Southern Blot Definition
Southern blot is the process of transfer of DNA fragments that are separated by electrophoresis onto a membrane for immobilization and identification.
- Southern blotting has been adopted as a routine procedure for the analysis of DNA samples for different applications.
- The technique was discovered by Edwin Southern, and the technique was named after him. The technique later gave rise to other techniques like Western Blotting and Northern Blotting that are based on the same principle.
- The most basic form of the technique is used to determine the size of a DNA fragment from a complex mixture of genomic DNA.
- The technique is also relatively quantitative and can be used to determine the number of copies of a segment present in a genome.
- Southern Blotting can be modified based on the choices of the membrane, transfer buffer, and method. The most commonly used membrane is the nitrocellulose membrane, as it is robust and can be reprobed a number of times.
- Similarly, the original protocol of the southern blotting utilizes the use of radioactive probes; however, other labeling systems utilizing fluorescence and chemiluminescence.
- Southern blotting has been modified in a number of ways to better serve the application and has been made more complex and efficient.
Principle of Southern Blot
- The principle of southern blotting is similar to the blotting technique involving the transfer of biomolecules from a membrane to another for detection and identification.
- The DNA to be analyzed is digested with restriction enzymes and fractionated by size by the process of agarose gel electrophoresis.
- The DNA strands are denatured by alkaline treatment and are transferred to a nylon or nitrocellulose membrane by the blotting process.
- The strands on the membrane are immobilized on the surface by baking or UV irradiation. The DNA sequences on the membrane can be detected by the process of hybridization.
- Hybridization reactions are specific as the probes used bind to target fragments consisting of complementary sequences.
- The probes used are labeled with different components that can be visualized by different methods depending on the type of probes used.
Requirements
Equipment
- Water Bath
- Agarose gel
- Power supply
- UV radiation
- Hybridization Oven
- Hybridization Bottles
- Trays
- Film processor
- Pipettes
- Centrifuge Tubes
- Glass Plate
- Whatman 3 mm chromatography paper
- Nylon membrane/ Nitrocellulose membrane
- Syringe
- Cellulose acetate membrane
Materials
- Restriction Enzymes
- Restriction enzymes buffer
- Agarose
- TBE buffer
- DNA loading buffer
- Tris base
- Sodium chloride
- Sodium hydroxide
- Sodium citrate
- DNA labeling kit
- Nucleic acid detection kit
- Sodium dodecyl sulfate (SDS)
- Polyvinylpyrrolidone
- Bovine serum albumin
- Formamide
- Phenol
Solutions and Buffers
- Denaturation buffer: NaOH and NaCl in the ratio 1:6
- Neutralization buffer: Tris-HCl and NaCl in the ratio 5:3
- SSC: 175.3 g of NaCl and 88.2 g of Sodium citrate to 1L of distilled water.
- Detection Buffer: Tris-HCl and NaCl in the ratio 5:1.
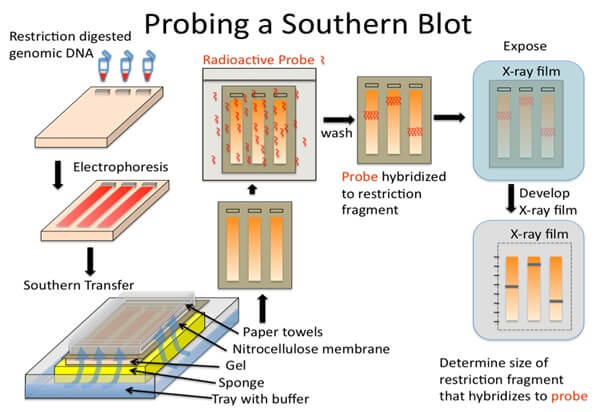
Procedure of Southern Blot
a. Restriction digestion of DNA
- About 10 µg of the extracted genomic DNA is digested with the appropriate restriction enzyme in a microcentrifuge tube.
- The tube is incubated overnight at 37°C. In some cases, the tubes are heated in a water bath at 65°C for 20 minutes after the incubation to denature the restriction enzymes.
- To the tubes, 10µl of the DNA sample buffer is added, and the mixture is poured on agarose gel for electrophoresis.
Image Source: Biology LibreTexts.
b. Electrophoresis
- The percentage and size of the gel are determined based on the size of the DNA fragments to be separated. The gel is then prepared accordingly.
- The electrophoresis buffer is prepared with ethidium bromide and poured into the tank in a way that is a few millimeters above the gel support.
- The gel cast is prepared along with a comb with teeth to form wells that can hold the sample volume. Once the comb is in place, the gel is slowly poured into the cast.
- Once the gel has set, the comb is removed, and the gel is placed on the tank.
- Running buffer is added to the tank to cover the gel.
- The samples are prepared by adding loading buffer and carefully pipetted into the wells.
- The tank is connected to the power supply and allowed to run overnight.
c. Denaturation
- The gel is removed from the electrophoresis apparatus and placed in a glass tray with 500 ml denaturation buffer (1.5 M NaCl and 0.5 M NaOH) for 45 minutes at room temperature.
- The denaturation buffer is poured off and replaced with a neutralization buffer. The gel is allowed to soak for 1 hour while slowly rotating on a platform rotator.
d. Blotting
- An oblong sponge that is slightly larger than the gel is placed on a glass dish which is filled with SSC to leave the soaked sponge about half-submerged in the buffer.
- Three pieces of Whatman 3mm paper are cut the same size as the sponge. These are placed on the sponge and wet with SSC.
- The gel is placed on the filter paper and squeezed out to remove bubbles by rolling a glass pipette over the surface.
- A nylon membrane, just large enough to cover the surface of the gel is placed on top of the gel. The membrane is further flooded with SSC, and few sheets of filter paper are placed on top of it.
- Finally, a glass plate is laid on top of the structure to hold everything in place. The DNA transfer is allowed to occur overnight.
e. Baking/ Immobilization
- The nylon membrane is removed from the blotting structure and attached to a vacuum or regular oven at 80°C for 2-3 hours.
- The DNA strands on the membrane can also be immobilized by exposing the membrane to ultraviolet radiation.
f. Hybridization
- The membrane is exposed to the hybridization probe, which can either be a DNA fragment or an RNA segment with a specific sequence that detects the target DNA.
- The probe nucleic acid is labeled so that it can be detected by incorporating radioactivity or tagging the molecules with fluorescent or chromogenic dye.
- The conditions during the process are chosen in a way that the probe hybridizes the target DNA with a complementary sequence on the membrane.
- The hybridization is followed by washing with a buffer to remove the probe that is bound nonspecifically or remain unbound so that only labeled probes remain bound to the target sequence.
g. Detection
- The hybridized regions on the membrane can be detected via autoradiography by placing the nylon membrane in contact with a photographic film.
- The images indicate the position of the hybridized DNA molecules, which can be used to determine the length of the fragments by comparing them with the marker DNA molecules of known length.
- Similarly, the images also provide information about the number of the hybridizing fragments and their size.
- If a fluorescent or a chromogenic dye is used, these can be visualized on X-ray film or by the development of color on the membrane.
Result Interpretation of Southern Blot
- The results of a Southern blot are observed in the form of bands on the membrane. The size of the DNA fragments can be determined by comparing their relative size with the DNA bands of known lengths.
Applications of Southern Blot
- Southern blotting has many applications in the field of gene discovery, mapping, evolution, and diagnostic studies.
- The technique can be used for DNA analysis to detect point mutations and other structural rearrangements in the DNA sequences.
- The method also allows the determination of molecular weights of the restriction fragments, which helps in the analysis of such fragments.
- Since the technique enables the detection of a particular DNA segment, it can be used in personal identification via fingerprinting.
- It can be used in disease diagnosis as well as prenatal diagnosis of genetic diseases.
Limitations of Southern Blot
- The method is costly as it requires expensive equipment and reagents as compared to other tests.
- It is a complex process consisting of multiple steps. The process is also labor intensive that requires trained personnel.
- It is a time-consuming process that can be replaced by other faster processes like Polymerase Chain Reaction.
- It is a semi-quantitative process that only provides estimated sizing of the DNA fragments.
- Southern blotting is not a suitable method for detecting mutations at the base-pair level.
- The sample requires a large amount of sample and higher quality of DNA via superior isolation methods.
References
- Brown T. Southern blotting. Curr Protoc Immunol. 2001 May; Chapter 10:Unit 10.6A. doi: 10.1002/0471142735.im1006as06. PMID: 18432697.
- Southern E. Southern blotting. Nat Protoc. 2006;1(2):518-25. doi: 10.1038/nprot.2006.73. PMID: 17406277.
- Glenn G, Andreou LV. Analysis of DNA by Southern blotting. Methods Enzymol. 2013;529:47-63. doi: 10.1016/B978-0-12-418687-3.00005-7. PMID: 24011036.
Sources
- https://www.mybiosource.com/learn/southern-blotting/ – 14%
- https://www.sciencedirect.com/topics/agricultural-and-biological-sciences/southern-blotting – 11%
- https://en.wikipedia.org/wiki/Southern_blot – 3%
- https://www.sciencedirect.com/science/article/pii/B9780124186873000057 – 1%
- http://www.radio.cuci.udg.mx/bch/ES/papers/SouthernBlot_CurrProtMolBiol_1999-u2.9.1.pdf – 1%
- https://magadhuniversity.ac.in/download/econtent/pdf/blotting.pdf – 1%

Can u upload a video on this procedures
Congratulations to your work
Great work for students and faculty
Thanks so much but how can l download the notes in pdf format
You can only copy and paste our content but cannot download PDF currently. We will make PDF options later on.