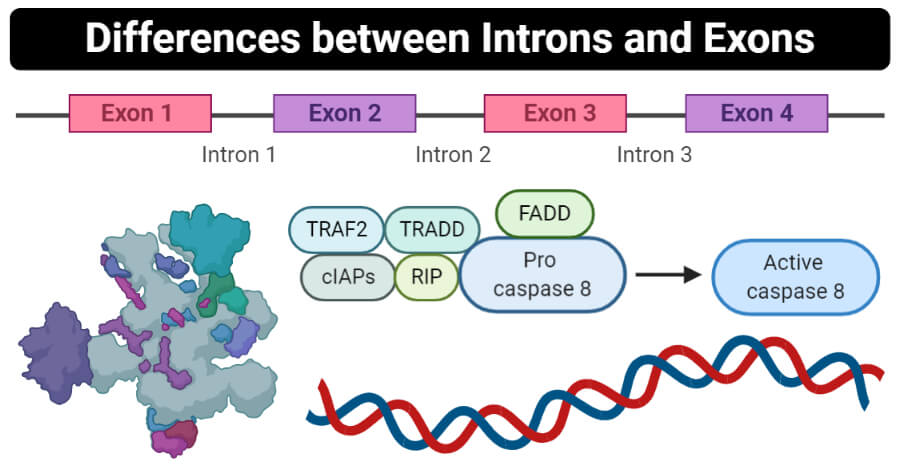
Introns Definition
Introns are non-coding DNA sequences within a gene that are removed by RNA splicing during maturation of the RNA product.
- The term ‘intron’ represents the intragenic region which is present within a gene.
- The term ‘introns’ indicates both the DNA sequences within the gene and the corresponding sequence in RNA transcripts.
- Introns are found in the genes of many eukaryotic organisms and also some viruses and are located in most genes including those that generate proteins, ribosomal RNA (rRNA), and transfer RNA (tRNA). These are, however, not found in prokaryotes.
- Introns are common in protein-coding nuclear genes of most jawed invertebrates but hey might be rare in some eukaryotic organisms.
- Similarly, the mitochondrial genomes of jawed vertebrates are almost entirely devoid of introns whereas those in other eukaryotes have many introns.
- During the generation of proteins from genes containing introns, RNA splicing occurs as a process of RNA processing that occurs after transcription and before translation.
- There are different types of introns based on their sequence analysis and the genetic and biochemical analysis of RNA splicing methods.
- The four more common types of introns include;
- Spliceosome introns in nuclear protein-coding genes that are removed by spliceosomes
- tRNA introns in nuclear and archaeal tRNA genes that are removed by proteins
- self-splicing group I introns removed by RNA catalysis
- self-splicing group II introns removed by RNA catalysis
- Different introns are also lost and gained throughout evolution as observed in different eukaryotes.
- Introns are crucial because the types of protein are greatly enhanced by alternative splicing in which introns take part in important roles. Alternative splicing is a controlled molecular mechanism producing multiple variant proteins from a single gene in a eukaryotic cell.
- The level of gene expression is greatly enhanced in the presence of introns.
- It has also been reported that spliced transcripts are exported faster from the nucleus to cytoplasm than the unspliced ones.
- However, the existence of introns in the genome might be a burden to some cells, because the cells have to consume a great deal of energy to copy and excise them exactly at the correct positions with the help of complicated spliceosomal techniques.
Exons Definition
Exons are protein-coding DNA sequences that require the necessary codons or information necessary for protein synthesis.
- The term ‘exon’ represents the expressed region present in the genome.
- The genes in eukaryotes are formed of coding exons separated by non-coding introns.
- During RNA splicing, the introns between the exons are removed to connect two different introns that then code for messenger RNA.
- The entire set of all exons present in the genome of the organisms is termed exosome.
- In genes coding for proteins, exons include both the protein-coding sequence and the 5’ and 3’ untranslated regions.
- Once these genes are transcribed, the resulting RNA has both exons and introns. The introns are then removed by RNA splicing resulting in mature mRNAs.
- The mature mRNA transcripts thus have exons and untranslated regions where the exons form a small part of the entire sequence.
- Exons are present in all organisms ranging from jawed vertebrates to viruses.
- In the human genome, only 1% of the total genome is formed of exons while the rest is occupied by introns and intergenic DNA.
- Sometimes, some introns are converted into exons by the process of exonization.
- Exons are crucial in protein synthesis as they are regions carrying codons that code for various proteins.
- The presence of exons and introns allows the process of alternative splicing that increases the variety of proteins produced from a single gene.
- Alternative splicing allows exons to be arranged in different sequences where different configurations result in different proteins.
- A process similar to alternative splicing is exon shuffling where exons or sister chromosomes are exchanged during recombination.
- Alternative splicing occurs commonly in a human gene that codes for a transmembrane protein involved in the regulation of potassium entry in the hair cell.
- This gene consists of 35 exons which can combine in different ways or configuration to form over 500 mRNAs by the reshuffling of about one to eight exons.
Introns vs Exons (Table Form)
| Basis for comparison | Introns | Exons |
| Definition | Introns are non-coding DNA sequences within a gene that are removed by RNA splicing during maturation of the RNA product. | Exons are protein-coding DNA sequences that require the necessary codons or information necessary for protein synthesis. |
| Type of sequence | Introns are the non-coding sequences that do not code for any protein. | Exons are protein-coding sequences that code for specific proteins. |
| Location in the DNA | Introns are present between two exons in a DNA sequence. | Exons are the sequences coding for proteins that are present between either the untranslated regions or two introns. |
| Distribution | These are found only in eukaryotic genomes. | These are found in both eukaryotic and prokaryotic genomes. |
| Location in the cell | Introns remain in the nucleus after being spliced out from the mRNA transcript during RNA processing. | Exons leave the nucleus to reach the cytoplasm after the mature mRNAs are synthesized. |
| Present in | Introns are present in the DNA and the mRNA transcripts but are not present in mature mRNAs. | Exons are present in DNA, mRNA transcripts, and mature RNAs. |
| Conserved | The sequences in introns are as conserved as the sequences of exons. Some introns might convert into exons by the process of exonization. | The sequences in exons are highly conserved. |
| Involved | Introns are not involved in protein synthesis. | Exons are involved in protein synthesis. |
| Quantity | More introns are present in the nuclear genome than exons. | Exons are present in lesser quantity than introns in the nuclear genome. |
| Human genome | About 24% of the human genome is composed of introns. | Only 1% of the human genome is composed of exons. |
| Alternative splicing | Introns are removed by alternative splicing. | Two or more exons are connected after alternative splicing. |
| Novel genes formation | Introns might result in novel genes as the short non-coding regions might evolve into real functional genes through a kind of continuous evolutionary process. | Exons might combine in a different configuration forming different sequences that code for different proteins. |
References and Sources
- Jo, B. S., & Choi, S. S. (2015). Introns: The Functional Benefits of Introns in Genomes. Genomics & informatics, 13(4), 112–118. https://doi.org/10.5808/GI.2015.13.4.112
- 4% – https://newoptionsnm.info/ekson-dan-intron-46/
- 4% – https://en.wikipedia.org/wiki/Intron
- 3% – https://egli-online.com/ekson-dan-intron-14/
- 2% – https://wiki2.org/en/RNA_splicing
- 2% – https://quizlet.com/22075306/bio-319-exam-3-eukaryotic-genomes-flash-cards/
- 1% – https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3150433/
- 1% – https://sciencing.com/introns-vs-exons-what-are-the-similarities-differences-13718414.html
- 1% – https://en.wikipedia.org/wiki/Exons
- 1% – https://en.wikipedia.org/wiki/Exome
- 1% – https://biologydictionary.net/exon/
- <1% – https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2958924/
