Enzymes are biological catalysts that speed up the rate of a specific chemical reaction in the cell. They catalyze the essential biochemical processes within living organisms. Enzymes work by binding to substrate molecules at specific sites called active sites, facilitating and accelerating chemical reactions.
The activity of enzymes must be regulated so that they function properly. There are various mechanisms to regulate the biological activity of enzymes. One of the important regulatory mechanisms is allosteric regulation in which a regulatory molecule, either an activator or inhibitor, binds to an enzyme at a site other than the enzyme’s active site. This specific binding site on the enzyme is known as the allosteric site. Both active sites and allosteric sites are present in allosteric enzymes. Allosteric regulation cannot be explained by the simple Michaelis–Menten (MM) model.
What is Allosteric inhibition?
Allosteric inhibition is a regulatory mechanism where an allosteric inhibitor binds to the specific allosteric site.
As a result of this binding, conformational change occurs in the enzyme’s structure, which indirectly reduces the enzyme’s activity. Allosteric inhibition is reversible. When the allosteric inhibitor molecule dissociates from the allosteric site, the enzyme can return to its original conformation and regain its activity.
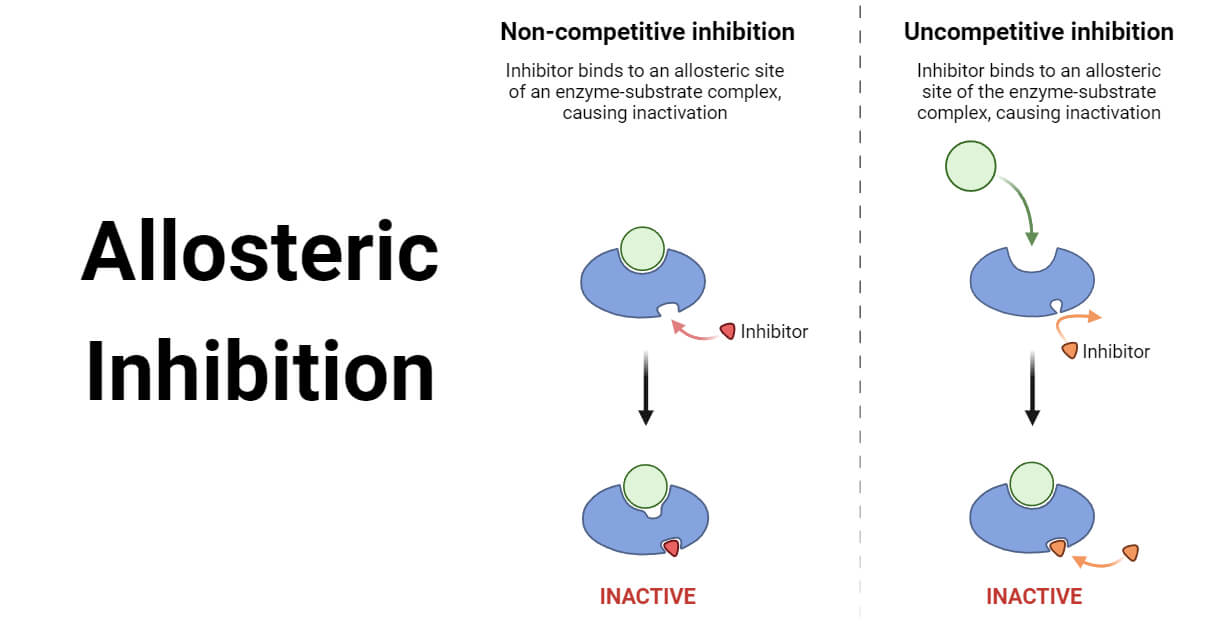
Allosteric inhibitors are regulatory molecules that interact with enzymes at allosteric sites and inhibit the binding of substrate or ligand molecules. Allosteric inhibitors do not compete with the substrate for binding at the active site of the enzyme. Instead, they bind to the allosteric site, which indirectly changes the enzyme’s shape or activity.
Mechanism of allosteric inhibition
Allosteric binding can occur in two ways: noncompetitive inhibition and uncompetitive inhibition. Both mechanisms regulate enzyme activity by inhibiting or reducing the desired chemical reaction.
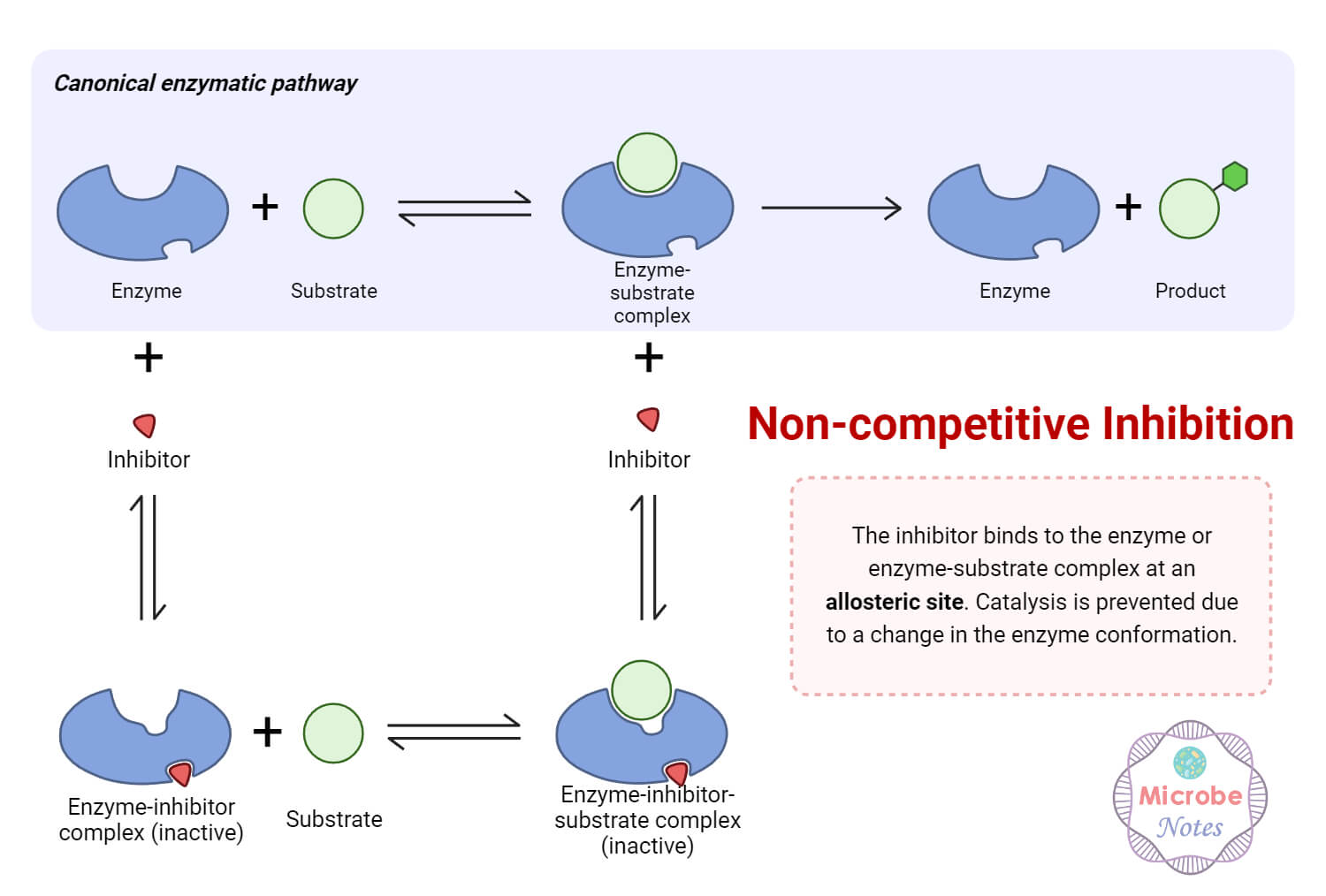
1. Noncompetitive Inhibition
- Noncompetitive inhibition occurs when the inhibitor molecule (I) binds to the allosteric site of the enzyme (E) before the substrate-binding event occurs.
- Once bound, the inhibitor changes the structure of the enzyme, which can alter the active site.
- As a result, the substrate (S) cannot bind effectively to the active site, which reduces the ability of enzymes to catalyze the reaction and form products (P).
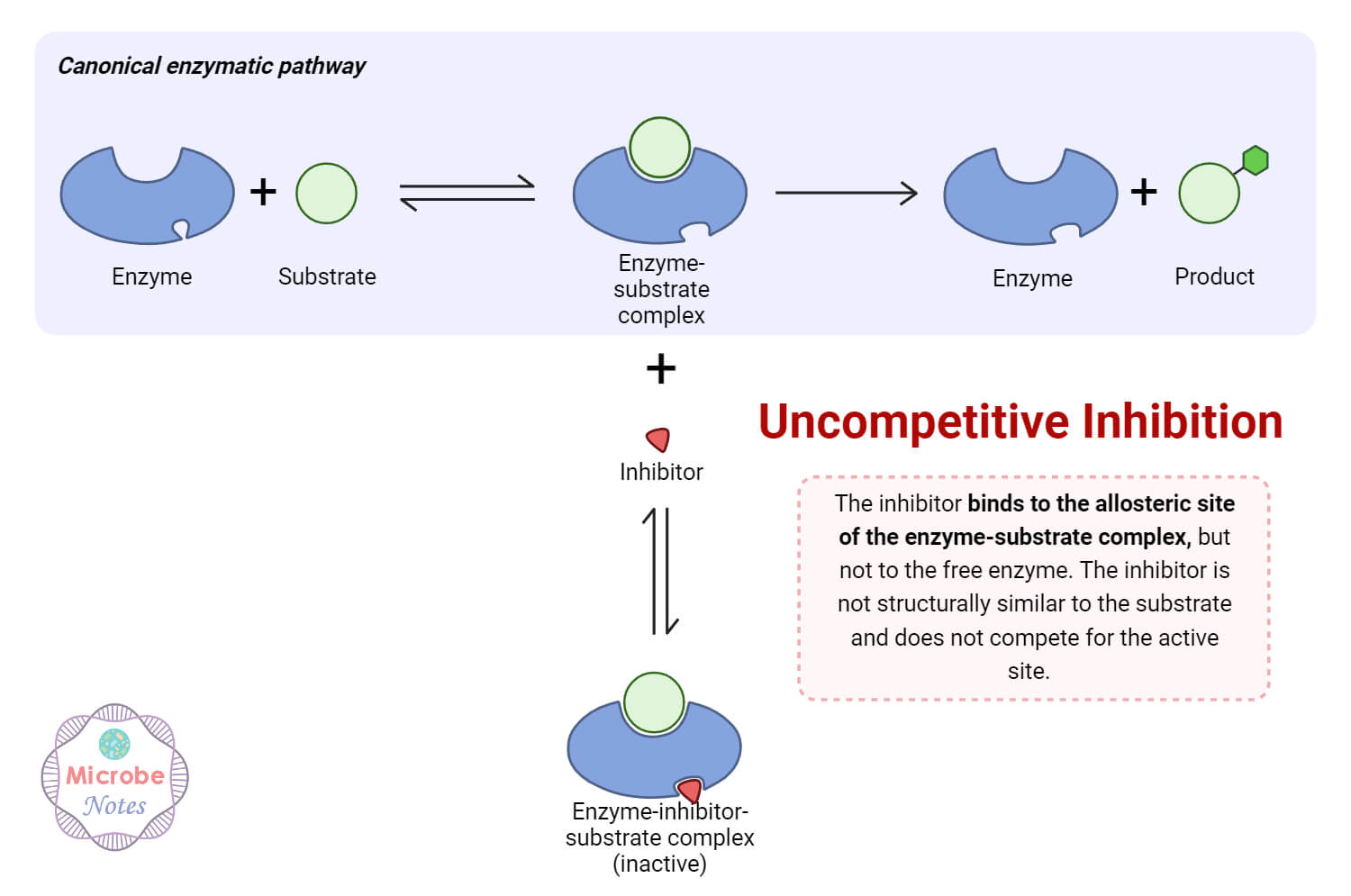
2. Uncompetitive Inhibition
- Uncompetitive inhibition occurs after the substrate binds to the enzyme and targets the enzyme-substrate (ES) complex.
- The inhibitor binds specifically to the ES complex and forms the enzyme-substrate-inhibitor (ESI) complex, which slows down the conversion of the complex into products.
- This type of inhibition mainly affects the catalytic step of product formation, making it slower or less efficient.
Cooperativity in allosteric enzymes
One of the special characteristics of allosteric enzymes is the presence of cooperativity in substrate binding. Cooperativity is a phenomenon displayed by enzymes with multiple binding sites where the affinity of the binding sites for a ligand is either increased or decreased when a ligand binds to a binding site.
There are two main models that explain the cooperativity in allosteric enzymes.
- Concerted Model:
- The concerted model, also called the MWC model, was introduced by Monod, Wyman, and Changeux in 1965.
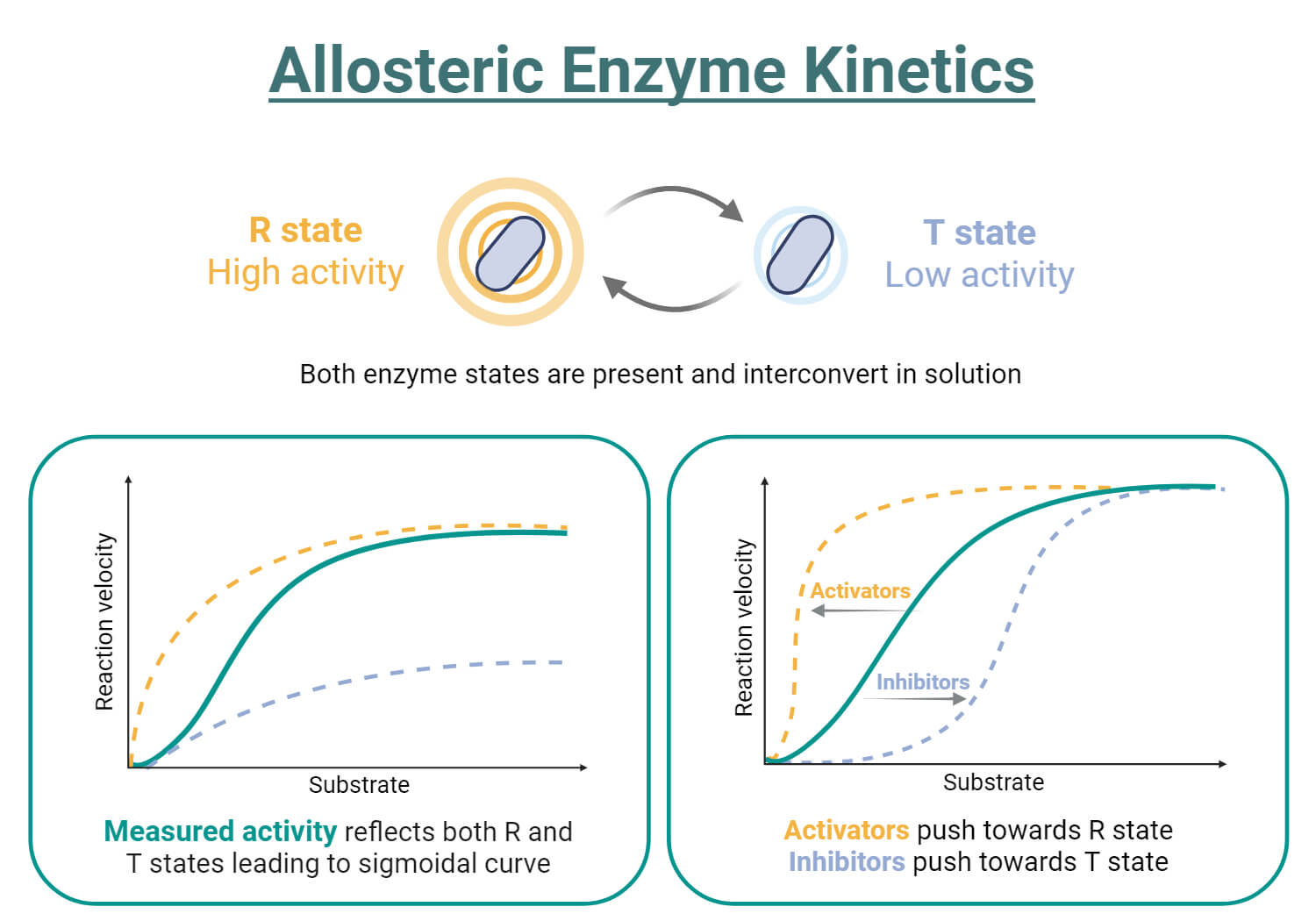
- According to the concerted model, enzymes with multiple subunits can exist in one of two conformational states: the relaxed (R) state or the tense (T) state.
- These states differ in the energy of their interactions between subunits but maintain symmetry, meaning all subunits are either in the T state or the R state.
- In the absence of ligands or substrates, the tense state is favored. As ligands bind, the protein transitions from the tense state to the relaxed state.
- Sequential Model:
- The sequential model, also called the KNF model, was proposed by Koshland, Nemethy, and Filmer in 1966.
- In this model, the binding of a ligand to the enzyme causes a conformational change only in the subunit to which the ligand binds.
- The conformational change in one subunit affects the neighboring subunits, leading to cooperative behavior.
- Unlike the MWC model, the KNF model does not maintain symmetry during the binding process as it contains a mixture of both R and T states.
Examples of Allosteric Inhibition
- Aspartate Transcarbamoylase (ATCase):
- ATCase is an enzyme involved in the synthesis of pyrimidine nucleotides, which are the building blocks of DNA and RNA.
- Cytidine triphosphate (CTP), one of the end products of this pathway, acts as an allosteric inhibitor.
- When CTP binds to the allosteric site of ATCase, it results in a conformational change that reduces the enzyme’s activity. This feedback inhibition prevents the overproduction of pyrimidine nucleotides when they are in excess.
- Phosphofructokinase (PFK-1):
- PFK-1 is a key enzyme in glycolysis, the pathway that breaks down glucose for energy production.
- High levels of ATP act as allosteric inhibitors of PFK-1.
- When ATP binds to the allosteric site, the activity of PFK-1 is reduced. This helps to regulate glycolysis and prevent excess glucose breakdown when the cell has abundant energy.
- Acetyl-CoA Carboxylase (ACC):
- ACC is an important enzyme that is involved in fatty acid synthesis.
- Allosteric inhibitors such as palmitoyl-CoA and long-chain fatty acyl-CoAs bind to the allosteric sites of ACC.
- These inhibitors signal that the end products are produced in excess and hence inhibit ACC, reducing fatty acid synthesis.
Significance of allosteric inhibition
- Allosteric inhibition is one of the important mechanisms to control enzyme activity.
- Allosteric inhibition is used in metabolic pathways for feedback control. It prevents the overproduction of metabolites by inhibiting enzymes when their products are in excess.
- Allosteric inhibitors can also be used to design target-specific proteins. This is useful in drug development.
- Allosteric inhibition also helps to maintain homeostasis by regulating processes like enzyme activity and cell signaling.
References
- 3.6: Allosteric Interactions – Chemistry LibreTexts
- 5.4: Enzyme Inhibition – Chemistry LibreTexts
- Allosteric Enzymes: Characteristics, Models, and Examples (conductscience.com)
- Byun, J. A., VanSchouwen, B., Akimoto, M., & Melacini, G. (2020). Allosteric inhibition explained through conformational ensembles sampling distinct “mixed” states. Computational and Structural Biotechnology Journal, 18, 3803-3818. https://doi.org/10.1016/j.csbj.2020.10.026
- Cooperman, B. S. (2013). Allosteric Regulation. Encyclopedia of Biological Chemistry, 71–74. doi:10.1016/b978-0-12-378630-2.00001-3
- Delaune, K. P., & Alsayouri, K. (2022). Physiology, Noncompetitive Inhibitor. In StatPearls. StatPearls Publishing.
- Liu J, Nussinov R (2016) Allostery: An Overview of Its History, Concepts, Methods, and Applications. PLoS Comput Biol 12(6): e1004966. https://doi.org/10.1371/journal.pcbi.1004966
- Lu, P. (2013). Allostery. Brenner’s Encyclopedia of Genetics, 83. doi:10.1016/b978-0-12-374984-0.00037-1
- Wenthur, C. J., Gentry, P. R., Mathews, T. P., & Lindsley, C. W. (2014). Drugs for Allosteric Sites on Receptors. Annual Review of Pharmacology and Toxicology, 54, 165. https://doi.org/10.1146/annurev-pharmtox-010611-134525
