental patterning or immune responses. Eg: – p53, Sox family, etc.
- They use various structures like Homeodomains and Zinc fingers.
Regulation of Activity
For organisms to maintain cellular homeostasis and adjust to the changing environment, TFs must be precisely regulated.
Prokaryotic Regulation of TF Activity
In bacteria, TFs function by responding directly to environmental physicochemical conditions.
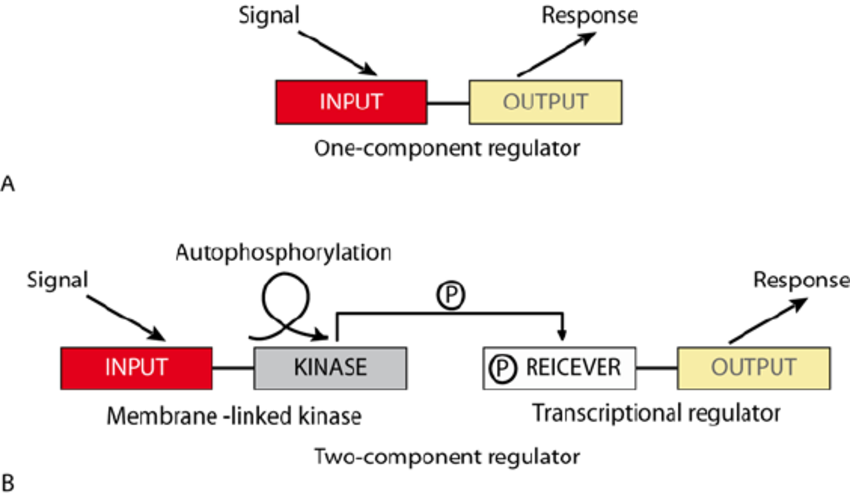
One-component systems (OCSs):
- These systems consist of a single TF containing both a sensory domain and a DNA-binding domain.
- The direct binding of a small effector molecule or a metabolite changes the conformation of TF that regulates its own DNA binding activity.
Two-Component Systems (TCSs):
- These systems consist of a sensor histidine kinase in the cell membrane and a separate cytoplasmic response regulator TF.
- When the sensor detects the external stimulus, the cytoplasmic catalytic domain autophosphorylates a conserved histidine residue.
- This phosphate group is transferred by histidine kinase to a conserved aspartate residue on the receiver domain of the response regulator, leading to conformational change of the TF to allow DNA binding and regulation of transcription of the target gene.
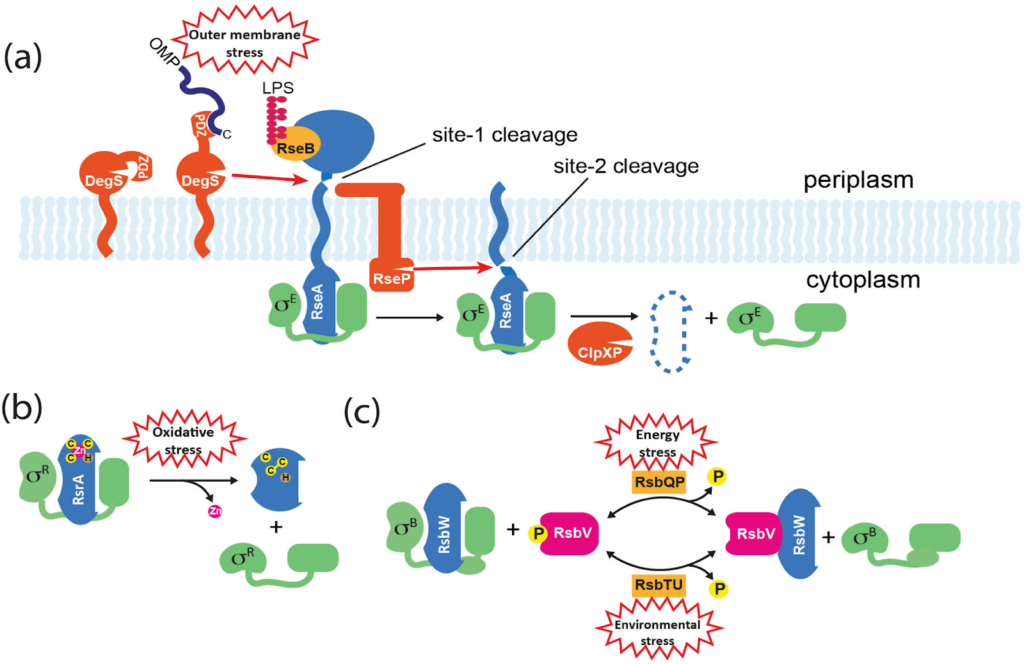
Anti-Sigma Factors:
- An environmental signal triggers the release of the sigma factors (σ), which are often held in an inactive state by anti-sigma factor proteins, allowing them to initiate a specific transcription by recruiting RNA polymerase to promoters.
Covalent Modifications:
- The LexA repressor is regulated by autoproteolysis (self-cleavage), which is triggered by contact with the RecA protein during the DNA damage response.
Intracellular Levels:
- The regulatory effect of the constitutively active TF is managed simply by altering its concentration, and controlling its localization, accessibility to DNA, or the availability of co-activators or co-repressors.
Eukaryotic Regulation of TF Activity
In eukaryotes, the TF regulation involves complex signaling cascades and multi-subunit complexes.
Ligand Binding:
- For Nuclear Hormone Receptors, this is a primary regulatory mechanism. When a specific ligand binds to the Ligand-Binding Domain, it triggers a conformational change that leads to the TF’s interaction with coactivators or corepressors.
Covalent Modifications:
- Signaling kinases like MAPK or p-TEFb perform phosphorylation to switch the DNA-binding affinity or the ability to recruit the transcription machinery.
- To facilitate the muscle differentiation process, coactivators acetylate the master regulator MyoD.
Dimerization and Combinatorial Logic:
- Many eukaryotic TFs function as homodimers or heterodimers. For instance, c-Myc pairing with Max can increase DNA-binding affinity and transactivation power.
Localization and Sequestration:
- TFs can be regulated by controlling their presence in the nucleus. These TFs may be held in the cytoplasm until a signal triggers their entry into the nucleus.
Recruitment of Co-factors:
- Eukaryotic TFs rarely contact RNA polymerase directly. Instead, their activity depends on recruiting large multi-protein coactivator or corepressor complexes (e.g., Mediator, SWI/SNF, or HATs) that modify chromatin or bridge the TF to the polymerase.
Co-activators and Co-repressors
Co-activators and co-repressors, collectively termed as co-regulators, are molecules or large protein complexes that modulate the rate of transcription by associating with sequence-specific TFs to interpret cellular signals and translate them into gene expression outputs.
Eukaryotic Co-regulators
- TFs rely on a diverse network of co-regulators to manage chromatin accessibility and bridge interactions with the polymerase.
- The Mediator Complex is a huge multi-subunit complex that acts as a universal bridge, connecting various TFs to RNA polymerase II to facilitate the formation of the preinitiation complex. It integrates signals from multiple enhancers and transmits them to the core promoter.
- Many co-activators have enzymatic activity, such as Histone Acetyltransferases (HATs) like p300/CBP, which acetylate histones to open compacted chromatin and make DNA accessible.
- Co-repressors often recruit Histone Deacetylases (HDACs) or methyltransferases to deposit repressive marks, such as H3K9me3, which silence gene expression.
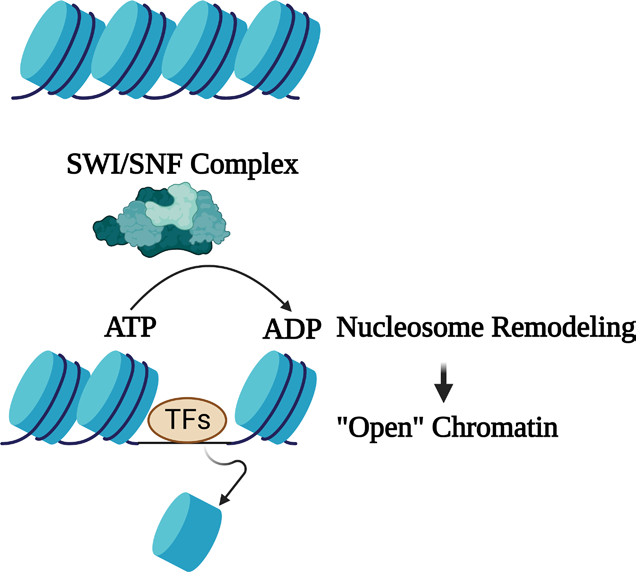
- Complexes like SWI/SNF use the energy of ATP hydrolysis to displace or reposition nucleosomes, clearing a path for the transcription machinery. To open closed heterochromatin, pioneer transcription factors recruit these remodelers.
- Many co-regulators interact with TFs through specific interfaces, such as the LxxLL motif, which was originally identified in nuclear hormone receptors but is now recognized across many unrelated TF families.
- Depending on the available cofactors and the local DNA sequence, a single protein like MAX can function as an activator when paired with MYC, or as a repressor when associated with MNT.
Prokaryotic Co-regulators
- Prokaryotes also use accessory proteins and molecules that function as co-regulators.
- Certain bacterial activators require a second protein to function effectively. For instance, the protein Sxy acts as a co-activator for the CRP (cAMP Receptor Protein) to help it bind to non-canonical DNA sites and activate specific genes.
- In complex promoters like araBAD, the CAP protein acts as a co-activator for AraC, facilitating the breaking of a DNA repression loop and increasing the rate of RNA polymerase open complex formation.
- Specific proteins can enhance a repressor’s affinity for its target. A classic example is the bacteriophage-encoded gp7 protein, which interacts with the host LexA repressor to double-lock phage promoters and maintain the lysogenic state.
- Many bacterial regulators only become active upon binding a small-molecule cofactor. For example, the PurR regulator acts as a repressor of purine biosynthesis only after binding a purine co-repressor.
- Anti-repressors, like CarS, can be produced in response to signals like light to displace repressors from their operators.
Pioneer Transcription Factors
Pioneer Transcription Factors (pTFs) are a specialized class of proteins that can bind DNA sequences that are buried within compacted heterochromatin.
Eukaryotic Pioneer Transcription Factors
- In eukaryotes, DNA is packaged into nucleosomes, which typically restrict access to transcription factors.
To bypass this restriction, pTFs have the following unique properties: –
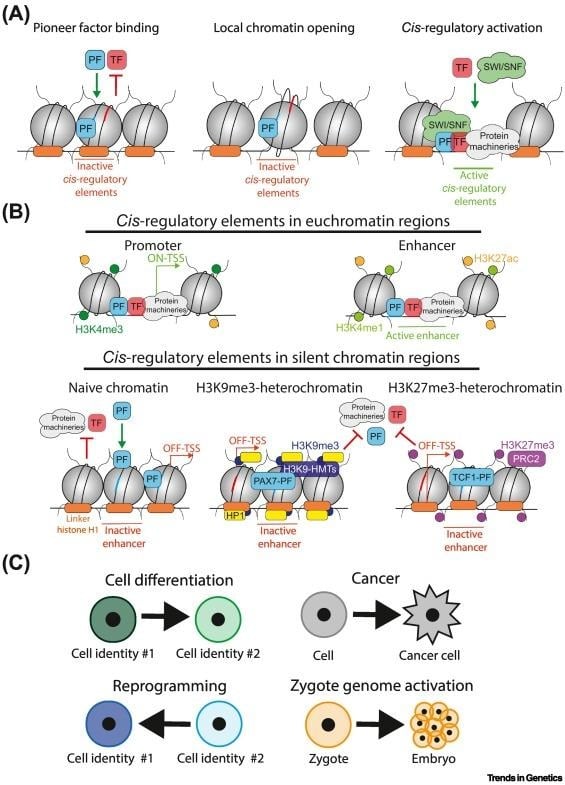
- Binding Closed Chromatin: pTFs can recognize and bind their target motifs within closed or unmarked heterochromatin, where the DNA is wrapped around nucleosomes.
- Initiating Remodeling: Their binding triggers local chromatin opening and remodeling, making the region accessible to non-pioneer factors.
- Stepwise Action: Pioneer activity involves an initial rapid but weak binding to heterochromatin, followed by a stabilization phase that facilitates nucleosome depletion and the deposition of active histone marks like H3K4me1 and H3K27ac.
- Epigenetic Memory: They establish stable changes, such as the loss of CpG methylation, which provides a form of memory to maintain the open state of enhancers for future gene expression.
Structural Mechanisms:
- Nucleosome Interaction: pTFs like FoxA possess a domain that mimics linker histone H1, allowing them to displace H1 and promote decompaction.
- DNA Sliding: Binding of pTFs like Oct4 can induce structural changes that cause the DNA to slide on the nucleosome, exposing internal binding sites for other factors like Sox2.
Biological Roles:
- pTFs are essential for cell fate decisions and cellular reprogramming. For instance, reverting fibroblasts to pluripotent stem cells.
- Some pTFs like GATA1, Sox2, Oct4, remain bound to chromosomes during mitosis, ensuring the original gene expression program is re-established in daughter cells.
Prokaryotic Functional Equivalents
- In prokaryotes, DNA is essentially naked compared to eukaryotes, but it is still structured and silenced by Nucleoid-Associated Proteins (NAPs) like H-NS, Fis, and IHF.
- In bacteria, anti-repression is the functional analog to pioneering.
- Certain activators, like NarL, function primarily by pushing NAPs out of the way to allow other factors, such as FNR, to recruit RNA polymerase and initiate transcription.
- Bacteria utilize a class of proteins called bEBPs that bind to enhancer-like distal sites. These proteins utilize ATP hydrolysis to induce large activation loops, which physically bring the remote activator into contact with the RNA polymerase-σ54 complex. This mechanism is functionally the closest prokaryotic equivalent to eukaryotic enhancer-dependent regulation.
- Bacteriophage factors utilize a mechanism known as sigma appropriation, where they remodel the σ70 subunit of RNA polymerase to redirect the entire transcription apparatus to a specific set of promoters.
Methods of Analysis of Transcription Factors
Transcription factors (TFs) are analyzed through computational identification, experimental mapping of DNA-binding sites, and structural characterization.
Identification and Classification:
- By searching for known DNA-binding domains using Hidden Markov Models (HMMs) against databases like Interpro or Pfam, putative TFs are identified.
- Prokaryotic pipelines like P2TF screen both predicted proteomes and reconstituted ORFeomes to recover mispredicted TFs.
In Vitro Specificity:
- Intrinsic DNA-binding preferences are defined using high-throughput assays like HTSELEX, Protein Binding Microarray, or Spec-Seq to generate Position Weight Matrices.
- A specialized method for prokaryotes, called Genomic SELEX, utilizes actual genome fragments to identify target sequences.
Genome-Wide Mapping:
- For identifying TF occupied regions in vivo, ChIP-seq is used as the gold standard.
- To pinpoint sites at single-nucleotide resolution, ChIP-exo uses exonuclease digestion.
Chromatin and 3D Analysis:
- For identifying pioneer factors, MNase-seq, DNase-seq, and ATAC-seq map nucleosome positions and open chromatin.
- Hi-C or micro-C monitors 3D interactions like DNA looping.
Functional Characterization:
- Cryo-Electron Microscopy (Cryo-EM) provides atomic detail of TFs bound to DNA or nucleosomes, while RNA-seq profiles gene expression in TF-deletion mutants.
- In bacteria, phenotype microarrays identify environmental signals that activate specific TFs.
Clinical Significance of Transcription Factors
TFs control the expression of thousands of other genes. Their clinical significance is as follows:
Clinical Phenotype hubs:
- TFs represent about 8% of all human genes, yet they are involved in a disproportionately high number of diseases.
- Approximately 19.1% of human TFs are linked to at least one clinical phenotype.
Highly Sensitive Mutations:
- Mutations in TFs are very harmful.
- These proteins exhibit a reduced level of common genetic variation because the body cannot easily tolerate changes to its core regulatory machinery.
Developmental Blueprints:
- TFs act as selector genes that specify cell types and body patterns. In conditions like anterior pituitary hypoplasia, 12 of the 15 known involved genes are TFs.
- Specific mutations in the factor HOXD13 are directly linked to limb malformations.
Cancer Drivers:
- TFs like TP53 are among the most significant markers in oncology. Furthermore, chromosomal breaks can create onco-fusion proteins, such as the ERG-EWSR1 fusion, that gain dangerous new abilities to drive aggressive tumour growth.
Immune and Metabolic Disorders:
- Variants in the IKZF1 family are tied to multiple autoimmune disorders.
- Obesity risks have been associated with DNA changes at the FTO locus, which affect the binding of the TF ARID5B.
Regenerative Potential:
- TFs are the primary tools used in the lab for cellular reprogramming.
- Factors like OCT4 and SOX2 can revert adult cells into pluripotent stem cells.
Diagnostic Markers:
- Conventional TFs are widely used as biomarkers for disease identification and remain primary targets for high-precision drug development.
Conclusions
Transcription factors are specialized proteins that regulate gene expression, and are structurally classified based on the sequence homology and structural motifs of their DNA-binding Domains. The eukaryotes use ten main structural families, while bacteria rely on simple helix-turn-helix shapes.
In humans, these proteins use large bridging complexes and 3D looping to connect distant enhancers to the transcription start sites. Pioneer factors are capable of binding and opening tightly packed DNA. The transcription factors are analyzed through computational identification, experimental mapping of DNA-binding sites, and structural characterization.
Mutations in them are severe, often leading to aggressive cancers or serious developmental defects. Beyond disease, factors like OCT4 and SOX2 are the primary tools used in modern medicine to reprogram adult cells back into pluripotent stem cells.
References
- Barral, A., & Zaret, K. S. (2024). Pioneer factors: roles and their regulation in development. Trends in Genetics, 40(2), 134–148. https://doi.org/10.1016/j.tig.2023.10.007
- Browning, D. F., Butala, M., & Busby, S. J. W. (2019). Bacterial Transcription Factors: Regulation by Pick “N” Mix. In Journal of Molecular Biology (Vol. 431, Number 20, pp. 4067–4077). Academic Press. https://doi.org/10.1016/j.jmb.2019.04.011
- Bylino, O. V., Ibragimov, A. N., & Shidlovskii, Y. V. (2020). Evolution of regulated transcription. Cells, 9(7), 1–38. https://doi.org/10.3390/cells9071675
- Dian, C. (n.d.). Adaptive Responses by Transcriptional Regulators to small molecules in Prokaryotes Structural studies of two bacterial one-component signal transduction systems DntR and HpNikR.
- G. Abril, A., Rodriguez Rama, J. L., Sanchez, A., & Villa, T. (2020). Prokaryotic sigma factors and their transcriptional counterparts in Archaea and Eukarya. Applied Microbiology and Biotechnology, 104. https://doi.org/10.1007/s00253-020-10577-0
- Ishihama, A. (2010a). Prokaryotic genome regulation: Multifactor promoters, multitarget regulators and hierarchic networks. In FEMS Microbiology Reviews (Vol. 34, Number 5, pp. 628–645). https://doi.org/10.1111/j.1574-6976.2010.00227.x
- Iyer, L. M., & Aravind, L. (2012). Insights from the architecture of the bacterial transcription apparatus. Journal of Structural Biology, 179(3), 299–319. https://doi.org/10.1016/j.jsb.2011.12.013
- Kanhere, A., & Bansal, M. (2005). Structural properties of promoters: Similarities and differences between prokaryotes and eukaryotes. Nucleic Acids Research, 33(10), 3165–3175. https://doi.org/10.1093/nar/gki627
- Li, H. X., Chai, Y. H., Sun, X. H., He, X. X., & Xi, Y. M. (2024). Epigenetic regulatory differentiation of hematopoietic stem cells to the red lineage. In Reproductive and Developmental Medicine (Vol. 8, Number 3, pp. 169–177). Lippincott Williams and Wilkins. https://doi.org/10.1097/RD9.0000000000000092
- Li, Y., Azmi, A. S., & Mohammad, R. M. (2022). Deregulated transcription factors and poor clinical outcomes in cancer patients. Seminars in Cancer Biology, 86, 122–134. https://doi.org/10.1016/j.semcancer.2022.08.001
- Litwack, G. (2018). Transcription. In Human Biochemistry (pp. 337–358). Elsevier. https://doi.org/10.1016/B978-0-12-383864-3.00012-0
- Mayran, A., & Drouin, J. (2018). Pioneer transcription factors shape the epigenetic landscape. In Journal of Biological Chemistry (Vol. 293, Number 36, pp. 13795–13804). American Society for Biochemistry and Molecular Biology Inc. https://doi.org/10.1074/jbc.R117.001232
- Ortet, P., De Luca, G., Whitworth, D. E., & Barakat, M. (2012). P2TF: a comprehensive resource for analysis of prokaryotic transcription factors. http://www.biomedcentral.com/1471-2164/13/628
- Paget, M. S. (2015). Bacterial sigma factors and anti-sigma factors: Structure, function and distribution. In Biomolecules (Vol. 5, Number 3, pp. 1245–1265). MDPI AG. https://doi.org/10.3390/biom5031245
- Peng, Y., Song, W., Teif, V. B., Ovcharenko, I., Landsman, D., & Panchenko, A. R. (n.d.-a). Detection of new pioneer transcription factors as cell-type-specific nucleosome binders.
- Razin, S. V., Ulianov, S. V., & Iarovaia, O. V. (2023). Enhancer Function in the 3D Genome. In Genes (Vol. 14, Number 6). MDPI. https://doi.org/10.3390/genes14061277
- Sinha, K. K., Bilokapic, S., Du, Y., Malik, D., & Halic, M. (2023). Histone modifications regulate pioneer transcription factor cooperativity. Nature, 619(7969), 378–384. https://doi.org/10.1038/s41586-023-06112-6
- Sperling, S. (2007). Transcriptional regulation at a glance. In BMC Bioinformatics (Vol. 8, Number SUPPL. 6). https://doi.org/10.1186/1471-2105-8-S6-S2
- Wan, X., Marsafari, M., & Xu, P. (2019). Engineering metabolite-responsive transcriptional factors to sense small molecules in eukaryotes: Current state and perspectives. In Microbial Cell Factories (Vol. 18, Number 1). BioMed Central Ltd. https://doi.org/10.1186/s12934-019-1111-3
- Wingender, E., Schoeps, T., Haubrock, M., & Dönitz, J. (2015). TFClass: A classification of human transcription factors and their rodent orthologs. Nucleic Acids Research, 43(D1), D97–D102. https://doi.org/10.1093/nar/gku1064
