Gene interactions refer to the relationships between genes that affect the phenotype of an organism. Gene interactions occur when allelic or non-allelic genes affect the expression of specific phenotypic traits in an organism.
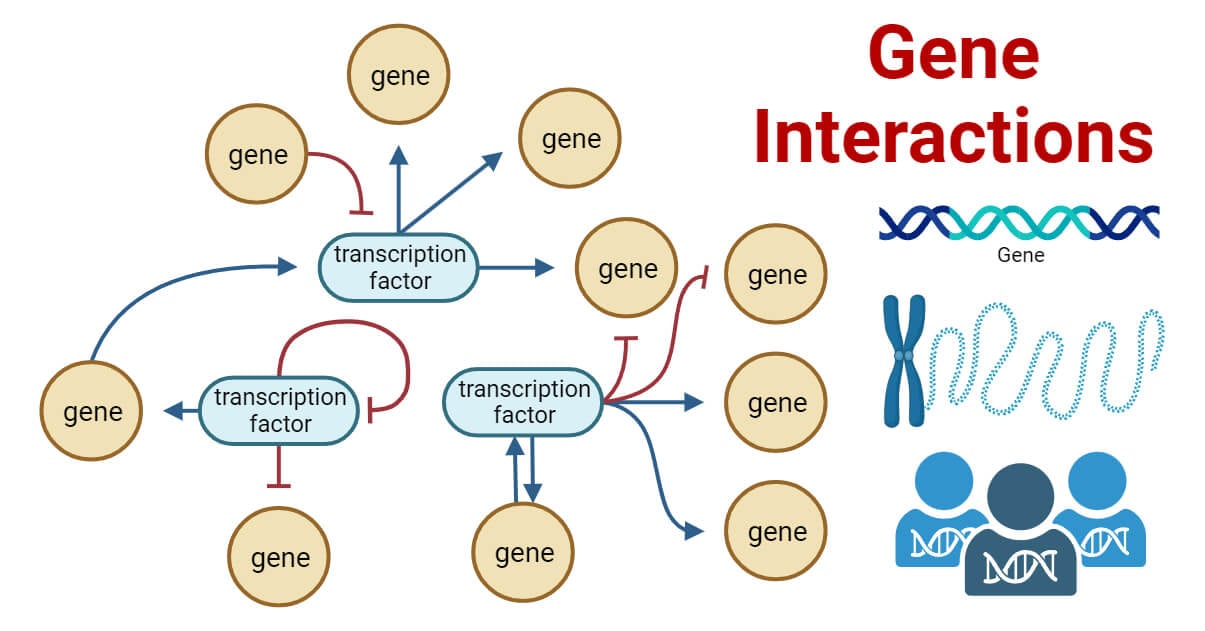
The study of gene interactions gained recognition following Gregor Mendel’s work on inheritance. Unlike Mendel’s initial observations, it was discovered that inheritance is not a simple one-gene, one-trait relationship. It involves the interaction of multiple genes, and their combined effects cannot be predicted by studying them individually.
Bateson and Punnett conducted breeding experiments with chickens, shortly after Mendel’s paper was rediscovered, and their research provided early evidence that traits can be influenced by multiple genes. They noticed that the observed ratios of inherited traits did not always follow Mendelian patterns, indicating that genes interact with each other.
Types of Gene Interactions
Gene interactions come in different forms, each providing valuable information about biological processes. Two main categories of gene interactions are allelic and non-allelic gene interactions.
Allelic Gene Interaction
Allelic gene interactions occur between different forms (alleles) of a single gene. Some of the types of allelic gene interactions are discussed below:
Complete Dominance
- Complete dominance is an allelic gene interaction that follows Mendel’s laws of inheritance. According to this pattern, each gene has two alleles, with one allele completely dominant over the other.
- The presence of the dominant allele masks the effect of the recessive allele, leading to the expression of a single phenotype.
- Archibald Garrod’s work in 1903 led to the discovery of complete dominance and its application to human diseases. Garrod studied alkaptonuria, a disease characterized by the excretion of homogentisic acid in urine. By analyzing family histories and urine samples, Garrod observed a 3:1 ratio in affected families, consistent with Mendelian inheritance.
Incomplete Dominance
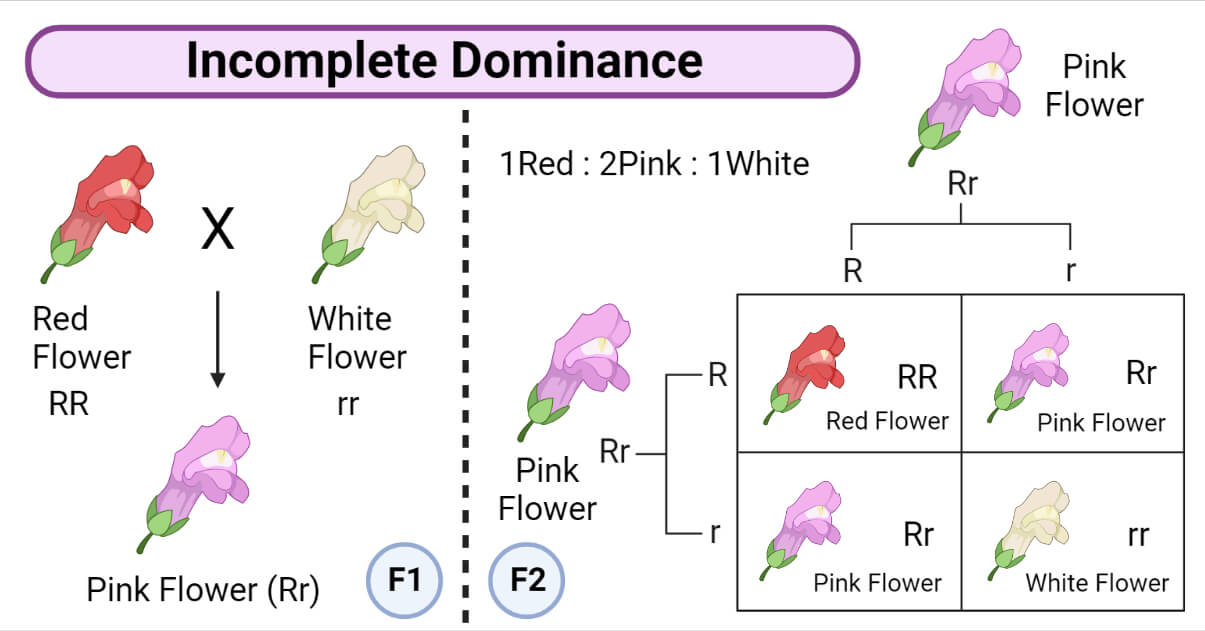
- Incomplete dominance occurs when two organisms with different traits are crossed, and their offspring show an intermediate phenotype that is a combination of the parental traits.
- Incomplete dominance can be observed when the enzymes involved in a particular trait have slight differences in their activity.
- For example, when red snapdragon plants are crossed with ivory snapdragon plants, the resulting F1 hybrid plants have pink flowers. When these F1 hybrids are self-bred, the F2 generation shows a phenotypic ratio of 1 red:2 pink:1 ivory.
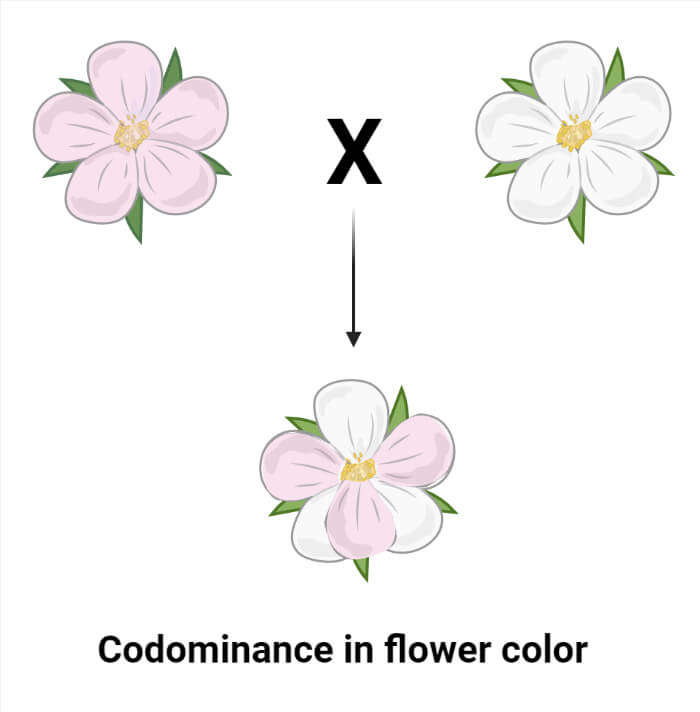
Codominance
- Codominance is a type of inheritance where two different alleles of a gene are expressed simultaneously and independently in an individual.
- In this case, neither allele is dominant over the other, and both traits associated with each allele are fully expressed. This results in the expression of multiple traits simultaneously, without any blending.
- Unlike complete dominance, where one allele completely masks the expression of the other, codominance allows both alleles to be visible in the phenotype of the organism.
- The ABO blood group system in humans provides a clear example of codominance, as the antigen alleles exhibit equal expression.
Lethal Allele
- Lethal alleles are mutant alleles that interfere with vital functions and can lead to the death of an organism.
- They are often associated with human diseases and are considered essential genes.
- Dominant lethal alleles that act early in life are lost in one generation as carriers die before passing them on. However, dominant lethal alleles that act later in life, after reproduction has occurred, can be inherited.
- On the other hand, recessive lethal mutations can persist longer as they can hide in heterozygotes, where a normal gene masks the mutant allele’s effect.
- An example of a recessive lethal mutation is the yellow-lethal mutation (AY) in mice. This mutation is both dominant and visible, causing the fur color to be yellow instead of the wild-type gray-brown (agouti). However, the AY mutation is also recessive lethal, meaning that individuals with two copies of the AY allele die during early embryonic development. When AYA+ heterozygotes are crossed, the progeny includes yellow (AYA+) and gray-brown (A+A+) mice in a ratio of 2:1. The AYAY homozygotes do not survive due to the lethal effect of the mutation.
Multiple Alleles
- Mendel’s initial work suggested that genes have only two alleles, but this is not always the case.
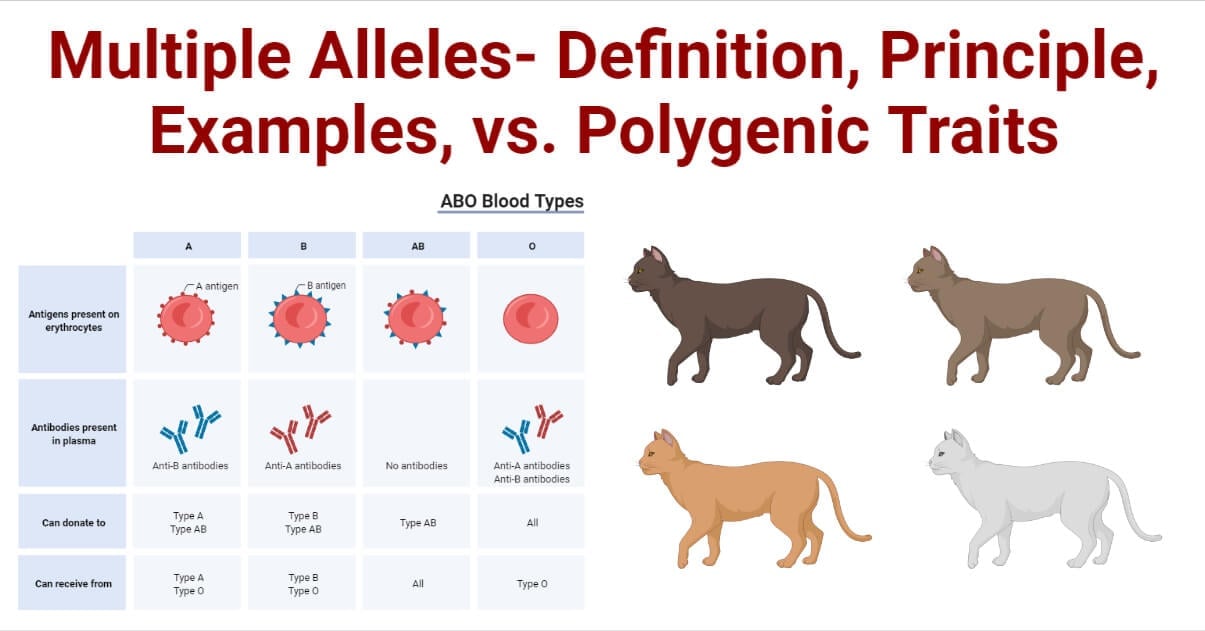
- Some genes have more than two allelic forms at the same chromosomal location. This leads to the presence of multiple alleles within a population.
- For example, in rabbits, the gene that controls coat color exhibits multiple alleles. This gene, represented by “c,” has four alleles: c (Albino), ch (Himalayan), cch (Chinchilla), and c+ (Wild-type). This example of a gene with multiple alleles in rabbits demonstrates the variation in coat color and the different effects that each allele can have on the phenotype.

Non-Allelic Gene Interaction
Non-allelic gene interactions refer to interactions between genes that are located on the same or different chromosomes but do not involve alleles of the same gene. Some of the types of non-allelic gene interactions are discussed below:
Complementary Genes
- Complementary gene interactions occur when two or more genes located at different loci and inherited from different parents interact to produce specific characteristics.
- These genes complement each other’s functions and require their combined presence for the phenotype to be expressed.
- When these genes are present individually, they do not have the ability to express the trait. However, when they come together through appropriate mating or crossing, they complement each other and the trait is expressed.
- For example, in sweet peas, C and P genes are involved in the synthesis of anthocyanin pigment, which gives the flowers their purple color. Both of these genes are required for the production of the pigment. If either gene C or gene P is absent or non-functional, the plant cannot produce anthocyanin and the flowers appear white instead of purple. This shows that the genes C and P are complementary to each other for the formation of anthocyanin pigment.
Epistasis
- Epistasis is a form of genetic interaction between non-allelic genes that occur when the effect of one gene is masked or modified by another gene.
- The gene exerting the effect is called the epistatic gene, while the gene being suppressed is the hypostatic gene.
- It was first described by William Bateson in 1907 to explain deviations from Mendelian inheritance.
- The term “epistasis” has two distinct meanings in the field of genetics. Mendelian and molecular geneticists use it strictly to refer to specific gene interactions where one locus masks another. In contrast, evolutionary and quantitative geneticists use “epistasis” to describe any form of gene interaction.
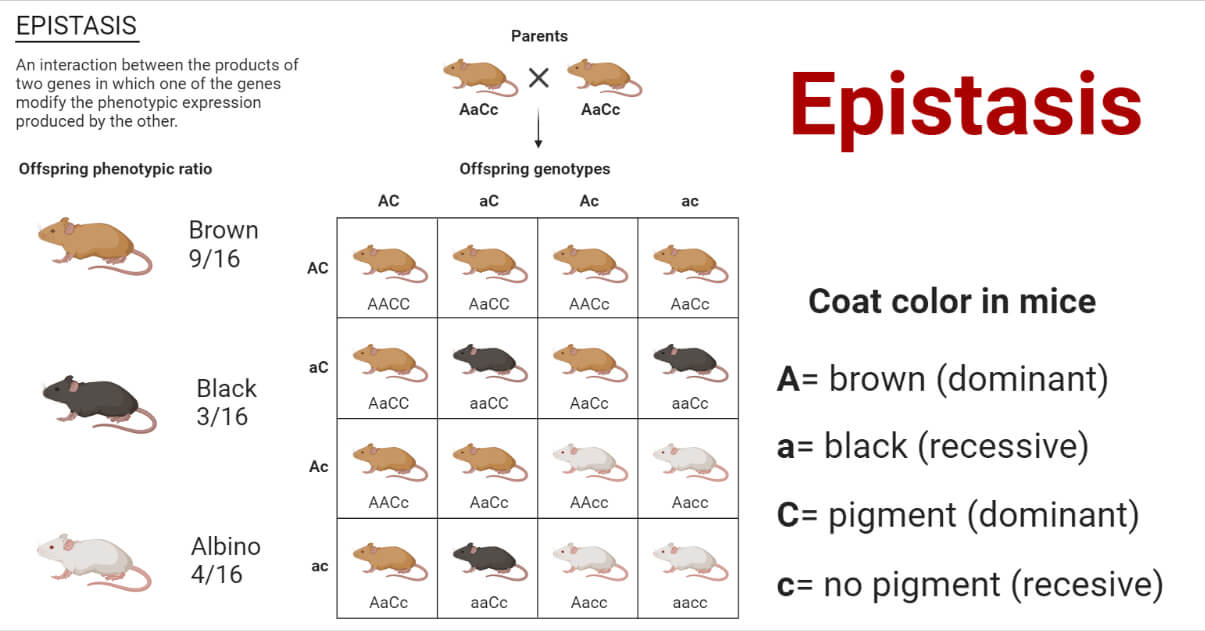
Epistasis can be of the following two types:
Dominant epistasis occurs when the presence of a dominant allele at one gene locus masks the effects of alleles at another locus.
Example: In foxgloves (Digitalis purpurea), two genes interact to determine petal coloration. The alleles d and D affect the red pigment intensity, and the alleles w and W determine where the pigment is synthesized in the petals.
The dominant allele W restricts pigment synthesis to throat spots, resulting in white petals with spots. F2 generation from the cross between D/d and W/w alleles indicates that the W allele shows dominant epistasis. It prevents red pigment synthesis in most petals but allows it in the throat spots.
Figure: In foxgloves, D and d cause dark and light pigments, respectively, whereas the epistatic W restricts pigment to the throat spots.
Recessive epistasis occurs when the recessive allele of one gene locus masks or prevents the phenotype expression of alleles at another gene locus.
For example, some Labrador retriever dogs have a yellow coat color due to recessive epistasis. The presence of the e (epistatic) allele interacts with the B (black coat) and b (brown coat) alleles to produce a yellow coat phenotype.
Mapping Gene Interactions
- Genetic interaction mapping is a technique used to study and characterize complex gene interactions.
- Analyzing and understanding these interactions helps to understand how genes function and define biological pathways.
- This can be done by deleting genes or inhibiting gene expression to observe the resulting changes in the phenotype.
- Different techniques like Pairwise deletion, gene expression inhibition, and high-throughput screening methods provide powerful tools for mapping gene interactions.
Significance of Gene Interactions
- Gene interactions can be studied in model organisms to understand the functional relationships between genes and their gene products.
- Gene interactions are important for understanding the key components and mechanisms involved in genetic pathways.
- Gene interactions play a crucial role in understanding the complex phenotypic variation in organisms. This helps us to understand how different genes work together to express different phenotypic traits.
- Gene interactions are important for understanding how complex genetic systems evolve by studying how genes interact and change over time.
- Gene interactions are complex and involve various molecular mechanisms. Understanding these complex interactions is essential for understanding how genes work together and their impact on different biological processes.
References
- 2.2: Multiple alleles, incomplete dominance, and codominance – Biology LibreTexts
- Boucher, B., & Jenna, S. (2013). Genetic interaction networks: Better understand to better predict. Frontiers in Genetics, 4, 68624. https://doi.org/10.3389/fgene.2013.00290
- Griffiths, A. J. F., Miller, J. H., Suzuki, D. T., Lewontin, R. C., Gelbart, W. M. (2000). An introduction to genetic analysis (7th ed.). New York: W. H. Freeman.
- Hartl D. L. (2014). Essential genetics: a genomics perspective (6th ed.). Jones and Bartlett.
- https://www.biologydiscussion.com/genetics/gene-interactions/gene-interactions-allelic-and-non-allelic-cell-biology/38795
- https://www.nature.com/scitable/topicpage/epistasis-gene-interaction-and-phenotype-effects-460/
- https://www.nature.com/scitable/topicpage/gene-interaction-and-disease-987/
- Kaushik, S., Kaushik, S., & Sharma, D. (2018). Functional Genomics. Reference Module in Life Sciences. doi:10.1016/b978-0-12-809633-8.20222-7
- Mani, R., St.Onge, R. P., Hartman, J. L., Giaever, G., & Roth, F. P. (2008). Defining genetic interaction. Proceedings of the National Academy of Sciences, 105(9), 3461–3466. doi:10.1073/pnas.0712255105
- Moore, J. H. (2013). Gene Interaction. Brenner’s Encyclopedia of Genetics, 200–201. doi:10.1016/b978-0-12-374984-0.00592-1
- Phillips, P. C. (1998). The language of gene interaction. Genetics, 149(3), 1167-1171. https://doi.org/10.1093/genetics/149.3.1167
- Phillips, P. C. (2008). Epistasis—The essential role of gene interactions in the structure and evolution of genetic systems. Nature reviews. Genetics, 9(11), 855. https://doi.org/10.1038/nrg2452

it is mainly for advanced research and learning
Nice.