RNA Capping is the addition of a 7-methylguanosine (m7G) structure to the 5′ end of RNA transcripts for regulating RNA stability, metabolism, and biological function.
In eukaryotes, co-transcriptional addition of a methylated guanosine serves as a unique molecular mark for mRNA.
- Cellular metabolites such as NAD and CoA are utilized by bacteria as non-canonical caps to modify their regulatory small RNAs and mRNAs.
- RNA caps are chemically diverse across all domains of life, encompassing canonical m7G structures alongside metabolite-derived caps and methylated phosphate structures.
Structure of the 5′ Cap
To regulate the life cycle of RNA, both eukaryotes and prokaryotes possess diverse 5′ terminal structures.
In Eukaryotes:
- The 5′ Cap is a highly regulated structure primarily found on mRNAs transcribed by RNA polymerase II.
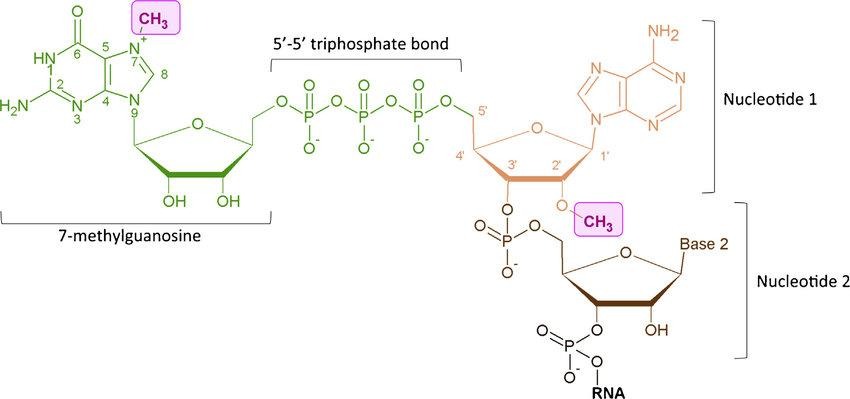
- Methylation of a guanosine nucleotide at the N7 position (m7G) provides a positive charge that is essential for recognition by nuclear and cytoplasmic cap-binding proteins.
- The m7G is linked to the first encoded nucleotide of the RNA chain via a reverse 5′ to 5′ triphosphate linkage (m7GpppN). This inverted structure protects the RNA from digestion by 5’–3′ exoribonucleases.
- Small nuclear RNAs (snRNAs) often undergo further modification to form a 2,2,7-trimethylguanosine (TMG) cap.
- Small RNAs like U6 possess a simpler structure where a methyl group is added directly to the terminal γ-phosphate of the initiating nucleotide.
In Prokaryotes:
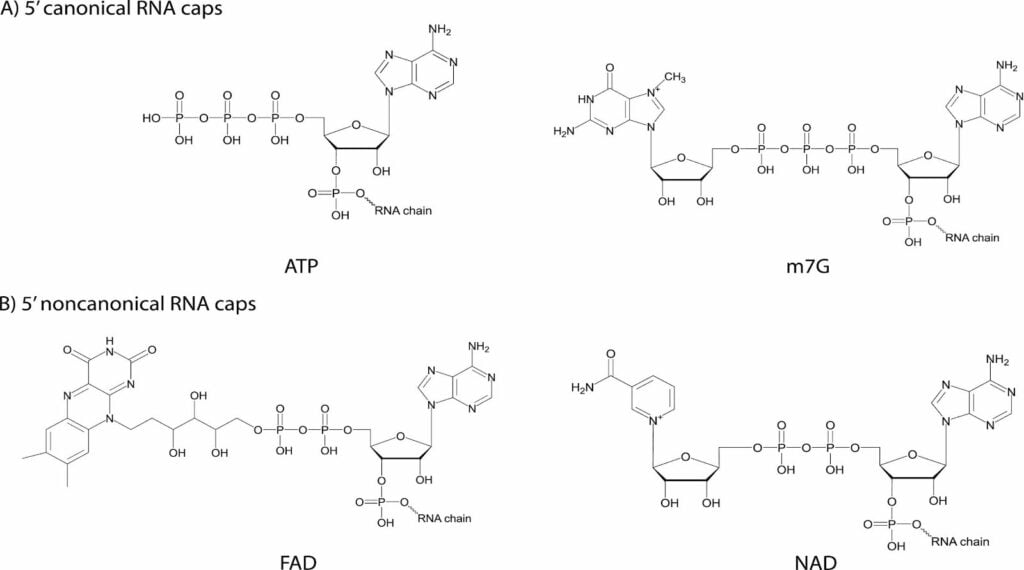
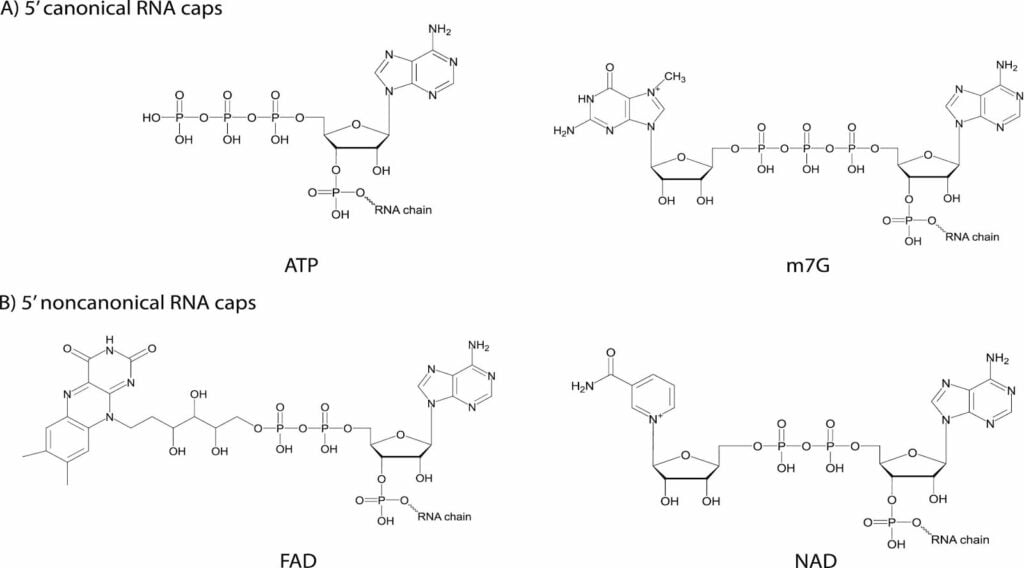
- Instead of possessing the m7G cap, bacteria utilize various cellular metabolites as non-canonical caps incorporated during transcription initiation.
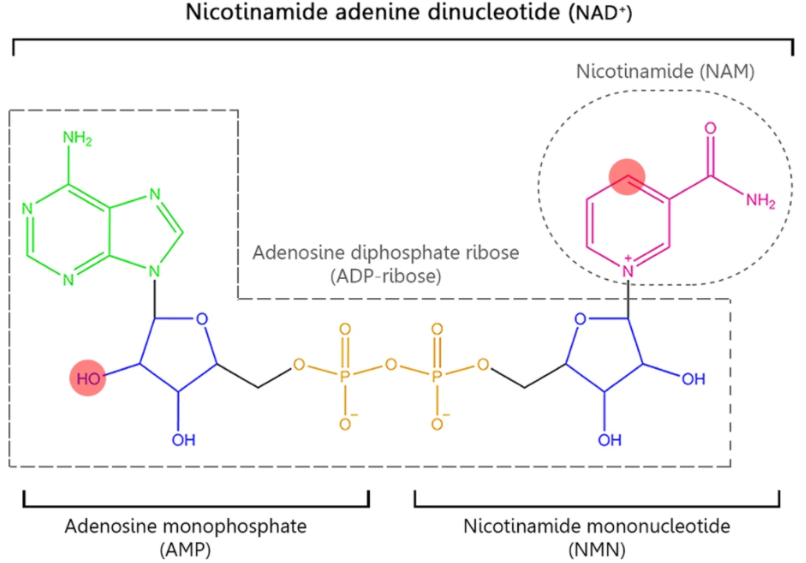
- Cofactors such as Nicotinamide Adenine Dinucleotide (NAD), Flavin Adenine Dinucleotide (FAD), and 3′-dephospho-Coenzyme A (dpCoA) are used for RNA capping in bacteria.
- These metabolites are attached via their adenosine moiety, which acts as the initiating nucleotide for RNA synthesis. For example, in an NAD cap, the nicotinamide riboside is linked to the adenosine via a pyrophosphate bond that begins the RNA chain.
- Bacteria also use uridine-containing sugars like UDP-Glucose (UDP-Glc) and UDP-N-acetylglucosamine (UDP-GlcNAc) as 5′ caps.
- Under stress conditions, bacteria incorporate molecules called alarmones like Ap4A or Ap3G (NpnN) at the 5′ end. Their concentration increases during metabolic shifts.
- Many metabolite caps, such as NAD, FAD, and UDP-sugars, have also been discovered on eukaryotic mRNAs and mitochondrial RNAs, though they are much less abundant than the canonical m7G cap.
The Timing of Capping
- It refers to the specific stage during transcription when the 5′ end of a nascent RNA molecule is chemically modified.
- The timing and mechanism of these events differ significantly for eukaryotes and prokaryotes.
In Eukaryotes
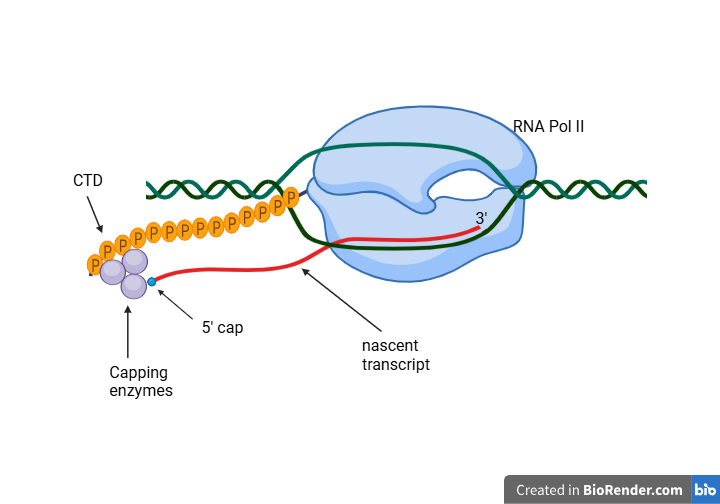
- Capping occurs in the nucleus while RNA Polymerase II is still synthesizing the RNA chain.
- This process is triggered when the nascent transcript reaches a length of approximately 20–30 nucleotides.
- Capping enzymes are recruited to the Serine-5 phosphorylated C-terminal domain (CTD) of RNA Polymerase II. This positioning allows the enzymes to act on the RNA immediately as it emerges from the polymerase exit channel.
- Some RNAs, such as processed snoRNAs or stored mRNAs, can undergo recapping in the cytoplasm after being decapped, serving as a secondary regulatory mechanism.
In Prokaryotes
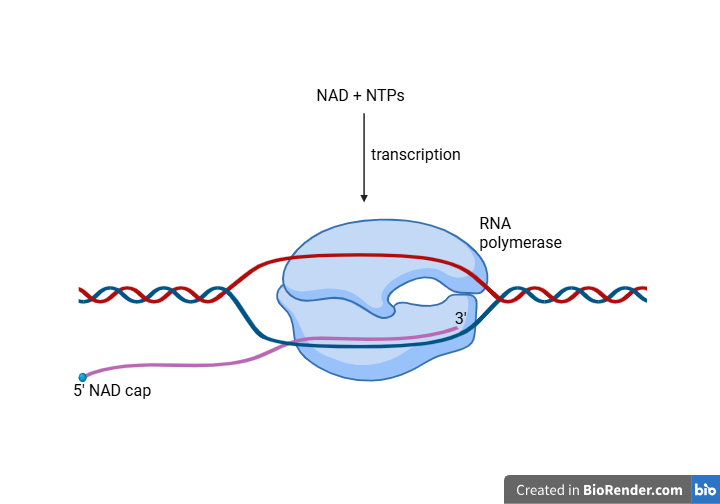
- In bacteria, the capping occurs immediately as transcription is initiated.
- The bacterial RNA Polymerase directly utilizes a metabolite such as NAD+, FAD or CoA as the initiating nucleotide in place of a standard ATP or GTP.
- Thus, the cap is incorporated into the RNA from the very beginning, and not added later.
- This is known as the Non-Canonical Initiating Nucleotide (NCIN) mechanism.
Enzymes Involved in Capping
- RNA capping is performed by specialized enzymes that modify the 5′ terminal of a nascent RNA transcript.
- These enzymes vary between the canonical system in eukaryotes and the recently discovered metabolite-based system in prokaryotes.
Eukaryotic Capping Enzymes
- To produce the m7G cap, the following sequential enzymatic activities are involved.
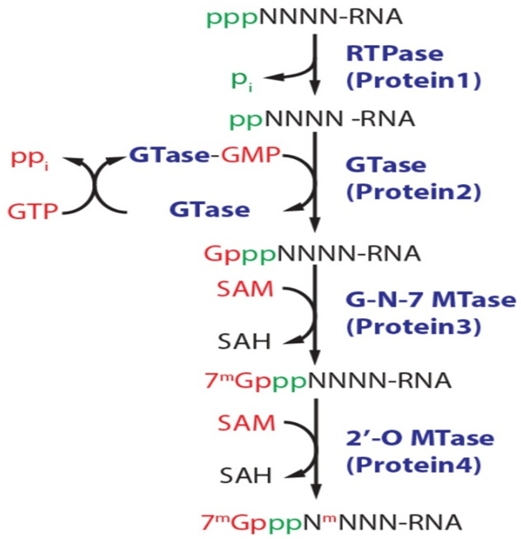
RNA Triphosphatase (RTPase): This enzyme removes the terminal (γ) phosphate from the 5′ triphosphate end (pppN) of the nascent RNA, resulting in a 5′ diphosphate RNA (ppN).
RNA Guanylyltransferase (GTase): This enzyme transfers a GMP group from GTP to the 5′ diphosphate RNA. This occurs via a two-step “ping-pong” mechanism involving a covalent lysyl-GMP intermediate.
Guanine-N7 Methyltransferase (G-N-7MTase): This enzyme adds a methyl group from S-adenosyl-L-methionine (SAM) to the N7 position of the added guanosine, creating the functional Cap 0 structure.
- In Metazoans, the additional enzyme Ribose Methyltransferases (CMTR1/CMTR2) methylate the 2′-O position of the first and second transcribed nucleotides to form Cap 1 and Cap 2 structures.
- In mammals/vertebrates, the RTPase and GTase activities are combined into a single bifunctional protein called RNGTT.
- In yeast, these activities are carried out by Cet1 (RTPase), Ceg1 (GTase), and Abd1 (N7MTase).
Prokaryotic Capping Enzymes
- RNA Polymerase (RNAP): In bacteria, the RNA polymerase itself acts as the primary capping enzyme by directly incorporating metabolites such as NAD+, FAD, or dpCoA, instead of a standard nucleoside triphosphate during the first step of transcription initiation.
- Lysyl-tRNA Synthetase (LysU): This enzyme is capable of installing alarmone caps (dinucleoside polyphosphates like Ap4A) onto 5′ triphosphorylated transcripts under stress conditions.
- NudC: The primary bacterial enzyme that regulates the stability and turnover of the capped RNAs by removing NAD+ caps.
Mechanism of Cap Synthesis
Eukaryotic Mechanism
In eukaryotes, the synthesis of the 7-methylguanosine (m7G) cap occurs co-transcriptionally through the following sequential enzymatic steps.
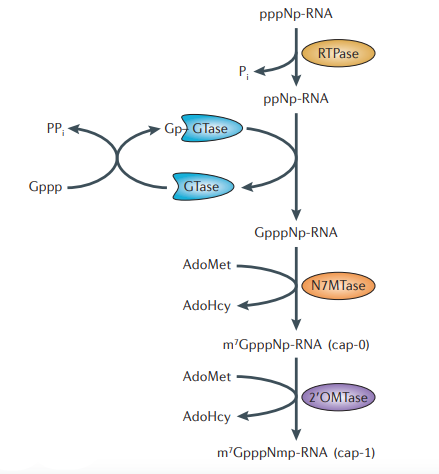
- Terminal Dephosphorylation: RNA triphosphatase (RTPase) removes the terminal γ-phosphate from the nascent 5′-triphosphorylated RNA (pppRNA), producing 5′-diphosphorylated RNA (ppRNA) and releasing inorganic phosphate (Pi).
- Guanylylation (GMP Transfer): The guanylyltransferase (GTase) catalyzes a “ping-pong” reaction. First, it reacts with GTP to form a high-energy covalent lysyl-GMP intermediate while releasing pyrophosphate (PPi). The enzyme then transfers this GMP group to the 5′-diphosphate end of the RNA, creating a unique reverse 5’–5′ triphosphate bridge (GpppN).
- N7-Methylation: The guanine-N7 methyltransferase (N7MTase) transfers a methyl group from S-adenosyl-L-methionine (SAM) to the N7 position of the added guanosine cap. This forms the Cap 0 structure (m7GpppN).
- Ribose Methylation: In higher eukaryotes, 2′-O-methyltransferases further modify the RNA by adding methyl groups to the ribose sugars of the first and second nucleotides, resulting in Cap 1 and Cap 2 structures, respectively.
Prokaryotic Mechanism
- Many non-canonical caps in bacteria and some eukaryotic organelles are incorporated at the very start of RNA synthesis.
- Transcription Initiation: Bacterial RNA polymerase (RNAP) initiates transcription by using a non-canonical initiating nucleotide (NCIN) such as NAD+, FAD or dpCoA, instead of a standard ATP or GTP.
- Direct Incorporation: The metabolite is incorporated into the RNA at the very beginning as part of the first phosphodiester bond formed.
- Determinants: The efficiency of this process is dictated by promoter sequences and the specific architecture of the “Rif-pocket” within the RNA polymerase, which stabilizes the metabolite during initiation.
Types of RNA Caps (Cap 0, Cap 1, Cap 2)
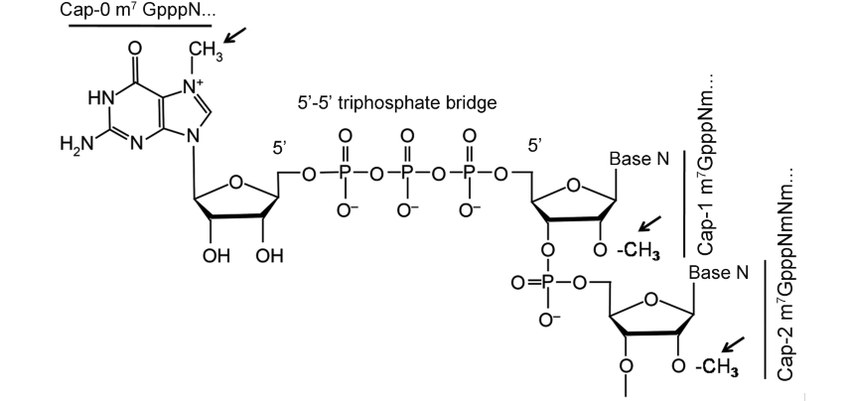
The standard eukaryotic cap consists of a guanosine methylated at the N7 position (m7G) and linked to the transcript via a 5’–5′ triphosphate bridge.
The following types are classified based on additional methylations:
- Cap 0 (m7GpppN): Only the terminal guanosine is methylated at the N7 position. It is the predominant form in lower eukaryotes, such as yeast, and serves as a precursor to more complex caps in higher organisms.
- Cap 1 (m7GpppNm): The ribose sugar of the first transcribed nucleotide is 2′-O-methylated. It is the most abundant cap in mammals and metazoans. It allows the innate immune system to distinguish cellular mRNA from viral RNA.
- Cap 2 (m7GpppNmpNm): The ribose sugars of both the first and second transcribed nucleotides are 2′-O-methylated. This type is common in higher eukaryotes and further aids in immune evasion.
Functions of the 5′ Cap
- RNA Stability and Protection: The cap protects the 5′ end of mRNA from degradation by 5′–3′ exoribonucleases, such as XRN1. Transcripts lacking this protection are rapidly targeted to cytoplasmic processing bodies (P-bodies) for decay.
- Translation Initiation: The m7G cap is recognized by the eukaryotic translation initiation factor 4E (eIF4E), which anchors the recruitment of the ribosome to the mRNA. It also facilitates mRNA “pseudo-circularization” through interactions between cap-binding proteins and poly(A)-binding proteins, enhancing translation processivity.
- Nuclear Export: The cap-binding complex (CBC) recognizes the structure in the nucleus and interacts with export factors like REF/Aly to guide the mRNA through the nuclear pore complex into the cytoplasm.
- Pre-mRNA Processing: The cap facilitates the recruitment of the spliceosome, promoting the efficient splicing of the first intron. It also plays a role in 3′ end processing by stabilizing the cleavage complex at the polyadenylation site.
- Immune Discrimination: Methylation of the first and second nucleotides (Cap 1 and Cap 2) serves as a molecular signature that allows the innate immune system to recognize the RNA as “self”. Caps lacking 2′-O-methylation can activate antiviral sensors such as RIG-I and MDA5, triggering interferon production.
- Transcriptional Regulation: The capping machinery itself, such as the RNMT-RAM complex, has been shown to stimulate the transcription process independently of the methylation event.
The Cap-Binding Complex
- It is a specialized protein assembly that recognizes and binds the 5′ cap of RNA.
- This complex connects transcripts to the cellular machinery for processing, transport, and translation.
- Viruses like Influenza use the PB2 subunit, which is unique and distinct from the host CBC, to bind and snatch host caps.
Eukaryotic Cap-Binding Complex
- The CBC is located predominantly in the nucleus.
- The mammalian CBC is a heterodimer is composed of NCBP2 (CBP20) and NCBP1 (CBP80). NCBP2 directly binds the methylated guanosine with its interacting partner NCBP1.
- NCBP2 has a high specificity for N7-methylated guanosine (m7G) with over 150-fold greater affinity for m7GpppG than for unmethylated GpppG.
- NCBP2 binds co-transcriptionally to facilitate splicing of the first intron, stabilizes the 3′ cleavage complex for polyadenylation, and interacts with export factors such as REF/Aly for nuclear export.
- Upon export, it initiates the pioneer round of translation, which is essential for quality control via nonsense-mediated decay (NMD).
- For steady-state protein synthesis, the CBC is replaced by the eIF4F complex (specifically eIF4E) after the first round of translation.
- NCBP3 can replace NCBP2 to form an alternative complex with NCBP1, which plays a specific role in clearing viral infections.
Decapping and Turnover
- These are the regulatory processes by which the 5′ cap is enzymatically removed for the rapid degradation of an RNA molecule.
- This step is critical for controlling the lifespan of mRNA and managing cellular responses to stress.
Eukaryotic Decapping and Turnover
- The primary enzyme for removing the standard m7G cap is Dcp2, which is a member of the Nudix hydrolase family.
- It hydrolyses the triphosphate bridge, releasing m7GDP and leaving a 5′ monophosphate terminus on the RNA.
- The DXO/Rai1 family of enzymes removes incomplete or unmethylated caps and prevents them from escaping normal mRNA decay.
- Metabolite caps like NAD+, FAD, and dpCoA are removed by specialized enzymes such as DXO and Nudt12.
- Once the cap is removed, the resulting 5′ monophosphate RNA is rapidly digested by the 5’–3′ exoribonuclease XRN1. This decay often occurs in cytoplasmic granular compartments known as P-bodies.
Prokaryotic Decapping and Turnover
- Bacteria utilize the phosphohydrolase NudC as their primary decapping enzyme for NAD-capped RNAs.
- NudC hydrolyzes the pyrophosphate bond within the NAD cap to release nicotinamide mononucleotide (NMN).
- The enzyme RppH (RNA pyrophosphohydrolase) acts as a functional analog to decapping by converting 5′ triphosphate RNA into 5′ monophosphate RNA. This conversion marks the RNA for further degradation.
- Removal of the bacterial cap triggers internal cleavage by the endonuclease RNase E or processive degradation by RNase J1.
- For stress-induced alarmone caps (NpnN), the enzyme ApaH functions as a decapping factor by hydrolyzing the polyphosphate chain.
Viral Capping Strategies
Viruses produce a 5′ terminal structure which is identical or similar to the host’s canonical m7G cap, to protect against exonucleases and serve as a marker of “self”.
Some of the capping strategies of the virus are as follows: –
- Conventional Viral Capping:
-Many large DNA viruses and some double-stranded RNA (dsRNA) viruses utilize a three-to-four-step enzymatic pathway identical to that of the host.
-The enzymes involved are RNA triphosphatase (RTPase), a guanylyltransferase (GTase), and one or more methyltransferases (MTases).
-In viruses like Vaccinia, these activities are often combined into a single multifunctional protein complex, e.g., the D1/D12 heterodimer, that acts as a cap assembly line.
-Poxviruses, Reoviruses, and Mimiviruses utilize this strategy.
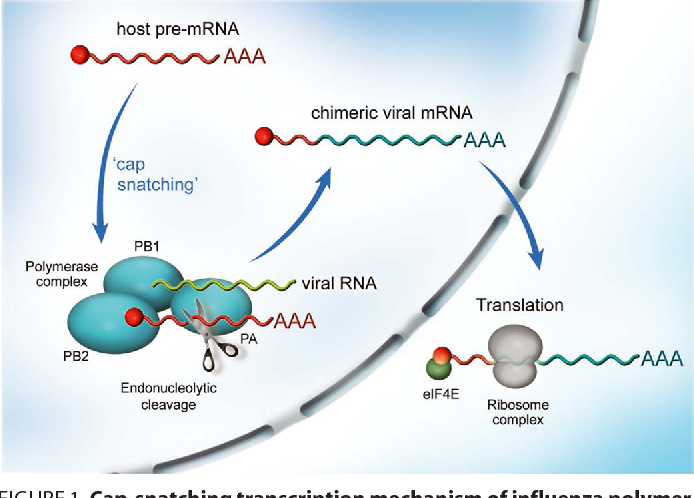
- Cap Snatching:
-Segmented negative-strand RNA viruses snatch the 5′ cap from host cellular mRNAs because they lack their own capping enzymes.
-During transcription, the PB2 subunit binds the 5′ cap of the host pre-mRNA.
-The PA subunit in Influenza cleaves the host mRNA 10–20 nucleotides from the 5′ end. This short, capped fragment is then used as a primer to initiate viral transcription by the RNA-dependent RNA polymerase of the PB1 subunit.
-The cap snatching process simultaneously provides the virus with a functional cap and degrades host mRNAs, effectively shutting down host protein synthesis in favor of viral production.
-This is the primary strategy for Influenza, Arenaviridae, and Bunyaviridae.
- Unconventional Synthesis Pathways:
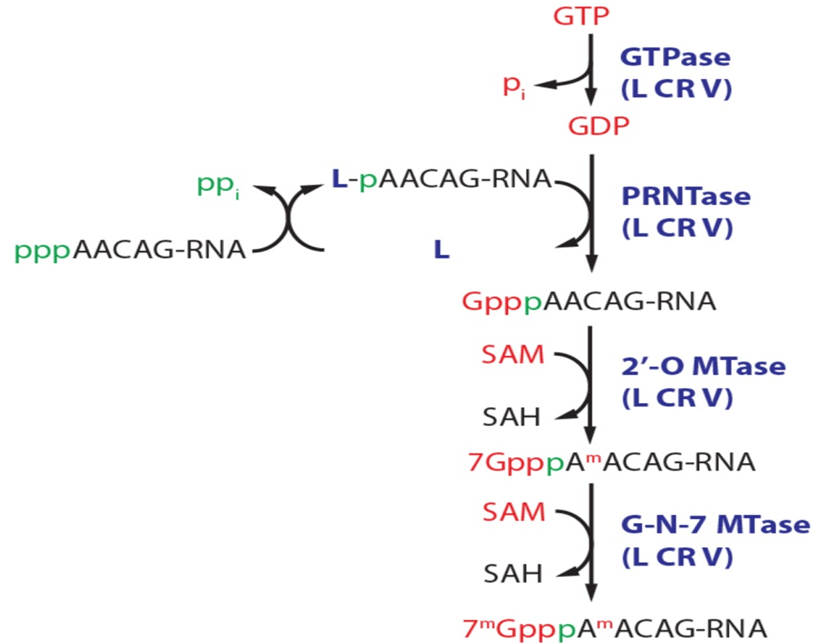
-Non-segmented negative-strand (NNS) viruses, such as Vesicular Stomatitis Virus (VSV), utilize a unique polyribonucleotidyltransferase (PRNTase) activity within their L protein. Rather than transferring GMP to RNA, the enzyme forms a covalent protein-RNA intermediate and transfers the nascent RNA onto a GDP molecule to form the cap.
-Alphaviruses, such as Chikungunya, methylate a GTP molecule first to form m7GTP, which then creates a covalent m7GMP-enzyme intermediate before being transferred to the 5′ end of the viral RNA.
-Coronaviruses, including SARS-CoV-2, employ a novel mechanism where the NiRAN domain of the nsp12 protein transfers nascent RNA to the nsp9 protein, forming a phosphoramidate bond, before transferring it to GDP.
- Non-Canonical Metabolite Capping:
-Instead of the standard m7G, some viruses utilize cellular metabolites as caps.
-Hepatitis C Virus (HCV) initiates its replication using FAD as a non-canonical initiating nucleotide, which also helps it evade the host immune sensor RIG-I.
-Dengue virus particles have also been found to contain transcripts capped with UDP-N-acetylglucosamine.
- Cap-Independent Strategies:
-Viruses in the Picornaviridae and Caliciviridae families covalently attach a small protein called VPg to the 5′ end of their RNA instead of a chemical cap.
-VPg can directly interact with host translation initiation factors like eIF4E to start protein synthesis.
-Some viruses use an Internal Ribosome Entry Site (IRES), a complex secondary RNA structure that recruits ribosomes directly without the need for a 5′ cap or its associated binding proteins.
Conclusions
RNA capping identifies transcripts and serves as a master regulator of an RNA molecule’s stability, transport, and overall function. The standard cap is a methylated guanosine (m7G) attached via a unique triphosphate bridge. Non-canonical versions can use cellular metabolites like NAD+ or FAD. Most eukaryotes add the cap co-transcriptionally when the RNA is very short.
However, the bacteria incorporate metabolite caps at the very first step of transcription initiation. Eukaryotes use a set of three specific enzymes, such as RTPase, GTase, and MTase, to build the cap, whereas bacteria typically use their RNA polymerase to initiate with a cap. Synthesis follows a strict chemical sequence of removing a phosphate, adding a guanosine through a high-energy intermediate, and finally attaching a methyl group. Caps are defined by their level of methylation, moving from the simple Cap 0 in yeast to the more complex Cap 1 and 2 used by higher organisms to avoid immune detection.
The cap protects the RNA from enzymatic degradation and anchors the machinery needed to translate it into proteins. Specialized protein complexes, such as CBC and eIF4F, bind to the cap to guide the RNA through the cell and ensure it reaches the ribosome safely. Enzymatic removal of the cap leads to the rapid degradation of RNA. This regulates RNA lifespan and maintains cellular balance. Viruses stay hidden and productive by encoding their own capping tools, snatching caps from host cells, or using proteins to mimic a cap’s function.
References
- Boivin, S., Cusack, S., Ruigrok, R. W. H., & Hart, D. J. (2010). Influenza A virus polymerase: Structural insights into replication and host adaptation mechanisms. In Journal of Biological Chemistry (Vol. 285, Number 37, pp. 28411–28417). American Society for Biochemistry and Molecular Biology Inc. https://doi.org/10.1074/jbc.R110.117531
- Cahová, H., Winz, M. L., Höfer, K., Nübel, G., & Jäschke, A. (2015). NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs. Nature, 519(7543), 374–377. https://doi.org/10.1038/nature14020
- Chen, Y., & Guo, D. (2016). Molecular mechanisms of coronavirus RNA capping and methylation. Virologica Sinica, 31(1), 3–11. https://doi.org/10.1007/s12250-016-3726-4
- Decroly, E., & Canard, B. (2017). Biochemical principles and inhibitors to interfere with viral capping pathways. In Current Opinion in Virology (Vol. 24, pp. 87–96). Elsevier B.V. https://doi.org/10.1016/j.coviro.2017.04.003
- Decroly, E., Ferron, F., Lescar, J., & Canard, B. (2012). Conventional and unconventional mechanisms for capping viral mRNA. In Nature Reviews Microbiology (Vol. 10, Number 1, pp. 51–65). https://doi.org/10.1038/nrmicro2675
- Doamekpor, S. K., Sharma, S., Kiledjian, M., & Tong, L. (2022). Recent insights into noncanonical 5′ capping and decapping of RNA. In Journal of Biological Chemistry (Vol. 298, Number 8). American Society for Biochemistry and Molecular Biology Inc. https://doi.org/10.1016/j.jbc.2022.102171
- Galloway, A., & Cowling, V. H. (2019). mRNA cap regulation in mammalian cell function and fate. In Biochimica et Biophysica Acta – Gene Regulatory Mechanisms (Vol. 1862, Number 3, pp. 270–279). Elsevier B.V. https://doi.org/10.1016/j.bbagrm.2018.09.011
- Jiao, X., Doamekpor, S. K., Bird, J. G., Nickels, B. E., Tong, L., Hart, R. P., & Kiledjian, M. (2017). 5′ End Nicotinamide Adenine Dinucleotide Cap in Human Cells Promotes RNA Decay through DXO-Mediated deNADding. Cell, 168(6), 1015-1027.e10. https://doi.org/10.1016/j.cell.2017.02.019
- Li, J., & Zhang, Y. (2012). Messenger RNA Cap Methylation in Vesicular Stomatitis Virus, a Prototype of Non‐Segmented Negative‐Sense RNA Virus. In Methylation – From DNA, RNA and Histones to Diseases and Treatment. InTech. https://doi.org/10.5772/54598
- Navas, L. E., & Carnero, A. (2021). NAD+ metabolism, stemness, the immune response, and cancer. In Signal Transduction and Targeted Therapy (Vol. 6, Number 1). Springer Nature. https://doi.org/10.1038/s41392-020-00354-w
- Park, G. J., Osinski, A., Hernandez, G., Eitson, J. L., Majumdar, A., Tonelli, M., Henzler-Wildman, K., Pawłowski, K., Chen, Z., Li, Y., Schoggins, J. W., & Tagliabracci, V. S. (2022). The mechanism of RNA capping by SARS-CoV-2. Nature, 609(7928), 793–800. https://doi.org/10.1038/s41586-022-05185-z
- Ramanathan, A., Robb, G. B., & Chan, S. H. (2016). mRNA capping: Biological functions and applications. In Nucleic Acids Research (Vol. 44, Number 16, pp. 7511–7526). Oxford University Press. https://doi.org/10.1093/nar/gkw551
- Sevajol, M., Subissi, L., Decroly, E., Canard, B., & Imbert, I. (2014). Insights into RNA synthesis, capping, and proofreading mechanisms of SARS-coronavirus. Virus Research, 194, 90–99. https://doi.org/10.1016/j.virusres.2014.10.008
- Teale, F. W. J. (2025). Flavin adenine dinucleotide (FAD) as a non-canonical RNA cap: Mechanisms, functions, and emerging insights. Archives of Biochemistry and Biophysics, 766(2), 110326. https://doi.org/10.1016/0968-0004(77)90259-6
- Wang, J., Alvin Chew, B. L., Lai, Y., Dong, H., Xu, L., Balamkundu, S., Cai, W. M., Cui, L., Liu, C. F., Fu, X. Y., Lin, Z., Shi, P. Y., Lu, T. K., Luo, D., Jaffrey, S. R., & Dedon, P. C. (2019). Quantifying the RNA cap epitranscriptome reveals novel caps in cellular and viral RNA. Nucleic Acids Research, 47(20). https://doi.org/10.1093/nar/gkz751
