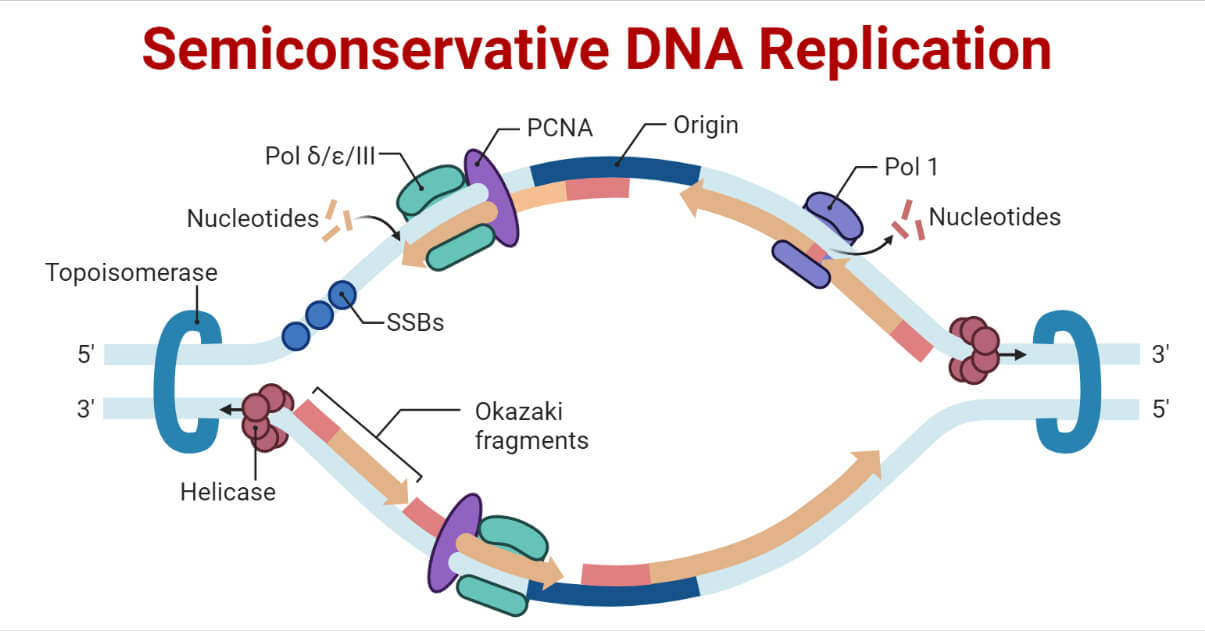
In the semiconservative model of DNA replication, two copies of the original DNA molecule are produced, each copy containing one original strand and one newly synthesized strand.
- Watson and Crick’s double helix model of the DNA molecule features an integrated template system for self-replication or autocatalysis.
- Because of the specificity of base pairing, Adenine to Thymine and Guanine to Cytosine, the base sequence along one chain automatically determines the base sequence along the other.
- This leads to each double helix chain serving as the template for synthesizing the other.
- For the replication of DNA molecules, Watson and Crick proposed that replication involved the disruption of hydrogen bonds followed by a rotation and separation of the two polynucleotide strands.
- Each purine and pyrimidine base of each polynucleotide strand attracts a complementary free nucleotide available for polymerization in the cell, and to hold it in place means of the specific hydrogen bonds.
- Once held in place on the parent template chain, the free nucleotides are linked together by forming the phosphodiester bonds that linked adjacent deoxyribose residues, forming a new polynucleotide molecule of a predetermined base sequence.
- The result of replication is two double-stranded DNA molecules with sequences identical to the original one.
- The original left strand is present in one of these daughter molecules, while the original right strand is in the other.
- Since each progeny retains half of the parent DNA molecule, this replication pattern is considered semi-conservative.
- The DNA replication in prokaryotes and eukaryotes is similar but not the same. The process is more complicated in eukaryotes than in prokaryotes.
In addition to the semiconservative mode, the next two models of DNA replication are equally possible. They are:
- Conservative replication:
The new DNA molecule would consist of two freshly synthesized strands, and both of the parent double helix’s strands are conserved.
- Dispersive replication:
The parent double helix is broken up during replication, and fragments of the parent strands are mixed with newly synthesized strands to form the two new double helices.
Enzymes Involved in DNA Replication and Their Functions
| Enzyme/protein | Specific Function |
| DNA Helicase | It opens the DNA helix by breaking hydrogen bonds between the nitrogenous bases. |
| Topoisomerase | It helps relieve the stress on DNA when unwinding by causing breaks and resealing the DNA. |
| Single-strand binding (SSB) proteins | Binds to single-stranded DNA to avoid DNA rewinding back. |
| Primase | Synthesizes RNA primers needed to start replication. |
| DNA pol I | Exonuclease activity removes RNA primer and replaces it with newly synthesized DNA. |
| DNA pol II | Repair function |
| DNA pol III (α, δ and ε) | Main enzyme that adds nucleotides. |
| Sliding Clamp | It helps hold the DNA polymerase in place when adding nucleotides. |
| DNA Ligase | Seals the gaps between the Okazaki fragments to create one continuous DNA strand. |
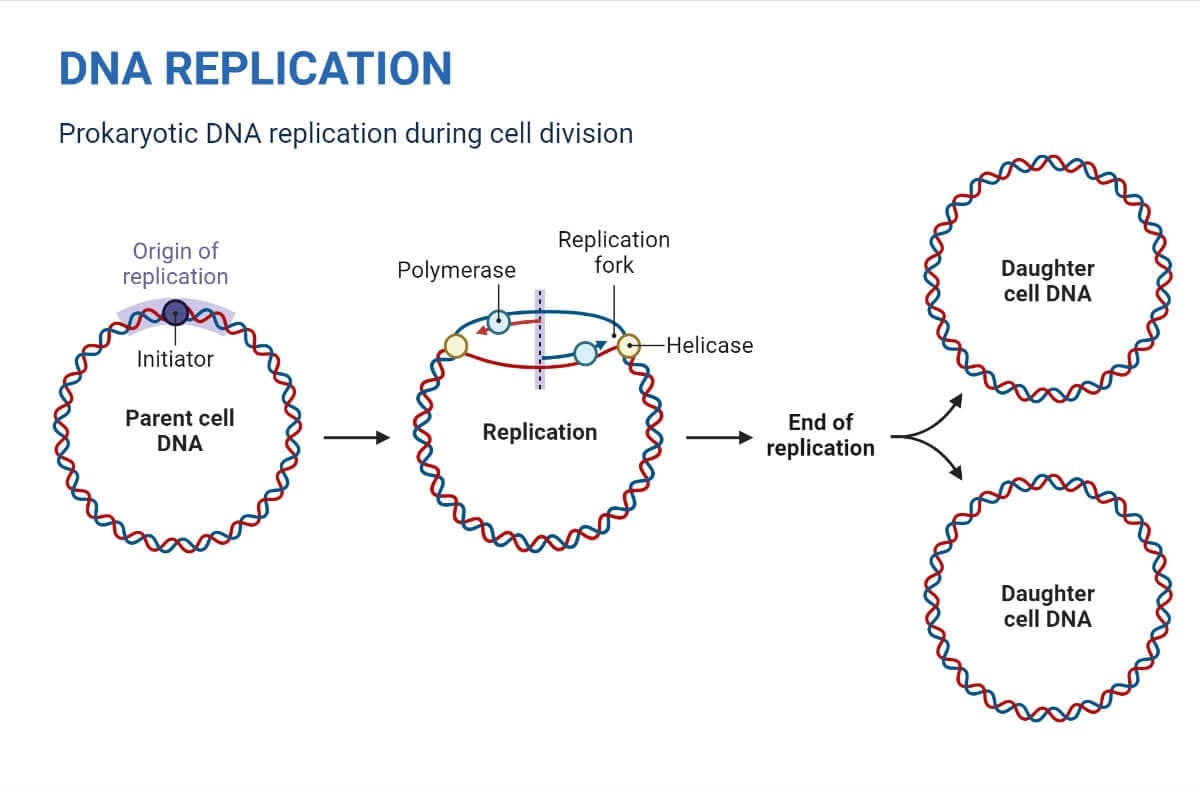
DNA Replication in Prokaryotes
The replication process in prokaryotes can be summarized in 3 major steps:
- Initiation
- Elongation
- Termination
Initiation of DNA Replication in Prokaryotes
- DNA replication begins from the origin of replication.
- On its one chromosome replication origin, OriC is the only replication origin found in Escherichia coli (as do most prokaryotes).
- There are several AT sequences in this origin of replication, which is about 245 base pairs long.
- OriC has several 13 base pair repeats and 9 base pair repeats.
- A cluster of 30 DnaA proteins initially attach to the 9 bp repetitions and bend the DNA to open the DNA helix at the 13 bp repeats.
- Then, DnaC proteins assist DNA helicase (DnaB) inappropriately loading onto the origin.
- DNA helicase unwinds the DNA by severing the hydrogen bonds between the nitrogenous base pairs. This process requires the hydrolysis of ATP.
- Y-shaped structures known as replication forks are formed as the DNA opens up.
- At the origin of replication, two replication forks are formed that grow bi-directionally as replication proceeds.
- Supercoiled DNA is formed when DNA helicase activity topologically stresses the unwinded strand.
- By temporarily nicking the DNA helix and then resealing it, Topoisomerase prevents the DNA double helix from being overwound as the DNA opens up before the replication fork.
- Single-strand binding (SSB) proteins coat the single strands of DNA near the replication fork to prevent the single-stranded DNA from winding back into a double helix.
- DNA polymerase cannot initiate DNA replication. It can only add nucleotides in the 5′ to 3′ direction. It requires a free 3′-OH group to which it can add nucleotides by creating a phosphodiester link between the 3′-OH end and the 5′ phosphate of the following nucleotide. If a free 3′-OH group is not present, it cannot add nucleotides.
- The enzyme RNA Primase synthesizes an RNA segment that is 5 to 10 nucleotides long and complementary to the template DNA. It is known as the primer because this sequence initiates the process for DNA synthesis.
Elongation of DNA Replication in Prokaryotes
- Once priming is complete, DNA polymerase III is loaded into the DNA, and elongation begins.
- The two template DNA strands have opposing orientations, one strand is in the 5′ to 3′ direction, and the other is oriented in the 3′ to 5′ direction.
- The only DNA strand that can be continually synthesized in the direction of the replication fork is the one complementary to the 3′ to 5′ parental DNA strand. This continuously synthesized strand is known as the leading strand and requires only one primer.
- Extending away from the replication fork is the other strand, the lagging strand, which is synthesized in a series of short fragments known as Okazaki fragments, consequently requiring many primers. It is complementary to the 5′ to 3′ parental DNA strand.
- A ring-shaped protein called the sliding clamp holds the DNA polymerase in place as it continues to add nucleotides.
- As synthesis proceeds, the RNA primers are replaced by DNA.
- By using DNA behind the RNA as its own primer and filling in the gaps left by removing the RNA nucleotides by adding DNA nucleotides, DNA Pol I’s exonuclease activity eliminates the primers.
- The gap between the two DNA fragments is sealed by DNA ligase, which helps form phosphodiester bonds.
- During elongation, two replication forks formed at the origin moves in the opposite direction around the circular chromosome.
Termination of DNA Replication in Prokaryotes
- Termination occurs when the two forks meet and fuse, creating two separate double-stranded DNA molecules.
- Replication terminator protein (Tus) binds to termination recognizing (ter) sequences to block replication forks approaching from one direction. These block the movement of DNA helicase.
- The other opposing replication fork halts when it collides with the first one.
- Bacterial circular chromosomes can tangle during replication or even join covalently due to an irregular number of crossovers.
- Topoisomerase IV decatenates tangled circles.
- Finally, the two DNA copies move into two different cells during cell division.
Proof-reading of replicated DNA
- It involves scanning the termini of nascent DNA chains for errors and correcting them during DNA replication.
- This process is carried out by 3′ to 5′ exonuclease activity of DNA polymerase.
- The process removes mispaired nucleotides at 3′ termini so that the correct nucleotide is added by 5′ to 3’ polymerase activity.
- In bacteria, all three DNA polymerases (I, II, and III) have the ability to proofread using 3′ to 5′ exonuclease activity.
- When an incorrect base pair is recognized, DNA polymerase reverses its direction by one base pair of DNA and excises the mismatched base. Following base excision, the polymerase can re-insert the correct base, and replication can continue.
- The extent of proofreading in DNA replication determines the mutation rate.
Evidence for Semi-Conservative Replication of DNA in Prokaryotes
- Using isotopically labeled DNA and an isopycnic density gradient centrifugation technique, M. Meselson, and F.W. Stahl successfully demonstrated the semiconservative nature of DNA replication in 1958.
- For multiple generations, Escherichia coli was cultured in a medium containing NH4Cl with 15N.
- When DNA from these cells is isolated and centrifuged on a salt cesium chloride (CsCl) density gradient, the DNA separates when its density reaches equal to that of the salt solution.
- The DNA of the cells cultured in 15N media had a higher density than the DNA of cells cultured in standard 14N medium.
- E. coli cells having just 15N in their DNA were transferred to a 14N medium and allowed to divide.
- The development of cell division was then observed by microscopic cell counts and by colony assay.
- DNA was extracted periodically and was compared to pure 14N DNA and 15N DNA.
- The density of the DNA was found to be intermediate after one replication.
- Conservative replication was disregarded since it would produce an equal amount of DNA with higher and lower densities (but no DNA with an intermediate density). The result was consistent with both semiconservative and dispersive replications.
- Semiconservative replication would produce double-stranded DNA with one strand of 15N DNA and one of 14N DNA, whereas dispersive replication would produce double-stranded DNA with both strands having mixtures of 15N and 14N DNA. Both of these would have appeared as DNA of an intermediate density.
- DNA from cells after two replications contained an equal amount of DNA with two different densities, one of which matched the intermediate density of DNA from cells grown for only one division in 14N medium and the other matching DNA from cells grown exclusively in 14N medium.
- This was contradictory with dispersive replication, which would have produced a single density, lower than the intermediate density of the one-generation cells but still higher than cells grown only on 14N DNA medium, as the original 15N DNA would have split equally among all DNA strands.
- The result matched the semiconservative replication hypothesis.
DNA replication in Eukaryotes
The replication process in eukaryotes can be summarized in 3 major steps:
- Initiation
- Elongation
- Termination
Initiation of DNA replication in Eukaryotes
- The eukaryotic chromosome has several origins of replication; humans can have up to 100,000 origins of replication.
- At the origin of replication, a pre-replication complex (pre-RC) is made with other initiator proteins. It involves:
- Six Origin Recognition Complex (ORC) proteins (ORC1-6)
- Cell division control (Cdc6) protein
- Chromatin Licensing and DNA Replication Factor (Cdt1) protein
- Six mini chromosome maintenance (MCM) proteins (MCM2-7)
- The process starts in the cell cycle’s G1 phase when the MCM2-7 replicative DNA helicase motors are recruited to each origin of replication.
- Two kinases initiate the activation of the complex once the pre-RC has been created. They are:
- Cyclin-dependent kinase 2 (CdK)
- Dbf4-dependent kinase (DdK)
- DdK and CdK recruit another protein called Cdc45 which then recruits all the DNA-replicating proteins.
- Cdc45, MCM2-7, and the four-subunit complex Go-Ichi-Ni-San (GINS) are combined to form the active CMG helicase during the S phase.
- CMG helicase unwinds the origin.
- Y-shaped structures known as replication forks are formed as the DNA opens up.
- At the origin of replication, two replication forks are formed that grow bi-directionally as replication proceeds.
- Supercoiled DNA is formed when DNA helicase activity topologically stresses the unwinded strand.
- More extensive duplex unwinding occurs due to the association of Replication factor A (RF-A) and Topoisomerase by altering the topology of DNA at the replication fork.
- Single-strand binding proteins coat the single strands of DNA near the replication fork to prevent the single-stranded DNA from winding back into a double helix.
- RNA synthesis of the primer is performed by Primase, which is tightly associated with DNA polymerase α.
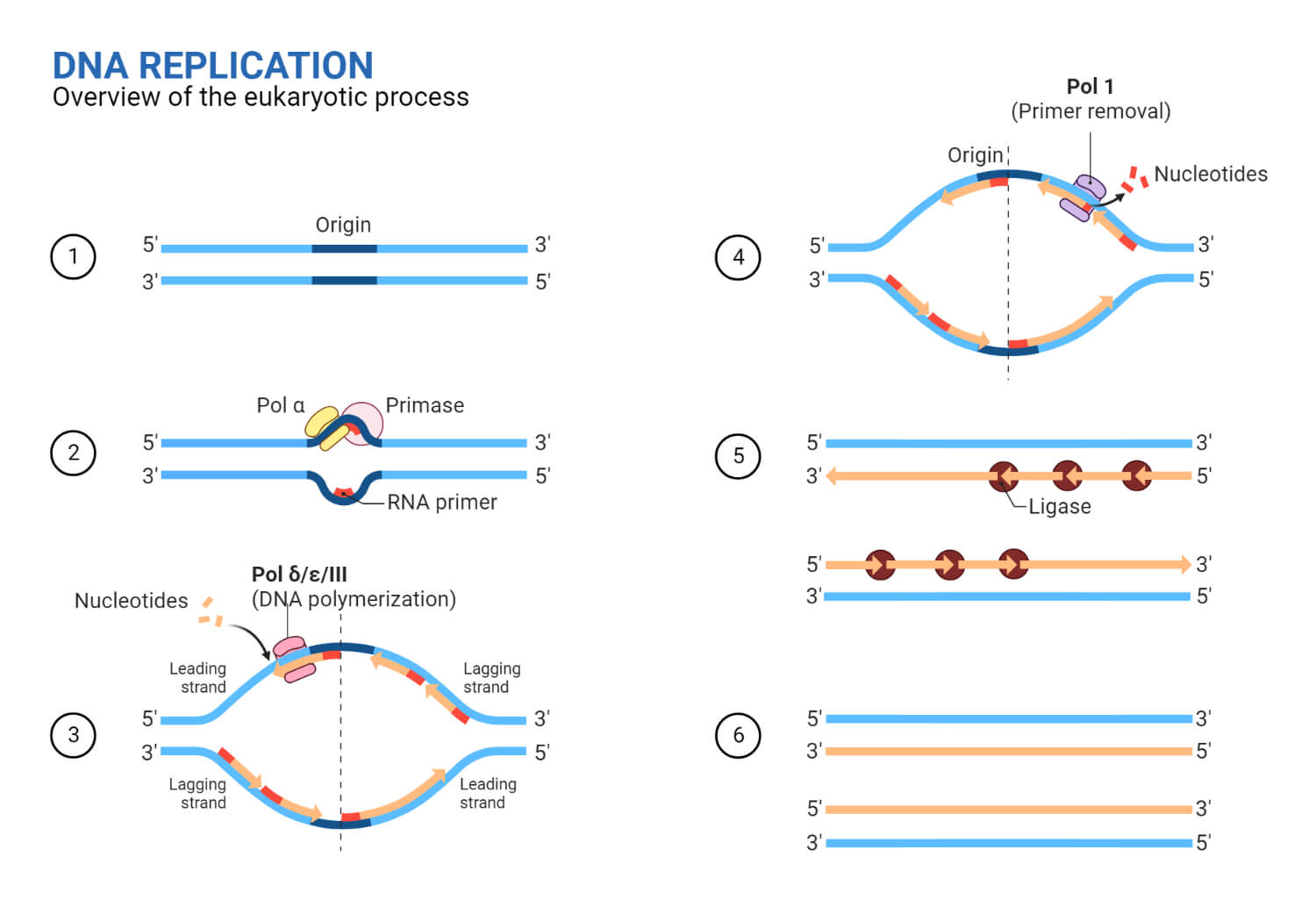
Elongation of DNA replication in Eukaryotes
- Once the initiation complex is formed and the cells pass into the S phase, the complex then becomes a replisome.
- The eukaryotic replisome complex is responsible for coordinating DNA replication.
- DNA polymerase ε synthesizes the leading strand continuously as it points in the same direction as DNA being unwinded.
- DNA polymerase α synthesizes the 20–30 nucleotides during Okazaki fragment initiation, which is further extended by DNA polymerase δ on the lagging strand, which is the opposite DNA template strand, in a fragmented or discontinuous manner.
- The DNA polymerase is held in place so that it does not slide off the DNA by a protein known as PCNA (Proliferating Cell Nuclear Antigen), which functions as a sliding clamp.
- RNase H removes the RNA primer, but one ribonucleotide remains attached to the DNA 3′ end of the Okazaki fragment.
- Flap endonuclease 1 (FEN 1) then removes the ribonucleotide that was left out.
- After the primer is removed, gaps between Okazaki fragments are filled by DNA polymerase δ.
- DNA ligase fills up the nick that forms between the Okazaki fragment and the lagging strand, resulting in the formation of a single lagging strand.
Termination of DNA replication in Eukaryotes
- Termination occurs when two converging replication forks meet. The location is not usually prespecified.
- Midpoint localization depends on the relative start times and progression speed of each converging replication fork.
- When two replication forks converge, the ensuing buildup of torsional strain gets transmitted behind the replication forks, which is dealt with by Topoisomerase II.
- The two replisomes pass each other in opposite directions, and it is thought that the CMG complex slides onto the double-stranded DNA of the last Okazaki fragment.
- A crucial step in the termination of eukaryotic replication is the removal of the CMG helicase from chromatin.
- CMG helicase is removed with the ubiquitination of MCM7, and then Cdc48 comes in and dismantles the replisome.
- Eukaryotic chromosomes are linear, so DNA replication does not reach the very end of the chromosome.
- Since Okazaki fragments require RNA primers attached ahead of the lagging strand, this results in the loss of DNA with each cycle.
- Eukaryotic cells address the problem of DNA shortening during replication using telomere regions of repetitive nucleotide sequences that code for no particular gene at the ends of chromosomes.
- In humans, this repetitive sequence is TTAAGGG. It is repeated 100 to 1000 times in telomere regions and prevents the chromosome ends from being damaged.
- Since the only DNA that gets lost is the meaningless telomere DNA.
- The cumulative DNA loss prevents further division where cells can no longer divide, reaching to point known as the hayflick limit.
- In the germ cell line, which passes DNA to the next generation, as well as in some types of stem cells and white blood cells, the telomerase enzyme prevents cell degradation by extending the telomeres’ repetitive sequence.
- This does not occur in somatic cells, so their telomeres shorten with each round of replication.
- When telomerase is mistakenly activated in somatic cells, it can sometimes lead to cancer.
Proofreading
- Three replicative polymerases replicate the eukaryotic genome; Pol α, Pol δ, and Pol ɛ.
- Due to the absence of internal proofreading, Pol α has low fidelity.
- With an estimated base substitution error rate of 104, Pol α contributes to the synthesis of around 1.5% of the eukaryotic genome and introduces hundreds of mismatches during each replication cycle.
- The exonucleolytic proofreading of Pol δ corrects errors made by Pol α.
- Pol δ-dependent replication errors, as well as errors induced by Pol α, could not be fixed by Pol ɛ; however, Pol δ can proofread errors introduced by Pol ɛ on the leading DNA strand.
Evidence for Semi-Conservative Replication of DNA in Eukaryotes
- By splitting root tip cells of the bean (Vicia faba) and utilizing the techniques of autoradiography and light microscopy, J.H. Taylor and P. Woods, in 1957 provided evidence in support of a semi-conservative form of DNA replication in eukaryotes.
- The root tip cells of the Vicia faba were given radioactive thymidine (3H) treatment to label the DNA molecule. They were subsequently grown in regular, unlabeled media with colchicine to observe the semi-conservative character of chromosome replication.
- To avoid sister chromatid separation during anaphase, colchicine was added.
- In the first generation, the original DNA strand with a 3H label was observed alongside another unlabeled DNA strand because radioactivity was evenly distributed in both strands.
- Only one strand from the second division of DNA exhibited radioactivity.
- This demonstrated the semi-conservative nature of the DNA.
Difference between Prokaryotic and Eukaryotic Replication
| Property | Prokaryotes | Eukaryotes |
| Origin of replication | Only one origin of replication per molecule of DNA. | Multiple origins of replication in each chromosome. |
| Location | Occurs in the cytoplasm | Occurs inside the nucleus. |
| Rate of replication | Replication is rapid.1000 nucleotides/s | Replication is slow.50 to 100 nucleotides/s |
| DNA polymerase types | 5 | 14 |
| Telomerase | Not present | Present |
| RNA primer removal | DNA pol I | RNase H |
| Strand elongation | DNA pol III | Pol α, pol δ, pol ε |
| Sliding clamp | Sliding clamp | PCNA |
References
- Bailey R., Moreno S. P. & Gambus A. (2015). Termination of DNA replication forks: “Breaking up is hard to do”. Nucleus. 6(3), 187-196, https://doi:10.1080/19491034.2015.1035843
- Bębenek, A., & Ziuzia-Graczyk, I. (2018). Fidelity of DNA replication-a matter of proofreading. Current genetics, 64(5), 985–996. https://doi.org/10.1007/s00294-018-0820-1
- Bell S.P. & Dutta A. (2002). DNA replication in eukaryotic cells. Annual Review of Biochemistry. 71, 333–74. https://doi:10.1146/annurev.biochem.71.110601.135425
- Clark D.P, Pazdernik N.J., & McGehee M. R. (2019). Chapter 10 – Cell Division and DNA Replication. In Molecular Biology. Third Edition. Academic Cell, pg 296-331. ISBN 9780128132883. https://doi.org/10.1016/B978-0-12-813288-3.00010-0.
- Dewar, J. M., & Walter, J. C. (2017). Mechanisms of DNA replication termination. Nature Reviews Molecular Cell Biology, 18(8), 507–516. https://doi.org/10.1038/nrm.2017.42
- Meselson, M., & Stahl, F. W. (1958). The Replication of DNA in Escherichia Coli. Proceedings of the National Academy of Sciences of the United States of America, 44(7), 671–682. https://doi.org/10.1073/pnas.44.7.671
- Pandey, M., Elshenawy, M. M., Jergic, S., Takahashi, M., Dixon, N. E., Hamdan, S. M., & Patel, S. S. (2015). Two mechanisms coordinate replication termination by the Escherichia coli tus–ter complex. Nucleic Acids Research, 43(12), 5924–5935. https://doi.org/10.1093/nar/gkv527
- OpenStaxCollege. (2022). DNA Replication in Prokaryotes. Accessed from: https://pressbooks-dev.oer.hawaii.edu/biology/chapter/dna-replication-in-prokaryotes/
- OpenStaxCollege. (2022). DNA Replication in Eukaryotes. Accessed from: https://openstax.org/books/biology-2e/pages/14-5-dna-replication-in-eukaryotes
- OpenStaxCollege. (2022). DNA Replication in Eukaryotes. Accessed from: https://pressbooks-dev.oer.hawaii.edu/biology/chapter/dna-replication-in-eukaryotes/#tab-ch14_05_01
- Taylor, J. H., Woods, P. S., & Hughes, W. L. (1957). The Organization and Duplication of Chromosomes as Revealed by Autoradiographic Studies using Tritium-Labeled Thymidinee. Proceedings of the National Academy of Sciences of the United States of America, 43(1), 122–128. https://doi.org/10.1073/pnas.43.1.122
- Verma P.S. & Agarwal V.K. (2005). Replication of DNA. In Cell Biology, Genetics, Molecular Biology, Evolution and Ecology. Multi-color Edition. S. Chand & Company Ltd. Ram Nagar, New Delhi, pg. 27-31. ISBN 81-219-2442-1
