Replication fork dynamics refers to the order, timing, and coordinated regulation of molecular processes that control the activity and movement of replication forks during chromosomal duplication.
It highlights the temporal and kinetic aspects of the process, such as: –
- The symmetry and speed of the fork progression.
- The rate of activation of the replication origin across the genome.
- The continuous replacement of damaged proteins with new ones during synthesis.
- The stalling of the fork during DNA damage.
The Architecture of the Replication Fork
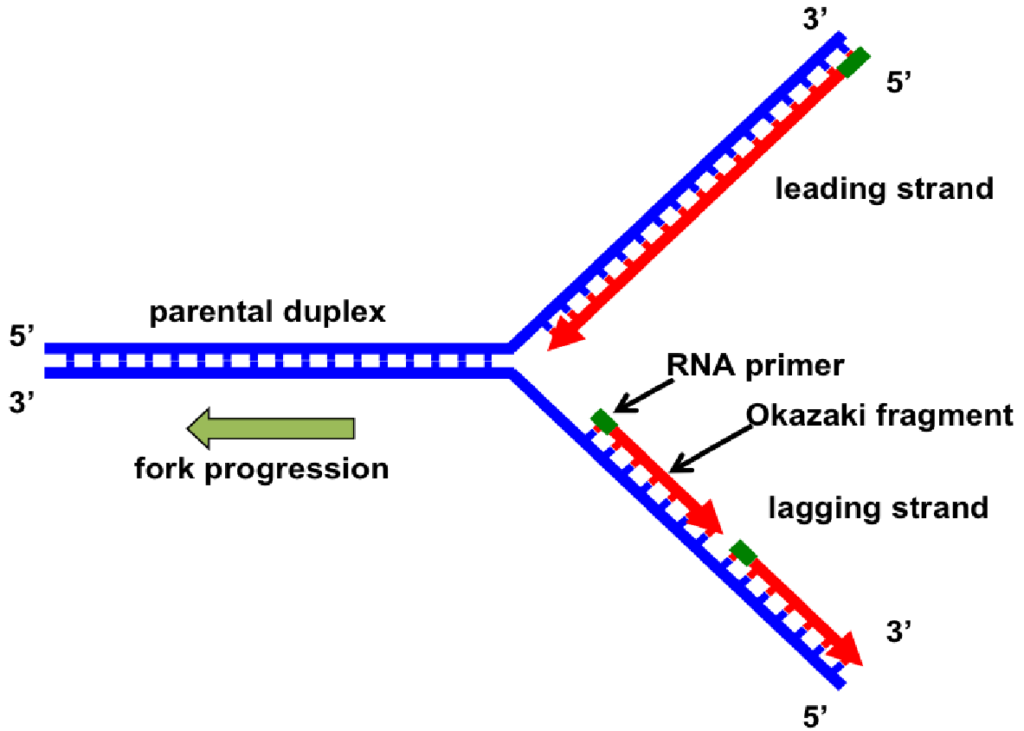
The architecture of the replication fork is a dynamic, Y-shaped complex structure in which the DNA double helix unwinds, and both strands are copied simultaneously through the coordination of various enzymes, such as helicase, primase, polymerases, ligase, single-stranded binding proteins, and topoisomerase.
Across all domains of life, the replication fork shows a conserved, functionally tiered architecture. However, the regulatory mechanisms and specific protein compositions involved in DNA synthesis are different.
- The central tier is formed by the hexameric DNA helicase that unwinds DNA and serves as a place to arrange the replication enzymes for new strand synthesis.
- The leading-strand DNA polymerase at the front tier continuously synthesizes the DNA in the fork’s direction.
- The lagging-strand DNA polymerases synthesize the short, discontinuous DNA segments, called the Okazaki fragments, at the back tier.
The replication fork’s architecture is defined by some of the following key characteristics: –
- Due to the antiparallel nature of DNA, the leading strand is synthesized continuously, while the lagging strand is synthesized discontinuously in short segments known as Okazaki fragments.
- The DNA replication must be completed even when the replication machinery comes across DNA damage and other barriers, causing the formation of small breaks or gaps in the newly synthesized strand, which are later repaired.
- For the coordinated, simultaneous working of the DNA polymerases on both strands, the lagging strand creates a loop. This is known as the “Trombone Model”.
- The tight coupling of helicase and leading-strand DNA polymerase ensures instant synthesis of the new strand as soon as the unwinding of the double helix takes place.
- Replisomes, such as polymerases and sliding clamps, are flexible structures that enable them to tackle the obstacles faced during the DNA replication process.
The Replisome Complex
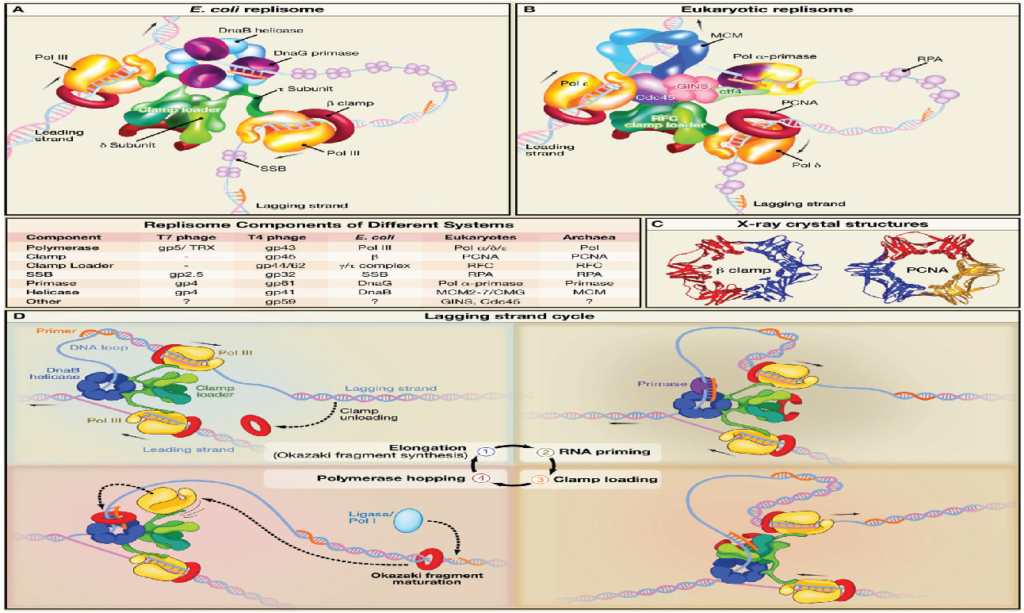
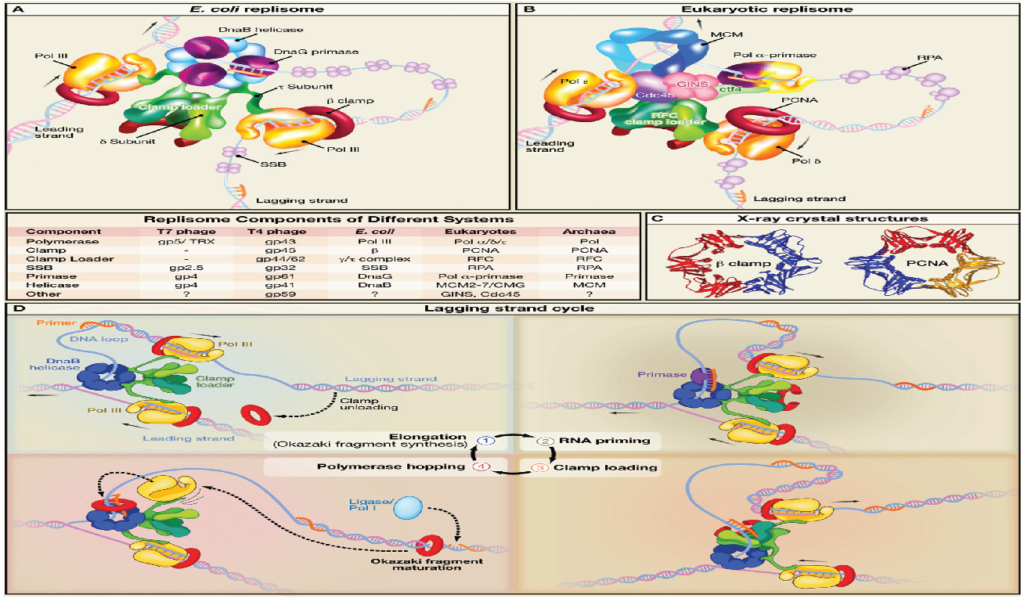
The replisome is a multiprotein molecular complex crucial for the precise and robust replication of DNA. It coordinates molecular motors and various specialized proteins responsible for unwinding the double helix of DNA and simultaneously synthesizing two new DNA strands.
The replisome’s core functional components that are conserved across all domains of life are as follows: –
- Replicative Helicase: It forms a hexameric ring that utilizes energy from the hydrolysis of ATP to move along DNA and unwind the double helix into two template strands.
- Topoisomerase/Gyrase: It prevents supercoiling of the DNA during unwinding by creating temporary cuts in the strands.
- Primase: It synthesizes short RNA primers on the single-stranded DNA templates to initiate DNA synthesis.
- DNA polymerase: It synthesizes the new DNA strands in the 5′ to 3′ direction.
- DNA Ligase: It functions by joining the synthesized Okazaki fragments on the lagging strand.
- Single-stranded DNA Binding Proteins (SSBs): They bind to the single-stranded DNA and stabilize it by avoiding re-annealing and the formation of secondary structure, and protect it from damage.
- Sliding Clamp: It is a ring-shaped protein that surrounds the DNA and tethers the polymerase to the template strand, thereby preventing the dissociation of the polymerase.
- Clamp Loader: It is an ATP-dependent, multiprotein complex that helps to open and position the sliding clamp onto DNA.
The operational mechanism of the replisome varies between eukaryotes and prokaryotes in the following ways:
| Components | Prokaryote (E. coli) | Eukaryote (S. cerevisiae) |
| Helicase | DnaB helicase is a homo-hexameric ring that moves in the 5′ to 3′ direction by encircling the lagging strand. | MCM helicase is a hetero-hexameric ring that is activated by recruiting Cdc45 and GINS, forming the active CMG complex. It moves in the 3′ to 5′ direction by encircling the leading strand. |
| Primase | DnaG primase synthesizes an RNA primer approximately 10-12 nucleotides long. | Pol α-primase complex creates an RNA-DNA hybrid primer. |
| Polymerase | A single Pol III complex synthesizes both strands of DNA. To increase replication efficiency, up to three Pol III can work simultaneously at one fork. | Specialized polymerases are employed for DNA synthesis. Pol α initiates new strand synthesis. Pol ϵ and Pol δ synthesize leading and lagging strands, respectively. |
| Sliding Clamp | The β clamp is a homodimer. | PCNA (Proliferating Cell Nuclear Antigen) is a homotrimer. |
| Clamp Loader | It is known as the γ/τ complex. | It is known as RFC (Replication Factor C). |
| SSB | EcoSSB functions as a homotetramer. | RPA (Replication Protein A) functions as a heterotrimer. |
Specialized factors of eukaryotes’ replisome: –
- The chromosomal structures are more complex in eukaryotes. Therefore, specialized factors such as CTf4 in the eukaryotic replisome connect Pol α–primase to the GINS complex, ensuring that primase activity is in sync with helicase activity.
- Phosphorylation of replisome proteins in eukaryotes helps to regulate the replication process in response to DNA damage and cell cycle checkpoints.
Okazaki Fragments
- Okazaki fragments are the short segments of DNA that are synthesized discontinuously on the lagging strand during DNA replication.
- The accumulation of DNA breaks as a result of incomplete Okazaki fragment processing causes neurodegenerative disorders and cancers in humans.
- To complete the replication, these fragments must undergo a pathway called Okazaki fragment maturation (OFM), where the primers are removed, the gaps are filled with DNA, and the fragments are joined by sealing the nicks.
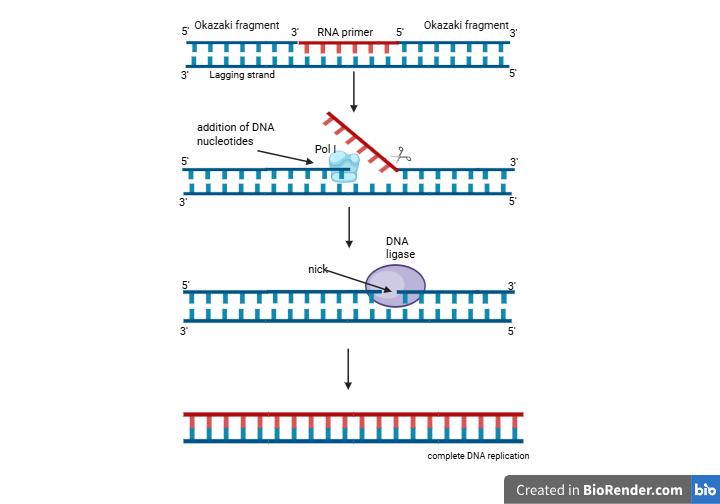
Okazaki Fragment Maturation in Prokaryote (e.g., E. coli): –
- Synthesis of Okazaki fragments, each starting with an RNA primer.
- Removal of RNA primer by the strand displacement activity of Pol I.
- 5′ flap endonuclease activity of Pol I cuts the 5′ flap structure formed by the displaced RNA primer.
- The gap is filled by Pol I’s DNA polymerase activity, and also ensuring accuracy by its 3′ to 5′ exonuclease proofreading activity.
- Pol I shifts the nick forward by replacing the RNA primer with the DNA. The process is known as ‘Nick translation’.
- DNA ligase seals the nick after the RNA primer is completely replaced by DNA.
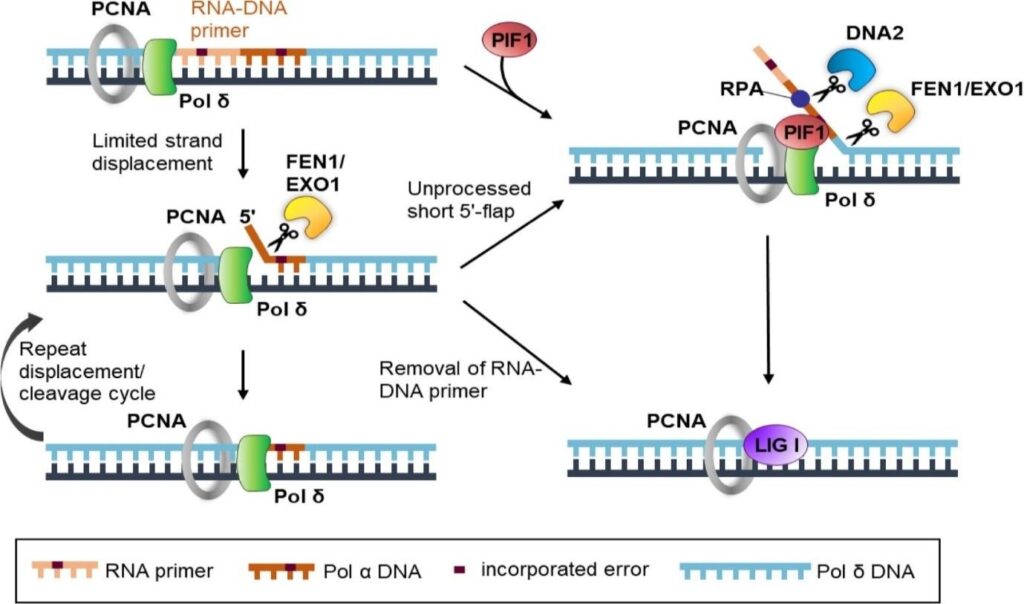
Okazaki Fragment Maturation in Eukaryotes (e.g., Yeast and Human): –
- The initial RNA-DNA primer is created by Pol α-primase.
- Pol δ performs strand displacement synthesis and nick translation.
- Nucleases such as Fen1 (known as Rad27 in yeast) cleave the short DNA flaps, whereas Dna2 cleaves the long DNA flaps into short ones for Fen1 to cleave them further.
- The final nick is sealed by DNA Ligase I (LIG1/Cdc9 in yeast).
- Accurate ligation is ensured by Mg2+ reinforced DNA-binding mode, employed by LIG1. Fen1 or mismatch repair (MMR) corrects the error if LIG1 fails to ligate accurately.
Single-Strand Binding Proteins (SSBs)
- These are the essential proteins that stabilize the single-stranded DNA (ssDNA) by binding to it.
- They prevent the formation of secondary structures.
- They protect the ssDNA from degradation.
- They coordinate other enzymes to the ssDNA during repair and replication.
- During homologous recombination, they help in the strand exchange process.
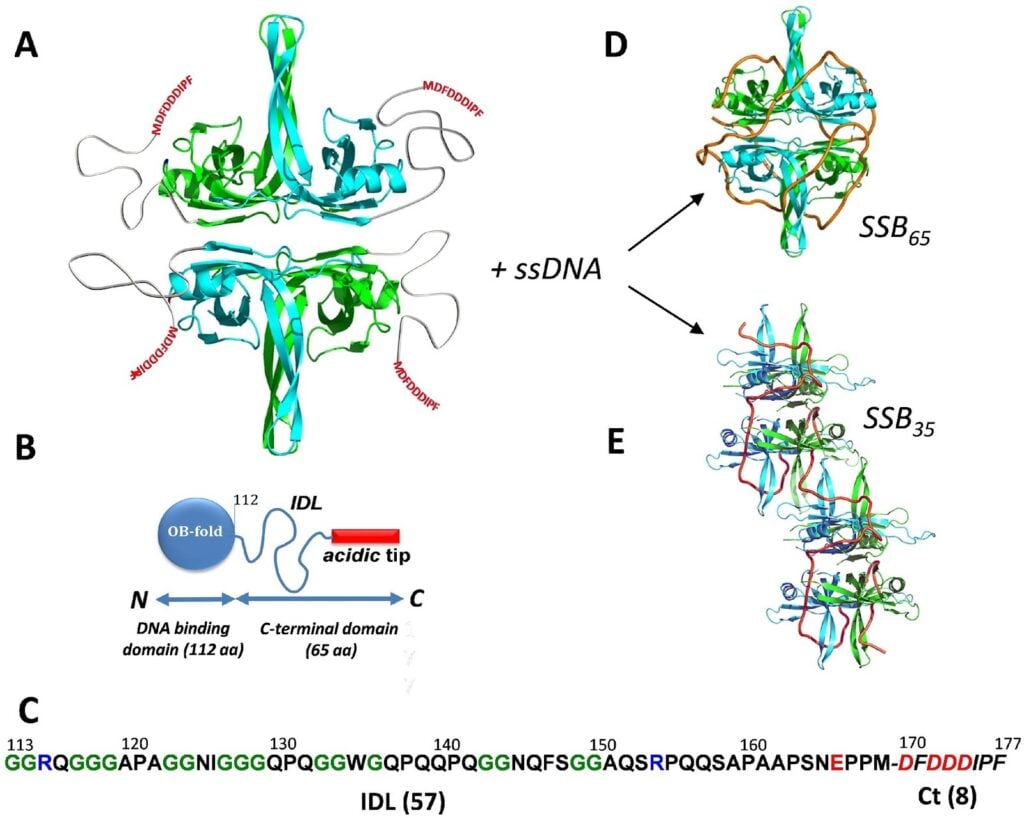
Key features of SSBs in Prokaryotes (E. coli):
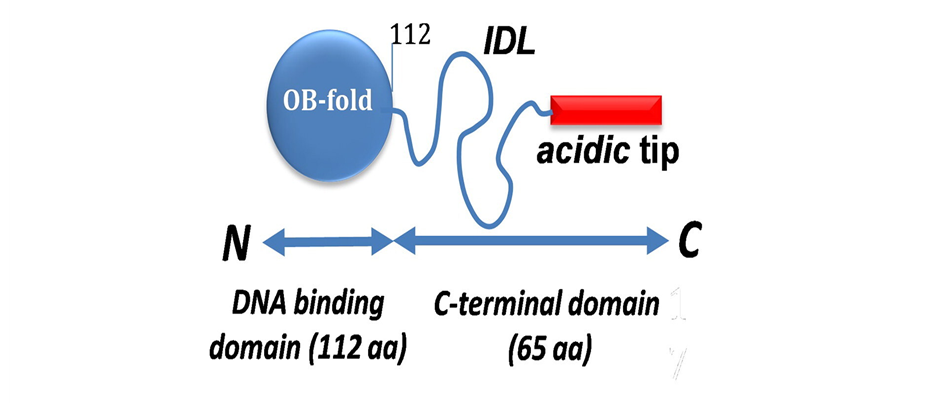
- SSBs function as a homotetramer, in which each monomer has an N-terminal Oligonucleotide Binding (OB) domain and a C-terminal tail.
- The N-terminal OB domain is responsible for DNA binding.
- C-terminal tail consists of a conserved 8-residue “acidic-tip” (SSB-Ct), and an intrinsically disordered linker (IDL).
- They can slide with a diffusion coefficient of ~270 nt2/s, along ssDNA.
- They are pushed unidirectionally along ssDNA by ATP-dependent motor proteins such as Rep and UvrD.
- Depending on binding density and salt concentration, ssDNA can be wrapped by SSB tetramers in various ways. At high salt (SSB), 65 mode is preferred, whereas at low salt (SSB), 35 mode is desired.
- SSBs suppress Illegitimate Recombination (IR) by working together with RecQ helicase.
Key features of SSBs in Eukaryotes:
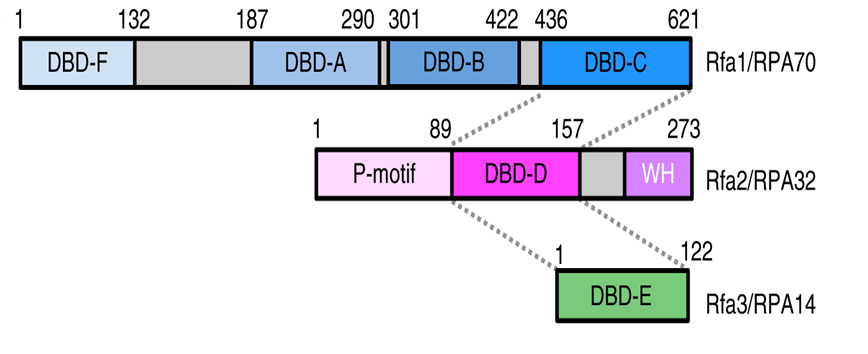
- Replication Protein A (RPA) is the functional analogue of bacterial SSB.
- It functions as a heterotrimer, consisting of RPA70, RPA32, and RPA14, as three different subunits.
- It prevents random base pairing (promiscuous annealing) by stabilizing ssDNA.
- It can slide with a diffusion coefficient of ~5000 nt2/s, along ssDNA.
- It can be pushed by Pif1 helicase along ssDNA.
- It suppresses microhomology-mediated end joining (MMEJ) and facilitates high-fidelity homologous recombination.
- Protein recruitment and post-translational modifications, such as phosphorylation of RPA32, control the regulation of RPA.
Replication Stress
- It is a phenomenon where the replication fork progression during DNA synthesis is stalled or slowed, disrupting accurate genome duplication.
- It is a leading cause of genomic instability, cellular ageing and senescence, cancer hallmarks, hereditary disorders, and stem cell exhaustion.
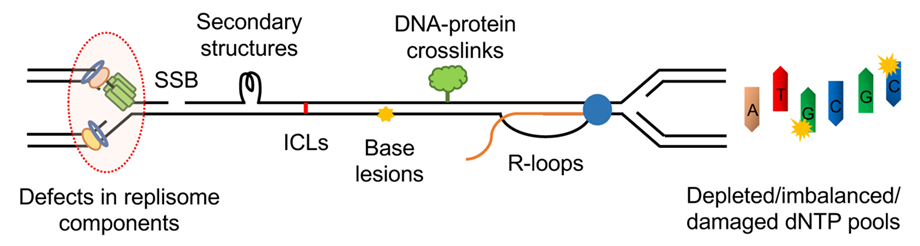
Sources of replication stress are: –
- Inter-strand cross-links (ICLs), DNA lesions from chemical mutagens or UV radiation, and secondary DNA structures, such as G-quadruplexes, cruciforms, and hairpins.
- Reduction of polymerase accuracy and efficiency due to uneven or low dNTP levels.
- Collisions between transcription and replication machinery are intensified by the R-loop formation.
- Naturally difficult regions to replicate, such as heterochromatin, fragile sites, and telomeres.
- Accelerated ageing and nuclear genome instability due to defects in mtDNA, which leads to a draining of cellular resources.
- Too many origins of replication drain replication resources, whereas too few origins of replication make the fork vulnerable.
Cellular response to replication stress: –
- In eukaryotes, ATR-mediated checkpoints halt the cell cycle for repair, stabilize the fork, and prevent excess origin firing.
- To bypass DNA damage, the fork is remodelled by reversing into four-way junctions (Chicken foot structure) to safeguard the DNA until the conflict resolves.
- The SOS response in bacteria, which is controlled by LexA and RecA, induces the enzymes required for DNA repair and allows continuous replication by bypassing DNA damage.
- A collapsed fork is restarted by Homologous Recombination (HR) and Break-Induced Replication.
- DNA lesions are bypassed by polymerases, which is allowed by Translesion Synthesis (TLS).
Transcription-Replication Conflicts
- It is the phenomenon that arises when the same DNA template is used to operate transcription as well as replication machinery.
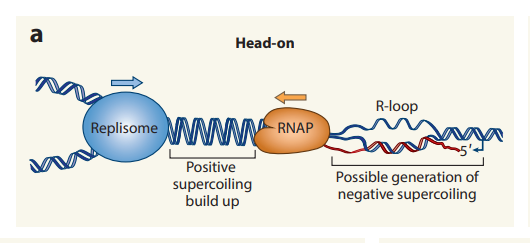
- This conflict can either occur Head-on (HO) or Codirectional (CD).
- The machinery moves towards each other in HO. This collision leads to instability, disassembly of the replisome, and halting of fork replication.
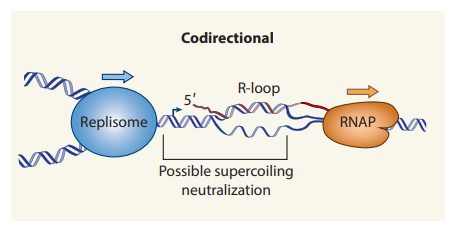
- Machinery moves in the same direction in CD, but it is still disruptive.
- CD encounters frequently occur in E. coli due to the faster movement of the replisomes than the RNA polymerase (RNAP).
- In eukaryotes, though both the machinery moves at the same speed, conflict is still raised due to high transcription density and the presence of long genes.
- Head-on conflicts in bacteria increase the rate of mutation, resulting in virulence and antibiotic resistance.
- The replication starts too often when the oncogenes like Myc and Cyclin E are overexpressed, leading to severe transcription-replication conflicts.
Mechanism of Transcription-Replication conflicts: –
- The DNA unwinding during both replication and transcription creates positive supercoiling. This supercoiling accumulates in the same region during Head-on conflicts, leading to fork instability and stalling.
- During transcription, R-loops are formed naturally and quickly resolved. Chromosomal rearrangements, fork stalling, and DNA breaks are caused when these R-loops persist longer and more frequently at HO conflict sites.
- During stress, RNAP slips backwards and gets immobilized on the DNA, preventing replication.
Resolution strategies and conflict avoidance: –
- The Head-on collision in bacteria is minimized through genome organization, by keeping highly transcribed genes codirectional with the replisome.
- In eukaryotes, the clashes are minimized by letting the cells replicate at specific times in highly transcribed regions or by organizing genes in separate nuclear domains.
- At DNA damage regions, proteins like CSB/Rad26 in eukaryotes, and Mfd in bacteria, remove the stalled RNAP.
- RNAP and R-loops are cleared by enzymes such as Rrm3, Pif1, and Senataxin (in eukaryotes), and Rep, UvrD, and PcrA (in bacteria).
- At conflict sites, positive supercoiling is resolved by topoisomerases I and II in eukaryotes, whereas DNA gyrase in bacteria resolves it.
Fork Collapse vs. Fork Stalling
Fork Stalling: –
- It is the temporary arrest of the replisome that occurs due to encountering physical barriers such as R-loops, RNAP, and Topoisomerase I cleavage complexes (Top1-ccs).
- In prokaryotes, the fork is slightly slowed by CD conflicts, whereas HO leads to extreme stalling of the fork.
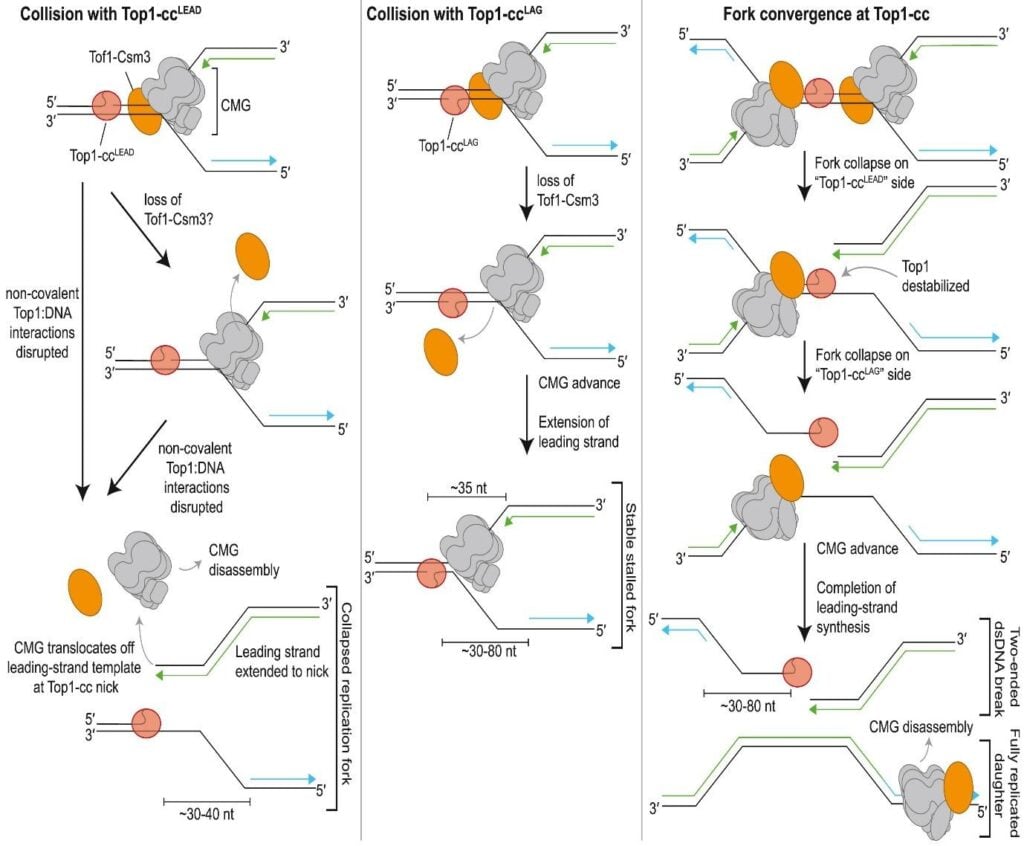
- In yeast, depending on whether the leading or the lagging strand encounters Top1-ccs, replication fork stalling varies.
- In both domains, accumulation of positive supercoiling during HO conflict is a primary reason for fork stalling.
- Oncogene-induced replication stress and breaks at Common Fragile Sites are caused by mismanaged stalling.
- For controlled stalling, the Tof1-Csm3 complex, which is known as “TIMELESS-TIPIN” in humans, is essential.
- In eukaryotes, collisions with Top1-ccs on the lagging strand result in stable stalled forks.
Fork Collapse: –
- It is the conversion of the stalled fork into double-strand breaks (DSBs) or the disassembly of the replisome.
- In eukaryotes, collision with Top1-ccs on the leading strand results in CMG helicase disassembly, replisome destabilization, and replication fork collapse.
- In the case of circular DNA, the two-ended DSB is formed when converging replication forks collide with a single Top1-ccs.
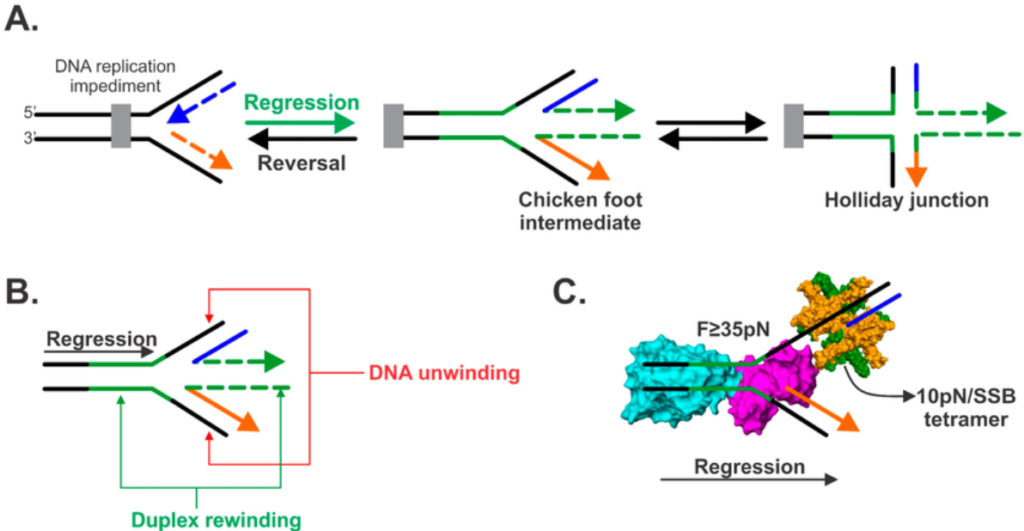
- A “Chicken Foot” intermediate is formed by reversal of the stalled forks to avoid fork collapse. RAD51 protects this structure from transcription-dependent damage in eukaryotes.
Rescue Mechanisms
This prevents lethal outcomes and preserves genome stability.
Rescue Mechanism in Prokaryotes (E. coli): –
- For Stalled forks:
- RecG helicase catalyzes fork regression and converts the stalled fork into a four-way Holliday junction (chicken foot intermediate).
- SSBs protect the ssDNA and recruit RecG to the stalled site.
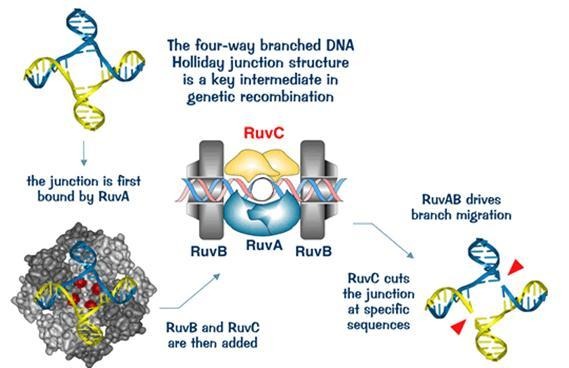
- RuvAB complex moves this Holliday junction along the DNA, and RuvC enzyme cleaves it at a specific point to restore the DNA structure and restart replication.
- For Collapsed forks:
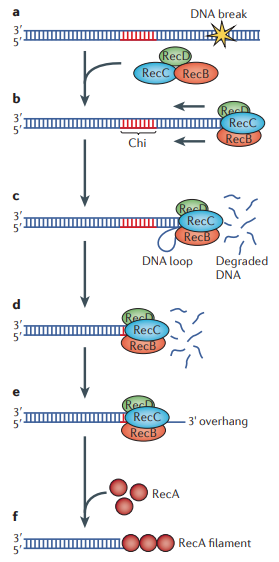
- RecBCD binds to the double-strand break and unwinds the DNA until the Chi-site is reached.
- RecA recombinase is then loaded onto the ssDNA by RecBCD, resulting in strand invasion for repair.
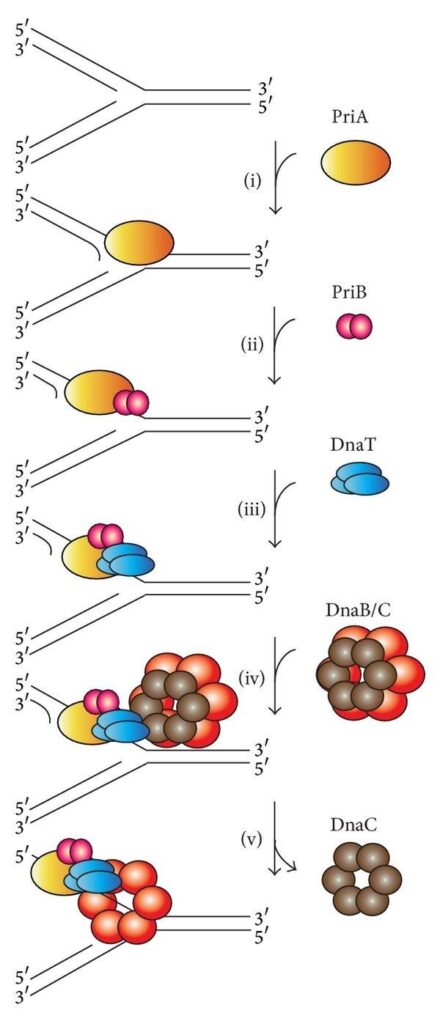
- Replication restart:
- Both stalled and collapsed fork rescue pathways create a substrate for the PriA helicase.
- PriA, along with PriB and DnaT, reloads the DnaB helicase onto the lagging strand to resume replication.
Rescue Mechanism in Eukaryotes: –
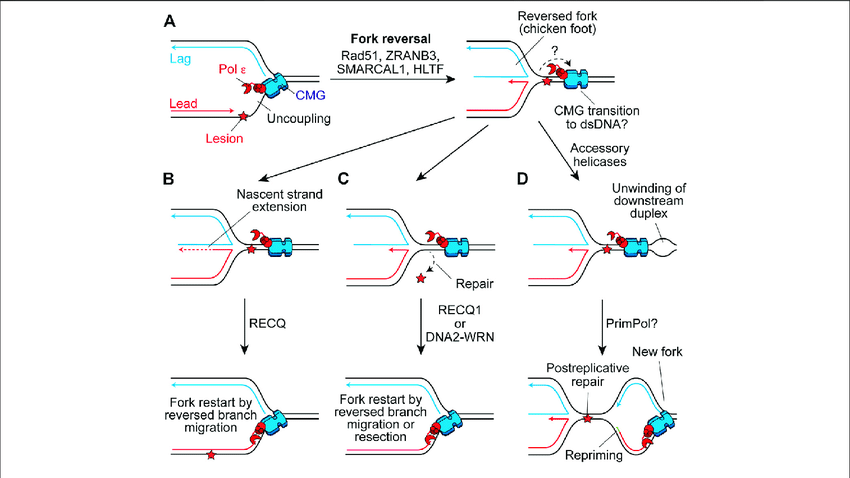
- In the initiation stage, SMARCAL1, ZRANB3, and HLTF, which are the SNF2-family enzymes, actively remodel stalled forks into four-way junctions (chicken foot).
- In the extension stage, PICH extends the reversed fork to complete and make it fully stable.
- RAD51 supports fork reversal by promoting strand annealing or stimulating translocase activity.
- Replication restart:
- After stress is cleared, RECQ1 helicase promotes fork restoration by reverse branch migration.
- Alternatively, to restart the fork, the regressed arm is partially degraded by DNA2 to create a specific substrate for RAD51-mediated strand invasion or translocases.
- Enzymes like PRIMPOL bypass the lesion and re-initiate the synthesis downstream when reversal is suppressed or impossible.
Checkpoint Activation
It is a cellular response that takes place when the DNA replication stress, such as protein roadblocks or template lesions, causes stalling of the replicative machinery.
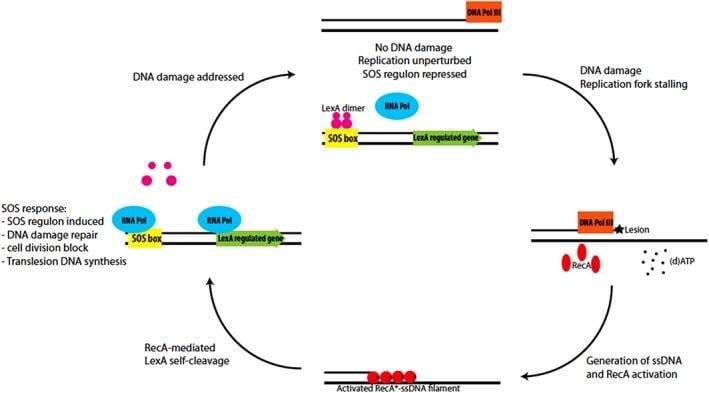
The SOS Response in Prokaryotes: –
- It is the primary checkpoint for replication stress in E. coli and B. subtilis, which is tightly regulated by RecA and LexA proteins.
- Accumulation of ssDNA due to stalling of the replication fork serves as a signal for checkpoint activation.
- An activated nucleoprotein filament (RecA*) is formed by the binding of RecA to the accumulated ssDNA in the presence of a nucleoside triphosphate.
- LexA represses expression of SOS genes by binding to the SOS box. RecA* triggers the self-cleavage of LexA, which leads to the induction of over 50 genes, including Translesion synthesis (TLS) polymerases and DNA repair.
- Cell division is halted by inhibitors, such as YneA (in B. subtilis) and SulA (in E. coli), to allow time for fork repair.
- DisA, an additional sensor in B. subtilis, detects branched DNA intermediates and suppresses c-di-AMP synthesis, leading to inhibition of cell proliferation for genome stability.
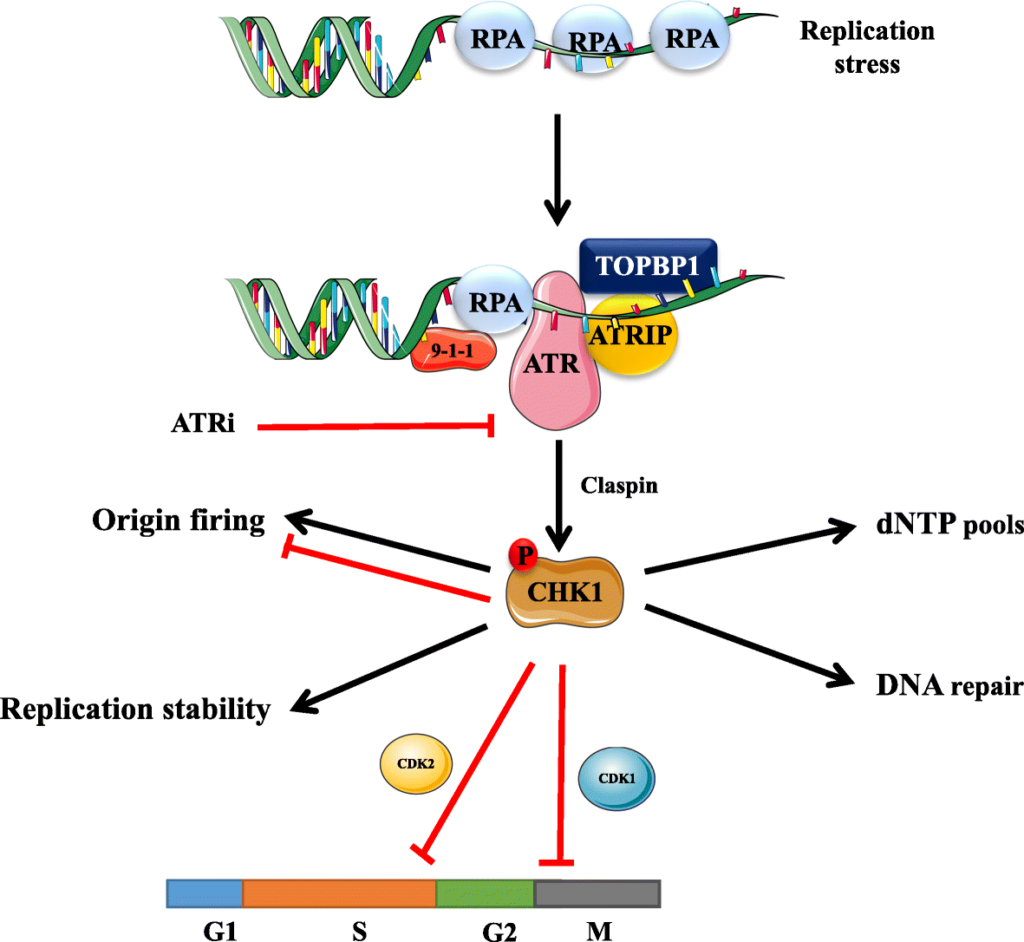
ATR-Chk1 Pathway in Eukaryotes: –
- The replication checkpoints in eukaryotes involve a cascade of ATR and Chk-1 kinases.
- Replication Protein A (RPA) coats the ssDNA generated by stalled forks, forming RPA-ssDNA complexes. ATR kinase is recruited to these sites via ATRIP.
- At the ssDNA-dsDNA junction, the 9-1-1 clamp (Rad9–Hus1–Rad1) is loaded by the Rad17-RFC complex.
- The 9-1-1 clamp recruits TopBP1, which stimulates the activity of ATR.
- The replisome protein And-1 (Ctf4 in yeast) is phosphorylated by ATR at residue T826. This facilitates the binding of Claspin to Chk1 and positions Chk1 at the stalled fork for its activation.
- Poly (ADP-ribose) (PAR) chains are synthesized by activated PARP1 when replication forks are damaged.
- The PbR motif (PAR-binding regulatory motif) helps PAR chains bind with Chk1. This holds Chk1 at the stalled fork, allowing ATR to phosphorylate and activate it.
- The activated Chk1 phosphorylates downstream targets for degradation. This leads to the cell cycle slowing and inhibition of new replication origin firing.
Clinical Relevance
Replication fork dynamics are of clinical importance in both prokaryotes and eukaryotes.
Clinical relevance in bacteria: –
- The antibiotics (like ciprofloxacin) that target DNA replication in Streptococcus pneumoniae cause fork stalling, which triggers the cell to uptake genes resistant to antibiotics from the environment, the phenomenon known as cell competence.
- The regulators that control competence are situated near OriC in many pathogens. When the replication is initiated continuously in the stalled fork, genes that regulate competence are copied many times. This is a “Gene Dosage Mechanism” activated by replication stress.
- To fight multidrug-resistant (MDR) pathogens, DNA ligase or sliding clamps are used as drug targets.
- Fork-induced competence acts as the general stress response in bacteria like S. pneumoniae, which lacks the SOS response.
Clinical relevance in Eukaryotes: –
- Genome instability is caused by replication stress, which is a key feature of cancer cells.
- Activation of oncogenes results in elevated replication stress in cancer cells. For survival, the cancer cells rely heavily on the replication stress response (RSR) pathways.
- By inhibiting kinases like ATR, Chk1, and WEE1, the stalled fork collapses and the cancer cell dies.
- The DNA fragments get accumulated in the cytoplasm when the replication stress response is blocked. The cGAS-STING signalling pathway is activated by these fragments, which enhances the efficacy of immune checkpoint inhibitors.
- ATR inhibition is highly sensitive in some cancers where ARID1A mutation or ATM deficiency is present. These cancer cells can be killed by blocking ATR, an approach for personalized cancer treatments.
Conclusions
Replication fork dynamics refers to the coordinated movement of the replication fork during DNA replication. The replisome complex, along with the stable structure of the fork, ensures processive and accurate DNA synthesis. Okazaki fragments must undergo the maturation process to complete lagging-strand replication. The single-stranded DNA (ssDNA) formed by the unwinding of the helicase is stabilized and protected from degradation with the help of the single-strand binding protein (SSBs or RPA in eukaryotes). Due to replication stress and transcription-replication conflicts, the fork can stall or collapse completely. These issues are resolved through checkpoint activation and the rescue mechanism. If these mechanisms fail, genomic instability arises, which leads to clinical consequences such as antibiotic resistance in bacteria and cancer induction in eukaryotes. Thus, replication fork biology serves as a bridge between fundamental science and medical practice.
References
- Adolph, M. B., & Cortez, D. (2024). Mechanisms and regulation of replication fork reversal. DNA Repair, 141. https://doi.org/10.1016/j.dnarep.2024.103731
- Bianco, P. R., & Lu, Y. (2021). Single-molecule insight into stalled replication fork rescue in Escherichia coli. Nucleic Acids Research, 49(8), 4220–4238. https://doi.org/10.1093/nar/gkab142
- Bocanegra, R., Ismael Plaza, G. A., Pulido, C. R., & Ibarra, B. (2021). DNA replication machinery: Insights from in vitro single-molecule approaches. In Computational and Structural Biotechnology Journal (Vol. 19, pp. 2057–2069). Elsevier B.V. https://doi.org/10.1016/j.csbj.2021.04.013
- Bonde, N. J., Kozlov, A. G., Cox, M. M., Lohman, T. M., & Keck, J. L. (2024a). Molecular insights into the prototypical single-stranded DNA-binding protein from E. coli. In Critical Reviews in Biochemistry and Molecular Biology (Vol. 59, Issues 1–2, pp. 99–127). Taylor and Francis Ltd. https://doi.org/10.1080/10409238.2024.2330372
- Bonde, N. J., Kozlov, A. G., Cox, M. M., Lohman, T. M., & Keck, J. L. (2024b). Molecular insights into the prototypical single-stranded DNA-binding protein from E. coli. In Critical Reviews in Biochemistry and Molecular Biology (Vol. 59, Issues 1–2, pp. 99–127). Taylor and Francis Ltd. https://doi.org/10.1080/10409238.2024.2330372
- Browning, K. R., & Merrikh, H. (2026). Replication-Transcription Conflicts: A Perpetual War on the Chromosome. Annual Review of Biochemistry Downloaded from Www.Annualreviews.Org. Guest, 58, 21. https://doi.org/10.1146/annurev-biochem-030222
- da Costa, A. A. B. A., Chowdhury, D., Shapiro, G. I., D’Andrea, A. D., & Konstantinopoulos, P. A. (2023). Targeting replication stress in cancer therapy. In Nature Reviews Drug Discovery (Vol. 22, Issue 1, pp. 38–58). Nature Research. https://doi.org/10.1038/s41573-022-00558-5
- Duckworth, A. T., Ducos, P. L., McMillan, S. D., Satyshur, K. A., Blumenthal, K. H., Deorio, H. R., Larson, J. A., Sandler, S. J., Grant, T., & Keck, J. L. (2023). Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli. Nature Communications, 14(1). https://doi.org/10.1038/s41467-023-38144-x
- Duderstadt, K. E., Reyes-Lamothe, R., van Oijen, A. M., & Sherratt, D. J. (2014). Replication-fork dynamics. Cold Spring Harbor Perspectives in Biology, 6(1). https://doi.org/10.1101/cshperspect.a010157
- Fajri, N., & Petryk, N. (2024). Monitoring and quantifying replication fork dynamics with high-throughput methods. In Communications Biology (Vol. 7, Issue 1). Nature Research. https://doi.org/10.1038/s42003-024-06412-1
- Feliciello, I., Ljubić, S., Đermić, E., Ivanković, S., Zahradka, D., & Đermić, D. (2024). Single-strand DNA-binding protein suppresses illegitimate recombination in Escherichia coli, acting in synergy with RecQ helicase. Scientific Reports, 14(1). https://doi.org/10.1038/s41598-024-70817-5
- Gao, Y., Cui, Y., Fox, T., Lin, S., Wang, H., De Val, N., Zhou, Z. H., & Yang, W. (2019). Structures and operating principles of the replisome. Science, 363(6429). https://doi.org/10.1126/science.aav7003
- Goehring, L., Huang, T. T., & Smith, D. J. (2023). Transcription-Replication Conflicts as a Source of Genome Instability. In Annual Review of Genetics (Vol. 57, pp. 157–179). Annual Reviews Inc. https://doi.org/10.1146/annurev-genet-080320-031523
- Guilliam, T. A. (2021). Mechanisms for Maintaining Eukaryotic Replisome Progression in the Presence of DNA Damage. In Frontiers in Molecular Biosciences (Vol. 8). Frontiers Media S.A. https://doi.org/10.3389/fmolb.2021.712971
- Herr, L. M., Schaffer, E. D., Fuchs, K. F., Datta, A., & Brosh, R. M. (2024). Replication stress as a driver of cellular senescence and aging. In Communications Biology (Vol. 7, Issue 1). Nature Research. https://doi.org/10.1038/s42003-024-06263-w
- Huang, Y. H., & Huang, C. Y. (2014). Structural insight into the DNA-binding mode of the primosomal proteins PriA, PriB, and AnaT. In BioMed Research International (Vol. 2014). Hindawi Publishing Corporation. https://doi.org/10.1155/2014/195162
- Kulczyk, A. W., Moeller, A., Meyer, P., Sliz, P., & Richardson, C. C. (2017). Cryo-EM structure of the replisome reveals multiple interactions coordinating DNA synthesis. Proceedings of the National Academy of Sciences of the United States of America, 114(10), E1848–E1856. https://doi.org/10.1073/pnas.1701252114
- Leman, A. R., & Noguchi, E. (2013). The replication fork: Understanding the eukaryotic replication machinery and the challenges to genome duplication. In Genes (Vol. 4, Issue 1, pp. 1–32). https://doi.org/10.3390/genes4010001
- Maslowska, K. H., Makiela-Dzbenska, K., & Fijalkowska, I. J. (2019). The SOS system: A complex and tightly regulated response to DNA damage. In Environmental and Molecular Mutagenesis (Vol. 60, Issue 4, pp. 368–384). John Wiley and Sons Inc. https://doi.org/10.1002/em.22267
- L., Zhang, J., He, K., & Zhang, J. (2019). Ataxia telangiectasia and Rad3-related inhibitors and cancer therapy: Where we stand. In Journal of Hematology and Oncology (Vol. 12, Issue 1). BioMed Central Ltd. https://doi.org/10.1186/s13045-019-0733-6
- Qiu, S., Jiang, G., Cao, L., & Huang, J. (2021). Replication Fork Reversal and Protection. In Frontiers in Cell and Developmental Biology (Vol. 9). Frontiers Media S.A. https://doi.org/10.3389/fcell.2021.670392
- Saxena, S., & Zou, L. (2022). Hallmarks of DNA replication stress. In Molecular Cell (Vol. 82, Issue 12, pp. 2298–2314). Cell Press. https://doi.org/10.1016/j.molcel.2022.05.004
- Shikazono, N., & Akamatsu, K. (2024). The role of DNA polymerase I in tolerating single-strand breaks generated at clustered DNA damage in Escherichia coli. Scientific Reports, 14(1). https://doi.org/10.1038/s41598-024-69823-4
- Slager, J., Kjos, M., Attaiech, L., & Veening, J. W. (2014). Antibiotic-induced replication stress triggers bacterial competence by increasing gene dosage near the origin. Cell, 157(2), 395–406. https://doi.org/10.1016/j.cell.2014.01.068
- Sun, H., Ma, L., Tsai, Y. F., Abeywardana, T., Shen, B., & Zheng, L. (2023). Okazaki fragment maturation: DNA flap dynamics for cell proliferation and survival. In Trends in Cell Biology (Vol. 33, Issue 3, pp. 221–234). Elsevier Ltd. https://doi.org/10.1016/j.tcb.2022.06.014
- Taylor, M. R. G., & Yeeles, J. T. P. (2019). Dynamics of Replication Fork Progression Following Helicase–Polymerase Uncoupling in Eukaryotes. Journal of Molecular Biology, 431(10), 2040–2049. https://doi.org/10.1016/j.jmb.2019.03.011
- van Eijk, E., Wittekoek, B., Kuijper, E. J., & Smits, W. K. (2017). DNA replication proteins as potential targets for antimicrobials in drug-resistant bacterial pathogens. Journal of Antimicrobial Chemotherapy, 72(5), 1275–1284. https://doi.org/10.1093/jac/dkw548
- Westhorpe, R., Roske, J. J., & Yeeles, J. T. P. (2024). Mechanisms controlling replication fork stalling and collapse at topoisomerase 1 cleavage complexes. Molecular Cell, 84(18), 3469-3481.e7. https://doi.org/10.1016/j.molcel.2024.08.004
- Wigley, D. B. (2013). Bacterial DNA repair: Recent insights into the mechanism of RecBCD, AddAB and AdnAB. Nature Reviews Microbiology, 11(1), 9–13. https://doi.org/10.1038/nrmicro2917
- Williams, J. S., Tumbale, P. P., Arana, M. E., Rana, J. A., Williams, R. S., & Kunkel, T. A. (2021). High-fidelity DNA ligation enforces accurate Okazaki fragment maturation during DNA replication. Nature Communications, 12(1). https://doi.org/10.1038/s41467-020-20800-1
- Yang, W., Seidman, M. M., Rupp, W. D., & Gao, Y. (2019). Replisome structure suggests mechanism for continuous fork progression and post-replication repair. In DNA Repair (Vol. 81). Elsevier B.V. https://doi.org/10.1016/j.dnarep.2019.102658
- Yao, N. Y., & O’Donnell, M. (2010). SnapShot: The Replisome. In Cell (Vol. 141, Issue 6, pp. 1088-1088.e1). Elsevier B.V. https://doi.org/10.1016/j.cell.2010.05.042
- Yates, L. A., Aramayo, R. J., Pokhrel, N., Caldwell, C. C., Kaplan, J. A., Perera, R. L., Spies, M., Antony, E., & Zhang, X. (2018). A structural and dynamic model for the assembly of Replication Protein A on single-stranded DNA. Nature Communications, 9(1). https://doi.org/10.1038/s41467-018-07883-7
- Zhang, Y., Wu, L., Wang, Z., Wang, J., Roychoudhury, S., Tomasik, B., Wu, G., Wang, G., Rao, X., & Zhou, R. (2022). Replication Stress: A Review of Novel Targets to Enhance Radiosensitivity-From Bench to Clinic. In Frontiers in Oncology (Vol. 12). Frontiers Media S.A. https://doi.org/10.3389/fonc.2022.838637
