In bioinformatics, sequence alignment is a fundamental process for comparing protein or DNA sequences, revealing meaningful structural, functional, and evolutionary relationships.

A Scoring system is required while comparing sequences to distinguish homologies from similarities that could have occurred by random chance. It facilitates assigning a quantitative score to an alignment, providing a numerical measure of similarity and also reflecting biological likelihood.
The scoring system consists of a Scoring matrix to score substitutions (matches and mismatches) and Gap penalties to score indels.
What are Scoring matrices?
A scoring matrix or a substitution matrix is a collection of scores for aligning nucleotide or amino acid sequences, where the scores generally represent the relative ease with which one nucleotide or amino acid may mutate into or substitute for another residue.
What do Scoring matrices do?
- Reward conserved substitutions and penalize unlikely ones
- Allow distinguishing biologically meaningful alignments from random matches
- Quantitatively measure the likelihood of a common evolutionary ancestor
The Identity matrix
An identity matrix is the simplest form of a scoring matrix with straightforward logic.
- Exact matches receive a positive score (e.g., +1)
- Mismatches receive a negative score or zero (e.g., −1 or 0)
- No distinction between different types of substitutions.
While it is easy to understand, the primary limitation is that it treats all substitutions (mismatches) as equal, ignoring that substitutions between all amino acids are not equally likely. Hence, it isn’t used to score protein sequence alignments.
The Concept of Amino Acid Substitution
Unlike DNA, the scoring for substitution of residues in protein sequence could be complex because the likelihoods of substitution or mutation between all amino acid pairs are not equal during evolution.
- Some substitutions are conservative (e.g., leucine → isoleucine), which are more likely to preserve protein function and are accepted by evolution. The scoring system should score these substitutions higher because they are more favorable and frequent.
- While others are radical (e.g., glycine → tryptophan), which could disrupt protein structure and function, making these less favorable events. Hence, these should be penalized.
Thus, an effective scoring system must move beyond a simple identity matrix’s match/mismatch model to assigning individual scoring to every pair based on the ease of the substitutions between them.
Log-Odds score
The scores in the substitution matrix represent the likelihood of a given alignment arising from a common evolutionary ancestor versus arising purely by chance. Each score is the logarithm of an odds ratio.
Odds ratio = probability of observing a substitution due to evolution [papb]probability of observing the same substitution by chance [pa,b]
By taking the logarithm of the odds ratio, these multiplicative probabilities are converted into additive scores, making it easier for algorithms to compute the total alignment score.
Hence,
Score[S(a,b)] = log(pap(b)p(a,b))
Interpretation:
- Positive score = This particular substitution occurs more frequently than expected by chance
- Negative score = This particular substitution occurs less frequently than expected by chance; substitution is unlikely.
This powerful log-odds approach is the foundation of scoring matrices used by different algorithms.
PAM (Point Accepted Mutation) Matrix family
Developed by Margaret Dayhoff and colleagues in the late 1960s and 1970s
First systematic method for scoring amino acid substitutions.
It is an explicit evolutionary model derived from studying point mutations through global alignments of 71 closely related protein families sharing at least 85% identity.
A ‘Point Accepted Mutation’ refers to a single amino acid substitution that has been accepted by natural selection because it is either beneficial or does not disrupt the protein’s essential function.
The PAM unit is a measure of Evolutionary distance and divergence, where 1 PAM is an interval in which an average of 1% of amino acids have undergone accepted mutations.
Sequences S1 and S2 are defined as “one PAM unit diverged” if an average of 1 point accepted mutation per 100 amino acid residues can convert the sequence S1 to S2.
Note: Two sequences can be more than 100 PAM units apart because one position can mutate multiple times.
The PAM 1 Matrix was the first PAM matrix created, which represents 1% divergence. It was constructed by following these steps:
- Initial Data Selection: The process began with a collection of 71 groups of closely related protein sequences (at least 85% identical) to ensure that no more than one point mutation occurred at one position.
- Evolutionary Mapping: Multiple global sequence alignments were created for these groups, and phylogenetic trees were constructed to track evolutionary history.
- Inferring Ancestors: Most likely ancestral sequences were inferred at the internal nodes of these trees to count the specific mutations occurring along each branch.
- Calculating Mutability: For each amino acid, its “relative mutability” (ma) was calculated, representing the probability that it will mutate within that 1 PAM unit of time.
- Filling the matrix: The PAM1 matrix table was filled after calculating the probability of any amino acid a mutating to amino acid b over 1 PAM unit of time.
- Log-Odds Conversion: These probabilities were converted into a log-odds scoring matrix, making the scores additive for easier computation.
Matrix Exponentiation
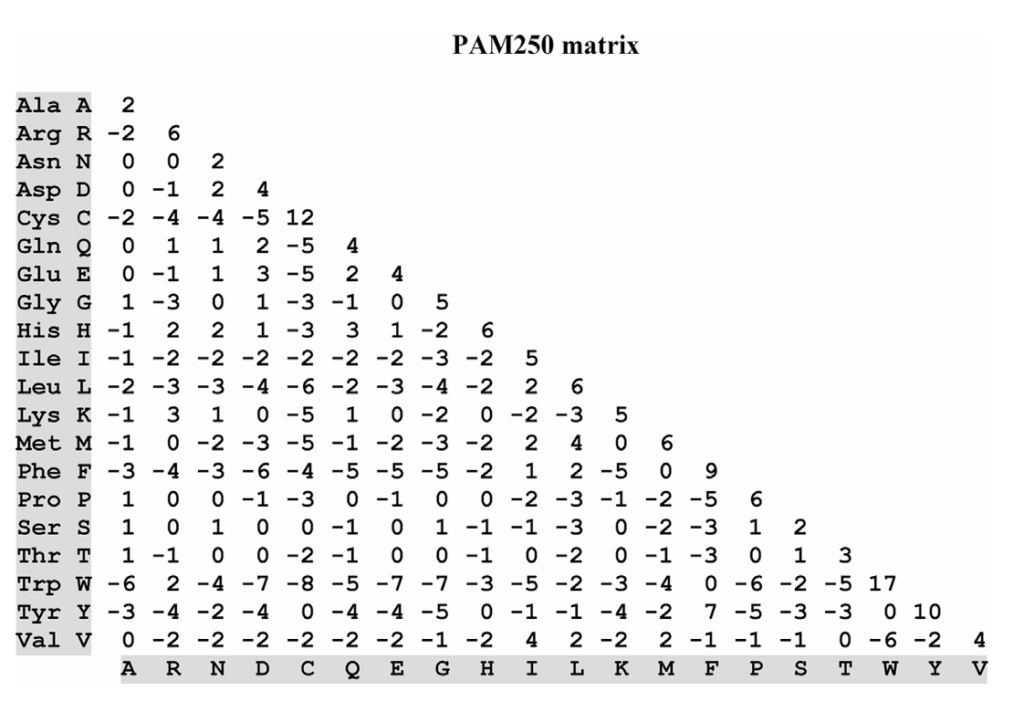
To deal with protein sequences that are more diverged and distantly related, other PAM matrices, such as PAM100 and PAM250, were generated by multiplying the PAM1 matrix by itself n times. For instance, PAM 250 is derived by multiplying the PAM1 matrix by itself 250 times, i.e., (PAM1)250.
Note: A large PAM distance does not correspond linearly to the percentage of different residues in sequences. For example, a PAM250 distance, representing 250 mutations per 100 residues, corresponds to sequences that are roughly 20% identical, not 0%. This is due to back mutations and silent mutations.
Usage:
- Lower-numbered matrices like PAM40 are used for closely related sequences
- Higher-numbered matrices like PAM250 are best for detecting distant sequence similarity during database searches

Limitations of PAM matrices
- Small Dataset: It was originally derived from a relatively small number of protein families.
- Uniformity: It assumes that all positions in a protein are equally mutable, ignoring that the functional and/or structural constraints can make some positions much more conserved than others.
- Error Amplification: Extrapolating to larger distances by repeatedly multiplying the PAM1 matrix can amplify any initial errors in the original PAM1 matrix.
- Less accurate for distant sequences: Because it is based on extrapolation of a matrix derived from closely related sequences, and also matrices for divergent sequences like PAM250 can sometimes cause alignment overextension, where the algorithm incorrectly extends a match into neighboring non-homologous regions.
BLOSUM (Blocks Substitution Matrix) Matrix family
Introduced by Steven and Jorja Henikoff in 1992, using a much larger and more diverse dataset than the PAM matrix.
BLOSUM is based on local multiple alignments of highly conserved regions known as “blocks”. These blocks are gapless segments shared by groups of closely as well as distantly related proteins.
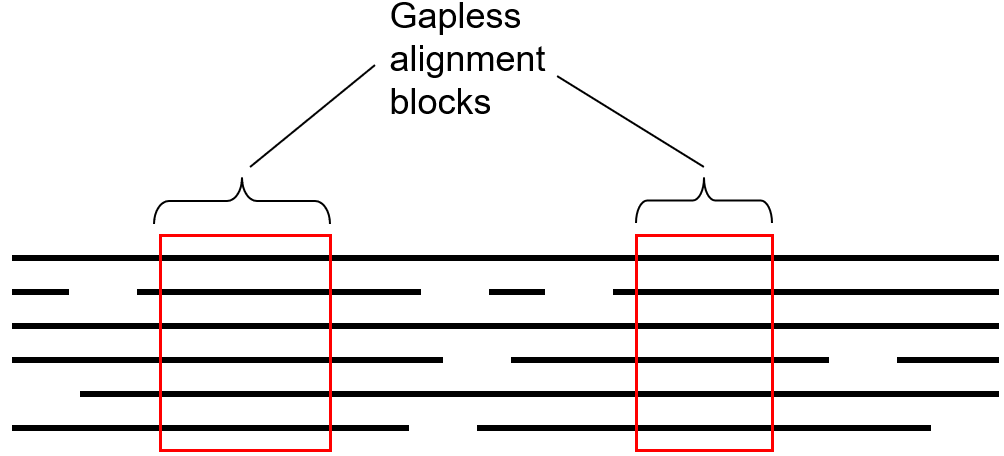
Steps in the construction of BLOSUM matrices
- Data source: BLOCKS database, a database of highly conserved aligned sequences, is used. Only the sequences within blocks are used to calculate substitution frequencies.
- Clustering: Related sequences with similarity higher than a set threshold percentage are clustered together. This prevents over-weighting contributions from very similar sequences.
For example, in constructing BLOSUM80, similar blocks with more than 80% similarity are clustered and treated as a single segment.
- Direct Calculation: Frequencies of amino acid substitutions are then counted by comparing sequences between different clusters, and log-odds scores are calculated directly from the observed frequencies.
Usage:
- Higher BLOSUM numbers (such as BLOSUM80) are optimized for closely related sequences
- Lower numbers (such as BLOSUM45) are more sensitive for detecting remote sequence similarities.
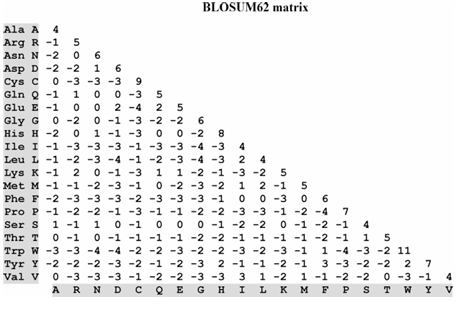

Source: Choudhuri, S. (2014). Bioinformatics for Beginners
Advantages of BLOSUM:
- Large dataset: BLOSUM is built using a significantly larger and more representative dataset (over 500 protein families) in comparison to PAM’s dataset (71 families).
- Empirical model: Unlike the extrapolations in PAM matrices, BLOSUM values are derived from direct observation for different similarity thresholds.
- High sensitivity: They generally perform better than PAM for detecting distant homologs.
Limitations of BLOSUM:
- Alignment Overextension: Deep matrices (such as BLOSUM62) can cause homologous overextension, where the algorithm incorrectly extends a high-scoring local alignment into neighboring non-homologous regions.
- Ineffectiveness for Short Sequences: Deeper matrices could lack the information content required to produce statistically significant scores for short protein sequences.
Differences Between PAM and BLOSUM
| PAM | BLOSUM | |
| Construction Model | Based on an explicit evolutionary model | Based on Empirical observed frequencies |
| Alignment Type | Based on Global alignments | Based on Local blocks |
| Data Source | Small set of closely related proteins | Large, diverse set of conserved blocks |
| Scaling | Extrapolated from PAM1 by multiplying it by itself n times. | Based on observed frequencies from alignments between certain threshold identity percentages. |
| Usage | Low Number = for closely related sequencesHigh Number = for distantly related sequences | Low Number = for distantly related sequences High Number = for closely related sequences |
Rough Equivalents:
- PAM120 ≈ BLOSUM80
- PAM160 ≈ BLOSUM62
- PAM250 ≈ BLOSUM45
Choosing the Right Matrix
Choosing the right substitution matrix is critical as it strongly influences the outcome of sequence alignment. The choice of the substitution matrix depends mainly on how similar you expect the sequences to be, i.e., the evolutionary distance between them.
The matrices are often categorized as deep and shallow in this regard.
-Shallow matrices: Matrices for comparing closely related sequences with high similarity(505 to 905), e.g., orthologs from closely related species.
- low-numbered PAM matrix (e.g., PAM40).
- high-numbered BLOSUM matrix (e.g., BLOSUM80)
-Deep matrices: Matrices for comparing distantly related Sequences with low similarity(20% to 30%), e.g., homologs across different phyla.
- high-numbered PAM matrix (e.g., PAM250).
- low-numbered BLOSUM matrix (e.g., BLOSUM62 or BLOSUM45).
BLOSUM62 is often chosen as the default matrix and is also the standard for popular search tools like BLAST because it is balanced to perform well across a wide range of evolutionary distances.
If the evolutionary distance between sequences is unknown, a common strategy is to perform a quick alignment with an identity matrix to estimate the distance, then select the corresponding matrix.
Also, newly developed specialized matrices can be chosen for proteins with biased compositions, for example, JTTtm and PHAT matrices for cysteine-rich transmembrane proteins.
Gap Penalty and Matrix Interaction
A complete scoring system for alignments needs a way of scoring gaps along with the substitution matrix. Gaps in alignments are created due to indels, which are less likely than substitutions, hence need to be penalized.
Gap penalties are adjusted by users according to their scoring scheme. The choice of the penalty value significantly changes the resulting alignment:
- No or Low Gap Penalties: Allow gaps to be inserted frequently to prevent mismatches; overrepresents similarity.
- High Gap Penalties: Force the algorithm to align dissimilar residues rather than creating a gap.
Gap penalty models
- Constant Penalty: This assigns a fixed penalty to a gap regardless of its length. This is rarely used in high-precision biological comparisons.
- Linear Gap Penalty: The penalty is directly proportional to the length of the gap. If the penalty per unit length is B and the gap length is I, the total penalty is BI. While computationally simple, it is often considered less realistic because it treats a single deletion of five residues as five separate, independent evolutionary events.
- Affine Gap Penalty: This is the most sophisticated model used in modern bioinformatics. It distinguishes between opening a new gap and extending an existing one.
Gap penalty(G) = A+ BI
Where,
A is called the Gap-opening penalty,
B is the Gap-extension penalty, and
I is the length of the gap, where A is much higher than B.
The rationale for the affine model is that a single, long indel event is evolutionarily more probable than many small, scattered indel events. Therefore, alignment algorithms are designed to encourage extending existing gaps rather than starting new ones.
Conclusion
The development of scoring matrices has proved to be crucial for sequence alignment. Its application has transformed a simple match/mismatch comparison to a more sophisticated, informed quantitative method for distinguishing true homology from random similarity. The PAM matrix family, which pioneered scoring matrices through explicit evolution-based extrapolation, had limitations due to the use of a small dataset of closely related sequences.
BLOSUM matrices were later developed through empirical observations using a larger, more diverse dataset from the BLOCKS database. This made BLOSUM a preferred choice for diverse sequences, with BLOSUM62 serving as the gold standard. The choice of an appropriate matrix, however, depends upon the evolutionary distance between the sequences being compared.
Thoughtful use of shallow and deep substitution matrices, combined with a carefully calibrated gap penalty scheme, is essential for producing accurate and meaningful alignments and for drawing reliable functional and evolutionary inferences.
References
- Choudhuri, S. (2014). Fundamentals of Genes and Genomes. In Bioinformatics for Beginners (pp. 1–25). Elsevier. https://doi.org/10.1016/b978-0-12-410471-6.00001-3
- Mount, D. W. (2008). Using BLOSUM in sequence alignments. Cold Spring Harbor Protocols, 3(6). https://doi.org/10.1101/pdb.top39
- Pearson, W. R. (2013). Selecting the right similarity-scoring matrix. Current Protocols in Bioinformatics, (SUPL.43). https://doi.org/10.1002/0471250953.bi0305s43
- Przytycka, T. (n.d.). Lecture 3 Scoring Matrices Position Specific Scoring Matrices Motifs. Principles of Computational Biology.
- Trivedi, R., & Nagarajaram, H. A. (2020). Substitution scoring matrices for proteins – An overview. In Protein Science (Vol. 29, Issue 11, pp. 2150–2163). John Wiley and Sons Inc. https://doi.org/10.1002/pro.3954
