Polyadenylation is a ubiquitous post-transcriptional modification, where a sequence of untemplated adenosine residues is added to the 3′ end of an RNA molecule.
While common to both prokaryotes and eukaryotes, its mechanisms and biological roles differ significantly between the two. In eukaryotes, it stabilizes mRNA, promotes its export from the nucleus, and enhances translation efficiency.
The length of the poly(A) tail is long and relatively stable, ranging from 200 to 250 nucleotides. Whereas, in prokaryotes, it serves as a signal for RNA degradation and quality control. Bacterial poly(A) tails are short and unstable, typically shorter than 20 nucleotides.
The Polyadenylation Signal
The polyadenylation signal (PAS) is a critical sequence motif that defines where an RNA molecule will be cleaved and modified with a poly(A) tail. Its characteristics vary significantly between domains of life.
Eukaryotic Polyadenylation Signal
- The most common, highly conserved PAS is a hexanucleotide sequence AAUAAA.
- The second most frequent signal is AUUAAA. Other single-nucleotide variants exist but are less common and often lead to alternative polyadenylation.
- This hexamer is typically found 20 to 30 nucleotides upstream of the actual cleavage site.
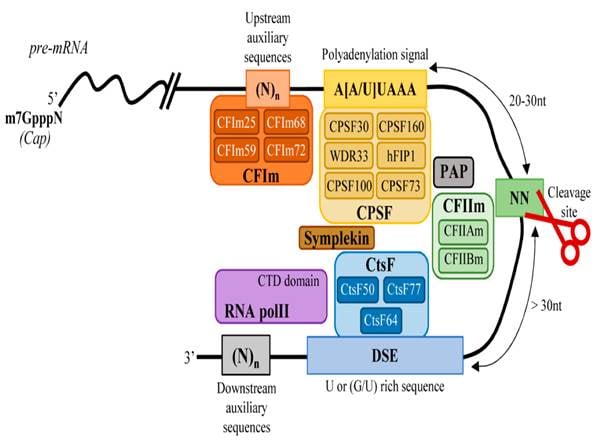
- The CPSF complex (Cleavage and Polyadenylation Specificity Factor) recognizes this signal, with the WDR33 and CPSF30 subunits directly binding the hexamer.
- Recognition of PAS is often supported by other elements such as Cleavage Factor I (CFIm) and Cleavage Stimulation Factor (CstF).
- CFIm recognizes UGUA motifs present in the Upstream Element (USE).
- Cleavage Site is usually a CA dinucleotide.
- CstF recognizes a GU- or U-rich region present in the Downstream Sequence Element (DSE).
Prokaryotic Polyadenylation Signal
- Bacteria, such as E. coli, lack a universal consensus sequence equivalent to the eukaryotic AAUAAA.
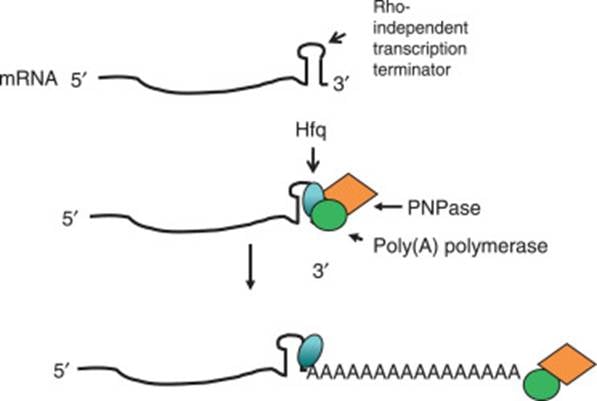
- Rho-independent transcription terminators (stable stem-loop structures) are preferred targets for polyadenylation by Poly(A) Polymerase I (PAP I).
- PAP I requires an accessible single-stranded 3′ end to work. If the terminators lack a short extension, PAP I is inhibited.
- Hfq is a hexameric RNA-binding protein that binds the stem-loop structure and unwinds it. This lets PAP I to initiate the addition of A residues.
- Bacterial polyadenylation is relatively non-specific. The 3′ ends of primary transcripts, processed RNAs, or decay intermediates can be polyadenylated by PAP I if they are accessible.
Key Protein Factors for Polyadenylation
Protein factors in polyadenylation differ significantly between eukaryotes and prokaryotes. Eukaryotes employ a large, multi-complex machinery, whereas prokaryotes use a smaller set of enzymes primarily focused on RNA turnover.
Eukaryotic Key Protein Factors
Polyadenylation machinery, known as the CPA machinery, consists of a core complex and dozens of associated factors. The following protein factors are given below:
- CPSF (Cleavage and Polyadenylation Specificity Factor): This central complex is responsible for recognizing the polyadenylation signal “AAUAAA” and defining the cleavage site.
-mPSF (Mammalian PSF): A core subcomplex within CPSF consisting of CPSF160, CPSF30, hFip1, and WDR33.
-WDR33 and CPSF30: The specific subunits that directly bind to the AAUAAA signal.
-CPSF73: The endonuclease that physically cuts the pre-mRNA transcript.
-hFip1: Directly recruits poly(A) polymerase to the processing site.
- CstF (Cleavage Stimulation Factor): It recognizes the GU- or U-rich downstream element (DSE) and interacts with CPSF to stabilize the machinery.
- CFIm and CFIIm (Cleavage Factors I and II): These are auxiliary complexes that bind sequence elements, such as UGUA, to help define the poly(A) site, especially for alternative polyadenylation.
- Poly(A) Polymerase (PAP): This enzyme catalyzes the non-templated addition of adenosine residues.
- PABPN1 (Nuclear Poly(A)-Binding Protein 1): It binds the growing tail to stimulate the polymerase and defines the appropriate tail length, i.e., ~250 nucleotides.
- Symplekin: It acts as a scaffold protein that physically connects the various sub-complexes, like CPSF and CstF.
Prokaryotic Key Protein Factors
Bacteria, particularly E. coli, employ a simpler system where polyadenylation functions as a signal for RNA decay rather than stabilization. The main protein factors are as follows:
- Poly(A) Polymerase I (PAP I): It is encoded by the pcnB gene. This primary enzyme is responsible for adding homopolymeric (all-adenosine) tails in bacteria.
- Polynucleotide Phosphorylase (PNPase): It is a dual-function enzyme. While primarily a 3′-5′ exonuclease that degrades RNA, it can also act as a polymerase to add heteropolymeric tails (containing all four nucleotides but rich in A).
- Hfq: This is an abundant RNA-binding protein and chaperone that facilitates polyadenylation by converting PAP I from a distributive to a processive enzyme and protecting long poly(A) tails from rapid degradation.
- Exoribonucleases (RNase II, RNase R, and PNPase): These enzymes counteract PAP I by removing poly(A) tails. However, poly(A) tails also allow these exonucleases to bind and degrade highly structured RNA fragments.
Mechanism of Reaction in Polyadenylation
It involves the non-templated addition of nucleotides to the 3′ end of an RNA molecule. The reaction process and biological outcome differ significantly between eukaryotes and prokaryotes.
Eukaryotic Mechanism
In eukaryotes, polyadenylation is a highly specific, two-step nuclear process coupled with transcription termination.
- Recognition: The CPSF complex, specifically subunits WDR33 and CPSF30, binds to the AAUAAA polyadenylation signal (PAS) on the pre-mRNA. Simultaneously, the CstF complex binds to a GU- or U-rich downstream sequence element (DSE).
- Cleavage: The CPSF73 subunit cleaves the pre-mRNA transcript at a specific cleavage site (usually a CA dinucleotide) located between the PAS and the DSE.
- Poly(A) Addition: Poly(A) polymerase (PAP) is recruited to the new 3′ end and catalyzes the addition of 200–250 adenosine residues using ATP as a substrate.
- Length Control: PABPN1 (nuclear poly(A)-binding protein 1) binds to the growing tail, stimulating PAP for rapid elongation and defines the appropriate tail length.
Prokaryotic Mechanism
In prokaryotes, primarily in E. coli, polyadenylation is generally less specific and serves as a signal for RNA decay.
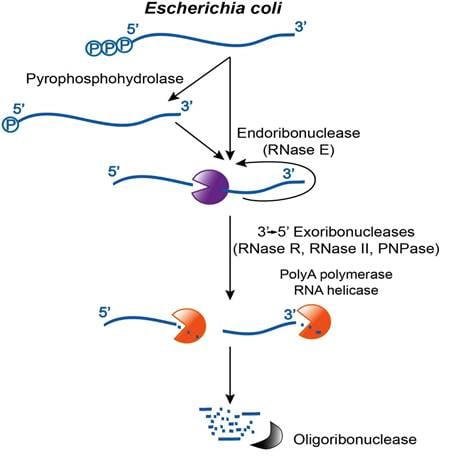
- Substrate Targeting: Prokaryotes lack a universal consensus signal. Instead, PAP I targets any accessible single-stranded 3′ end. Preferred targets include Rho-independent terminators (stem-loops) and fragments already partially processed by endonucleases like RNase E.
- Tailing Reaction:
-Homopolymeric tails: PAP I uses ATP to polymerize a sequence of adenosine residues onto the RNA primer.
-Heteropolymeric tails: Polynucleotide phosphorylase (PNPase) can add mixed tails, containing all four nucleotides (though often A-rich), when it stalls during its usual degradative function.
- Processivity: PAP I is inherently a distributive enzyme that adds only a few nucleotides before detaching. However, the Hfq protein can bind the tail and transform PAP I into a more processive enzyme to extend the tail.
- Coupling with Decay: The reaction provides a single-stranded extension for 3′-5′ exoribonucleases like PNPase or RNase R to bind and degrade highly structured RNA fragments.
Biological Functions of Polyadenylation
Polyadenylation controls the functionality and fate of RNA in all domains of life.
In Eukaryotes
- mRNA Stability: The poly(A) tail protects mature mRNA from non-specific degradation in the cytoplasm.
- Nuclear Export: It is critical for the transport of mRNAs from the nucleus to the cytoplasm.
- Translation Efficiency: The tail enhances translation by facilitating the formation of the closed-loop complex, which recruits ribosomes.
- Gene Regulation (APA): Alternative Polyadenylation (APA) produces different mRNA isoforms from the same gene. This can change the coding sequence to create different protein isoforms or alter the 3′ UTR to escape regulation by microRNAs or RNA-binding proteins.
- Developmental Control: In oocytes and embryos, cytoplasmic polyadenylation unmasks dormant mRNAs, allowing them to be translated at specific stages of development.
In Prokaryotes
- RNA Degradation: It promotes the rapid decay of mRNA fragments and other transcripts.
- Exonuclease “Toeholds”: Poly(A) tails provide a single-stranded extension, called a toehold, that allows exonucleases like PNPase to bind and degrade tightly folded structured RNAs that are otherwise resistant to decay.
- Quality Control: It targets abnormally folded or defective RNAs, such as non-functional tRNAs or rRNAs, for recycling.
- Regulation of Small RNAs: Polyadenylation controls the intracellular concentration of small regulatory RNAs (sRNAs) and antisense RNAs.
- Metabolic Signaling: It helps regulate the levels of essential decay enzymes, like RNase E and PNPase, and manages toxin-antitoxin systems and plasmid copy numbers.
Alternative Polyadenylation
Alternative polyadenylation (APA) is a mechanism where a single gene utilizes multiple polyadenylation sites (PAS) to produce distinct mRNA isoforms.
In Eukaryotes
- It is a post-transcriptional gene regulatory process that affects most of the protein-coding genes.
- The types of APA are: –
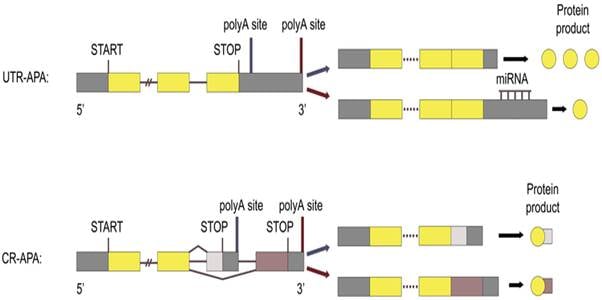
- 3′ UTR APA: It is the most common type of APA. Here, different PASs are chosen within the same terminal exon. This changes the length of the 3′ untranslated region (UTR) without altering the protein sequence.
- Coding Region APA (CR-APA): mRNA gets polyadenylated within an internal intron or exon, which produces protein isoforms with different C-termini.
- Especially in proliferating or cancer cells, increased mRNA stability and higher protein expression can result from the shortening of the 3′ UTR, as this reduces binding sites for microRNAs and RNA-binding proteins.
- Depending on the specific tissue or cell type, the same gene can produce different mRNA isoforms, resulting in different 3′ UTR lengths. For instance, the nervous system preferentially uses longer PASs, whereas proliferating and cancer cells often switch to shorter PASs to evade regulation. Therefore, APA is highly cell-type and tissue-specific.
- The choice of PAS is determined by the concentration of core factors like CstF64 and CFIm25, and the speed of RNA polymerase II during transcription.
In Prokaryotes
- Prokaryotic transcripts are also polyadenylated at multiple sites.
- Unlike eukaryotes, prokaryotes lack a universal consensus signal like AAUAAA. Instead, polyadenylation can occur at almost any accessible 3′ end.
- Tailing is detected at transcription terminators, intergenic regions, and at internal endoribonucleolytic cleavage sites, which are generated by enzymes like RNase E.
- Polyadenylation at these various sites in bacteria serves as a signal for rapid RNA degradation and quality control.
- The equilibrium between PAP I adding tails and exoribonucleases removing them determines which fragments are targeted for complete decay.
Regulation of Efficiency in Polyadenylation
It is achieved through the complex interplay of cis-acting sequence elements and the concentration and activity of trans-acting protein factors.
Regulation in Eukaryotes
- The efficiency of polyadenylation is primarily determined by the strength of core signals, such as the AAUAAA hexamer, and auxiliary elements like UGUA or GU-rich regions.
- The abundance of core processing factors like CstF64 or CFIm determines site selection. For instance, high factor levels often promote the use of weaker proximal polyadenylation sites.
- The speed of RNA Polymerase II directly impacts efficiency. Slower elongation or pausing provides more time for the machinery to recognize and process proximal or weak signals.
- Regulatory proteins can either compete with basal factors to block processing or recruit the polyadenylation machinery to specific sites through direct protein-protein interactions.
- Modifications such as phosphorylation and sumoylation of factors like PAP can increase or decrease their catalytic activity.
Regulation in Prokaryotes
- Efficiency is tightly controlled by the low abundance of PAP I, with only about 32–50 molecules typically present in an E. coli cell to prevent toxic over-polyadenylation.
- The net efficiency of tailing is an equilibrium between the synthesis rate of PAP I and the tail removal rate by exoribonucleases like RNase II and PNPase.
- The RNA chaperone Hfq enhances efficiency by binding to the RNA, converting PAP I from a distributive to a processive enzyme, and protecting the growing tails from degradation.
- Polyadenylation efficiency is inversely related to growth rate, with PAP I levels and overall RNA tailing increasing significantly in slowly growing bacteria.
- The presence of Rho-independent terminators serves as a signal that can influence how efficiently PAP I targets and tails specific transcripts.
Deadenylation and Decay
Deadenylation is the enzymatic shortening or removal of the poly(A) tail, which serves as a critical control point for RNA stability and decay in all domains of life.
Eukaryotic Deadenylation and Decay
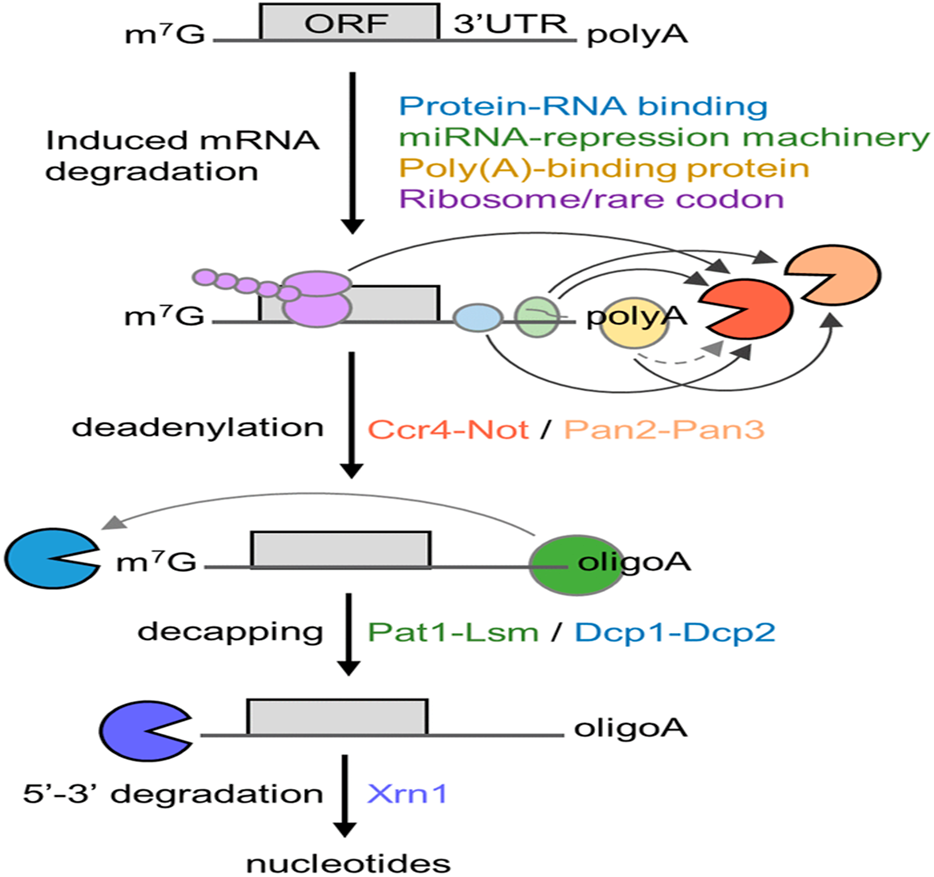
- Deadenylation is typically the initial and rate-limiting step in the 5′-3′ mRNA degradation pathway.
- This task is performed by Pan2-Pan3 and Ccr4-Not, which are the two major complexes.
- These complexes are recruited to specific mRNAs through signals in the 3′ UTR, by the miRNA repression system involving Ago and GW182, or through general interactions with the cytoplasmic poly(A)-binding protein (PABPC1).
- The Lsm1-7/Pat1 complex binds once the tail is shortened to about 10–15 nucleotides, leading to 5′ cap removal by the Dcp1-Dcp2 decapping complex.
- Decapping exposes the 5′ end to Xrn1, which is an exoribonuclease that carries out the final 5′-3′ degradation of the mRNA body into nucleotides.
Prokaryotic Deadenylation and Decay
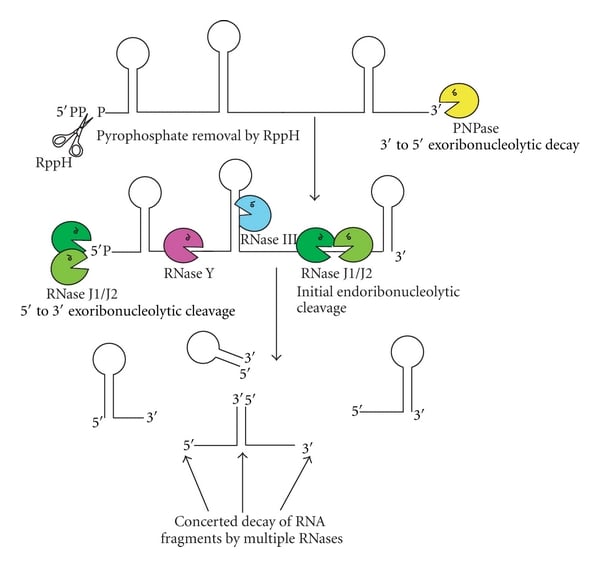
- In bacteria, polyadenylation usually acts as a signal for rapid RNA turnover and quality control.
- The length and abundance of tails represent a balance between the synthetic activity of PAP I and the degradative activity of exoribonucleases like PNPase and RNases.
- Poly(A) tails provide unstructured, single-stranded extensions, called toeholds, that allow enzymes like PNPase and RNases to bind and degrade highly structured RNAs, such as stem-loops, that they otherwise could not digest.
- The addition and subsequent removal of tails predominantly promote mRNA degradation in mitochondria and chloroplasts as well.
Cytoplasmic Polyadenylation
Cytoplasmic polyadenylation is the enzymatic elongation of a poly(A) tail on an mRNA transcript after it has been exported from the nucleus to the cytoplasm. It is a critical regulator of gene expression, particularly in development and neuronal function.
- It translationally activates dormant mRNAs that were exported with short poly(A) tails.
- It is essential for oocyte maturation, early embryonic development, cell cycle progression, and synaptic plasticity.
- Key sequence elements:
-Cytoplasmic Polyadenylation Element (CPE): It is a U-rich motif with a consensus sequence “UUUUUAU” in the 3′ UTR that determines which mRNAs should undergo cytoplasmic polyadenylation.
-Poly(A) Signal (PAS): It is the hexanucleotide “AAUAAA” which is also found in nuclear polyadenylation. In cytoplasmic polyadenylation, it works with CPE to promote proper tail extension and regulate mRNA activity.
- Key Protein Factors:
-CPE Binding Protein (CPEB): It is the central sequence-specific factor that binds CPE and promotes tail elongation.
-Cleavage and Polyadenylation Specificity Factor (CPSF): It is a multi-subunit complex that recognizes PAS in mRNAs. In addition to the endonuclease CPSF73, three cytoplasmic CPSF subunits bind the PAS and facilitate the process.
-Gld2 (PAPD4) and Gld4 (PAPD5): These non-canonical poly(A) polymerases catalyze the addition of adenosine residues.
-Symplekin: It acts as a scaffold for the assembly of the cytoplasmic complex.
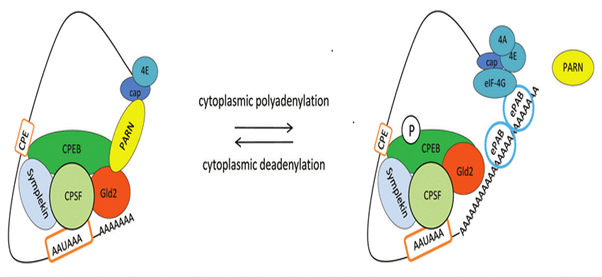
- Mechanism Example: In a repressed state, CPEB recruits deadenylases like PARN to keep the tail short. Upon signaling (e.g., hormone stimulation), CPEB is phosphorylated, PARN is expelled, and Gld2 elongates the tail, which triggers translation.
Clinical Significance of Polyadenylation
Defects and dysregulation in polyadenylation are directly linked to various diseases, ranging from genetic blood disorders to cancer and viral infections.
Clinical Significance in Eukaryotes
- Widespread shortening of 3′ UTRs through alternative polyadenylation (APA) is a hallmark of cancer. By removing miRNA binding sites, these shortened transcripts activate oncogenes and help cells evade normal regulatory controls.
- Specific mutations in the polyadenylation signal (AAUAAA) cause serious diseases, such as Alpha-thalassaemia (hemoglobin deficiency) and hereditary thrombophilia (excessive blood clotting).
- Mutations in PABPN1 cause Oculopharyngeal Muscular Dystrophy (OPMD) by disrupting proper tail length control.
- Dysregulation of cytoplasmic polyadenylation is linked to Fragile X syndrome, Alzheimer’s disease, Huntington’s disease, and autism.
- Viruses, like Influenza, use the NS1 protein to block host polyadenylation factors such as CPSF and PABP, preventing the host from producing mature antiviral mRNAs while allowing viral replication.
- APA profiles are being explored as biomarkers for drug sensitivity (e.g., to HDAC inhibitors) and as targets for new RNA-based therapies.
Clinical Significance in Prokaryotes
- In E. coli, polyadenylation regulates the FliA regulon, which controls motility and cell adhesion. These factors are essential for establishing infections in hosts.
- Polyadenylation manages the levels of antitoxin RNAs, such as Sok and CopA, which determine bacterial survival and the stability of antibiotic-resistance plasmids.
- PAP I levels increase during slow growth or stress conditions, potentially helping pathogens survive adverse environments inside a host.
- Inhibiting the polyadenylation endonuclease (CPSF3) in pathogens like Cryptosporidium has been shown to effectively control infection, marking it as a promising drug target.
Conclusions
Polyadenylation involves the enzymatic addition of untemplated adenosine residues to the 3′ terminus of RNA. This promotes transcript stability and maturation in eukaryotes while primarily serving as a signal for exonucleolytic degradation in prokaryotes.
In mammalian systems, this processing is directed by highly conserved sequence motifs like AAUAAA hexamer, whereas bacterial enzymes typically target accessible, single-stranded 3′ ends without requiring a universal consensus signal. The reaction is carried out by complex protein machineries, including the CPSF and associated factors in eukaryotes, whereas in bacteria, the simpler PAP I and PNPase enzymatic systems are utilized.
Mechanistically, eukaryotic nuclear processing requires coordinated endonucleolytic cleavage before tail extension, while prokaryotic tailing often provides critical toeholds for the breakdown of otherwise resistant structured RNA fragments. These modifications are indispensable for eukaryotic mRNA nuclear export and translational efficiency, yet in prokaryotes, they function as vital regulators of RNA turnover and metabolic quality control. Through alternative polyadenylation, eukaryotic genes further expand their regulatory potential by producing distinct mRNA isoforms, while bacteria add tails to multiple internal sites to speed up decay.
The efficiency of these reactions is modulated by the cellular abundance of trans-acting factors and the strength of transcript-specific cis-elements, with deadenylation governing the rate-limiting step of decay in eukaryotes.
Consequently, dysregulation within these polyadenylation pathways leads to various human pathologies, including oncogenesis and genetic blood disorders, as well as bacterial stress adaptation and pathogenesis.
References
- Bandyra, K. J., Bouvier, M., Carpousis, A. J., & Luisi, B. F. (2013). The social fabric of the RNA degradosome. In Biochimica et Biophysica Acta – Gene Regulatory Mechanisms (Vol. 1829, Numbers 6–7, pp. 514–522). https://doi.org/10.1016/j.bbagrm.2013.02.011
- Charlesworth, A., Meijer, H. A., & De Moor, C. H. (2013). Specificity factors in cytoplasmic polyadenylation. Wiley Interdisciplinary Reviews: RNA, 4(4), 437–461. https://doi.org/10.1002/wrna.1171
- M., Faini, M., Muckenfuss, L. M., Aebersold, R., & Jinek, M. (2018). Structural basis of AAUAAA polyadenylation signal recognition by the human CPSF complex. Nature Structural and Molecular Biology, 25(2), 135–138. https://doi.org/10.1038/s41594-017-0020-6
- Guo, S., & Lin, S. (2023). mRNA alternative polyadenylation (APA) in regulation of gene expression and diseases. In Genes and Diseases (Vol. 10, Number 1, pp. 165–174). KeAi Communications Co. https://doi.org/10.1016/j.gendis.2021.09.005
- Gutierrez, P. A., Wei, J., Sun, Y., & Tong, L. (2022). Molecular basis for the recognition of the AUUAAA polyadenylation signal by mPSF. https://doi.org/10.1261/rna
- E., & Kaberdin, V. R. (2018). RNA polyadenylation and its consequences in prokaryotes. In Philosophical Transactions of the Royal Society B: Biological Sciences (Vol. 373, Number 1762). Royal Society Publishing. https://doi.org/10.1098/rstb.2018.0166
- Huang, Y. S., Mendez, R., Fernandez, M., & Richter, J. D. (2023). CPEB and translational control by cytoplasmic polyadenylation: impact on synaptic plasticity, learning, and memory. In Molecular Psychiatry (Vol. 28, Number 7, pp. 2728–2736). Springer Nature. https://doi.org/10.1038/s41380-023-02088-x
- Ivshina, M., Lasko, P., & Richter, J. D. (2014). Cytoplasmic polyadenylation element binding proteins in development, health, and disease. In Annual Review of Cell and Developmental Biology (Vol. 30, pp. 393–415). Annual Reviews Inc. https://doi.org/10.1146/annurev-cellbio-101011-155831
- Jester, B. C., Romby, P., & Lioliou, E. (2012). When ribonucleases come into play in pathogens: A survey of gram-positive bacteria. In International Journal of Microbiology. https://doi.org/10.1155/2012/592196
- Kushner, S. R. (2013). Poly(A) Tails. Brenner’s Encyclopedia of Genetics: Second Edition, 384–386. https://doi.org/10.1016/B978-0-12-374984-0.01183-9
- Lutz, C. S., & Moreira, A. (2011). Alternative mRNA polyadenylation in eukaryotes: An effective regulator of gene expression. In Wiley Interdisciplinary Reviews: RNA (Vol. 2, Number 1, pp. 22–31). https://doi.org/10.1002/wrna.47
- Maes, A., Gracia, C., Innocenti, N., Zhang, K., Aurell, E., & Hajnsdorf, E. (2017). Landscape of RNA polyadenylation in E. coli. Nucleic Acids Research, 45(5), 2746–2756. https://doi.org/10.1093/nar/gkw894
- Marsollier, A. C., Joubert, R., Mariot, V., & Dumonceaux, J. (2018). Targeting the polyadenylation signal of pre-mRNA: A new gene silencing approach for facioscapulohumeral dystrophy. In International Journal of Molecular Sciences (Vol. 19, Number 5). MDPI AG. https://doi.org/10.3390/ijms19051347
- Millevoi, S., & Vagner, S. (2009). Molecular mechanisms of eukaryotic pre-mRNA 3’ end processing regulation. Nucleic Acids Research, 38(9), 2757–2774. https://doi.org/10.1093/nar/gkp117616.
- Mitschka, S., & Mayr, C. (2022). Context-specific regulation and function of mRNA alternative polyadenylation. In Nature Reviews Molecular Cell Biology (Vol. 23, Number 12, pp. 779–796). Nature Research. https://doi.org/10.1038/s41580-022-00507-5
- Mohanty, B. K., & Kushner, S. R. (2011). Bacterial/archaeal/organellar polyadenylation. In Wiley Interdisciplinary Reviews: RNA (Vol. 2, Number 2, pp. 256–276). https://doi.org/10.1002/wrna.51
- Pavanello, L., Hall, M., & Winkler, G. S. (2023a). Regulation of eukaryotic mRNA deadenylation and degradation by the Ccr4-Not complex. In Frontiers in Cell and Developmental Biology (Vol. 11). Frontiers Media SA. https://doi.org/10.3389/fcell.2023.1153624
- Pavanello, L., Hall, M., & Winkler, G. S. (2023b). Regulation of eukaryotic mRNA deadenylation and degradation by the Ccr4-Not complex. In Frontiers in Cell and Developmental Biology (Vol. 11). Frontiers Media SA. https://doi.org/10.3389/fcell.2023.1153624
- Rehfeld, A., Plass, M., Døssing, K., Knigge, U., Kjær, A., Krogh, A., & Friis-Hansen, L. (2014). Alternative polyadenylation of tumor suppressor genes in small intestinal neuroendocrine tumors. Frontiers in Endocrinology, 5(APR). https://doi.org/10.3389/fendo.2014.00046
- Schönemann, L., Kühn, U., Martin, G., Schäfer, P., Gruber, A. R., Keller, W., Zavolan, M., & Wahle, E. (2014). Reconstitution of CPSF active in polyadenylation: Recognition of the polyadenylation signal by WDR33. Genes and Development, 28(21), 2381–2393. https://doi.org/10.1101/gad.250985.114
- Them, S. L., Wallace, R. B., Pressley, L., Clegg, J. B., Weatherall, D. J., & Higgs, D. R. (1988). The Polyadenylation Site Mutation in the a-Globin Gene Cluster In a previous study. we described a form of nondeletion a-thalassemia (Vol. 71, Number 2). https://ashpublications.org/blood/article-pdf/71/2/313/596508/313.pdf
- Thore, S., Raoelijaona, F., Talenton, V., Fribourg, S., & Mackereth, C. D. (2023). Molecular details of the CPSF73-CPSF100 C-terminal heterodimer and interaction with Symplekin. Open Biology, 13(11). https://doi.org/10.1098/rsob.230221
- Venkat, S., Tisdale, A. A., Schwarz, J. R., Alahmari, A. A., Carlo Maurer, H., Olive, K. P., Eng, K. H., & Feigin, M. E. (2020). Alternative polyadenylation drives oncogenic gene expression in pancreatic ductal adenocarcinoma. Genome Research, 30(3), 347–360. https://doi.org/10.1101/gr.257550.119
- Wahle, E., & Winkler, G. S. (2013). RNA decay machines: Deadenylation by the Ccr4-Not and Pan2-Pan3 complexes. In Biochimica et Biophysica Acta – Gene Regulatory Mechanisms (Vol. 1829, Numbers 6–7, pp. 561–570). https://doi.org/10.1016/j.bbagrm.2013.01.003
- Xiang, Y., Ye, Y., Lou, Y., Yang, Y., Cai, C., Zhang, Z., Mills, T., Chen, N. Y., Kim, Y., Ozguc, F. M., Diao, L., Karmouty-Quintana, H., Xia, Y., Kellems, R. E., Chen, Z., Blackburn, M. R., Yoo, S. H., Shyu, A. Bin, Mills, G. B., & Han, L. (2018). Comprehensive characterization of alternative polyadenylation in human cancer. Journal of the National Cancer Institute, 110(4), 379–389. https://doi.org/10.1093/jnci/djx223
- Yuan, F., Hankey, W., Wagner, E. J., Li, W., & Wang, Q. (2021). Alternative polyadenylation of mRNA and its role in cancer. In Genes and Diseases (Vol. 8, Number 1, pp. 61–72). Chongqing University. https://doi.org/10.1016/j.gendis.2019.10.011
- Zhang, X., Virtanen, A., & Kleiman, F. E. (2010). To polyadenylate or to deadenylate: That is the question. In Cell Cycle (Vol. 9, Number 22, pp. 4437–4449). Taylor and Francis Inc. https://doi.org/10.4161/cc.9.22.13887
