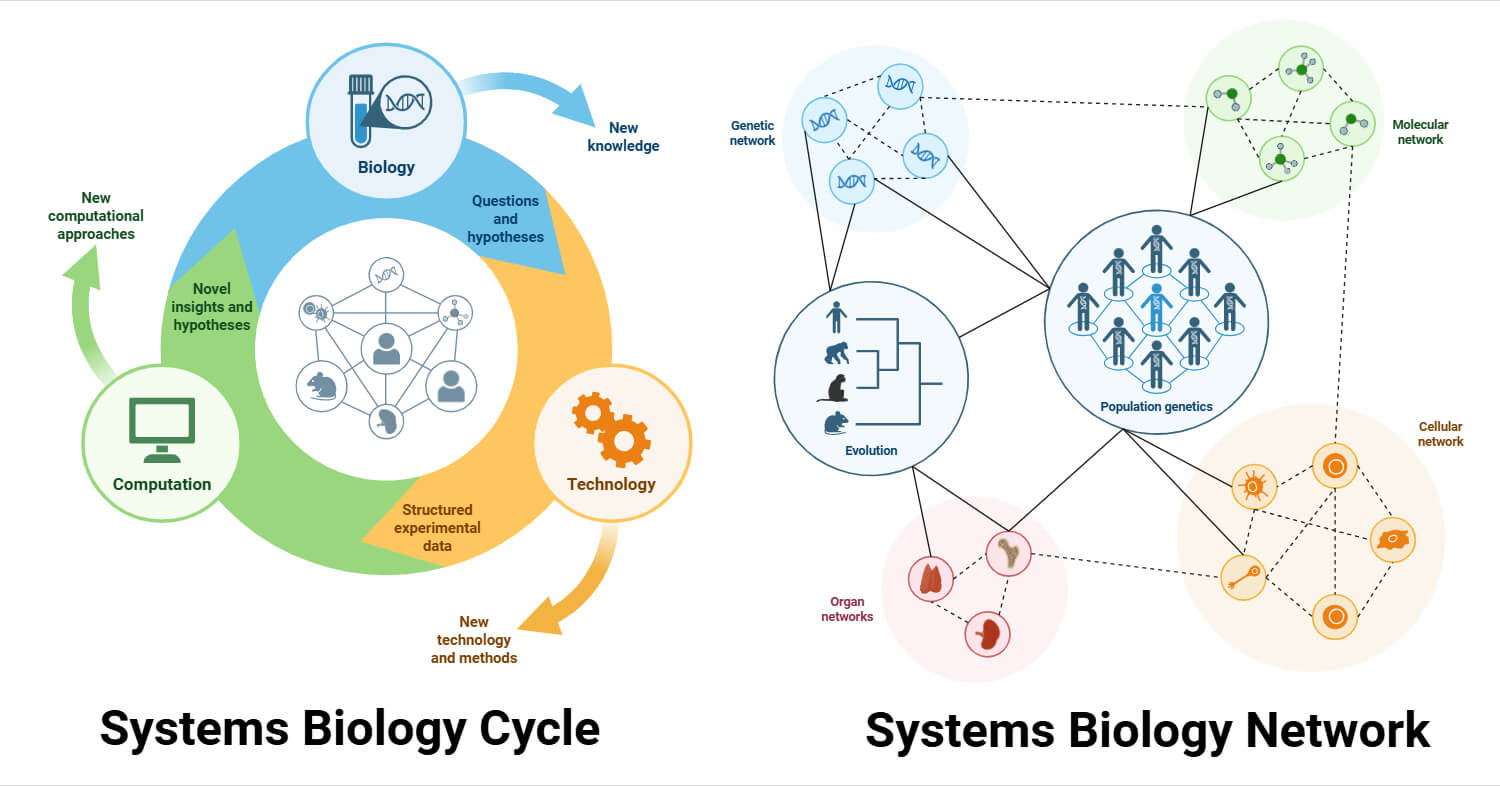
Systems biology is a comprehensive quantitative analysis technique in which the entire biological system functions logically over time. Its main goal is to synthesize all biological data to develop a unified picture of the structure, dynamics, logistics, and logic of living beings.
This analysis technique has been used to revolutionize novel computational tools required for biological intervention and to explore new boundaries of the biological world. It uses molecular biology and biochemistry of cellular components to understand how physiological functions at the cell/tissue, and organ levels arise.
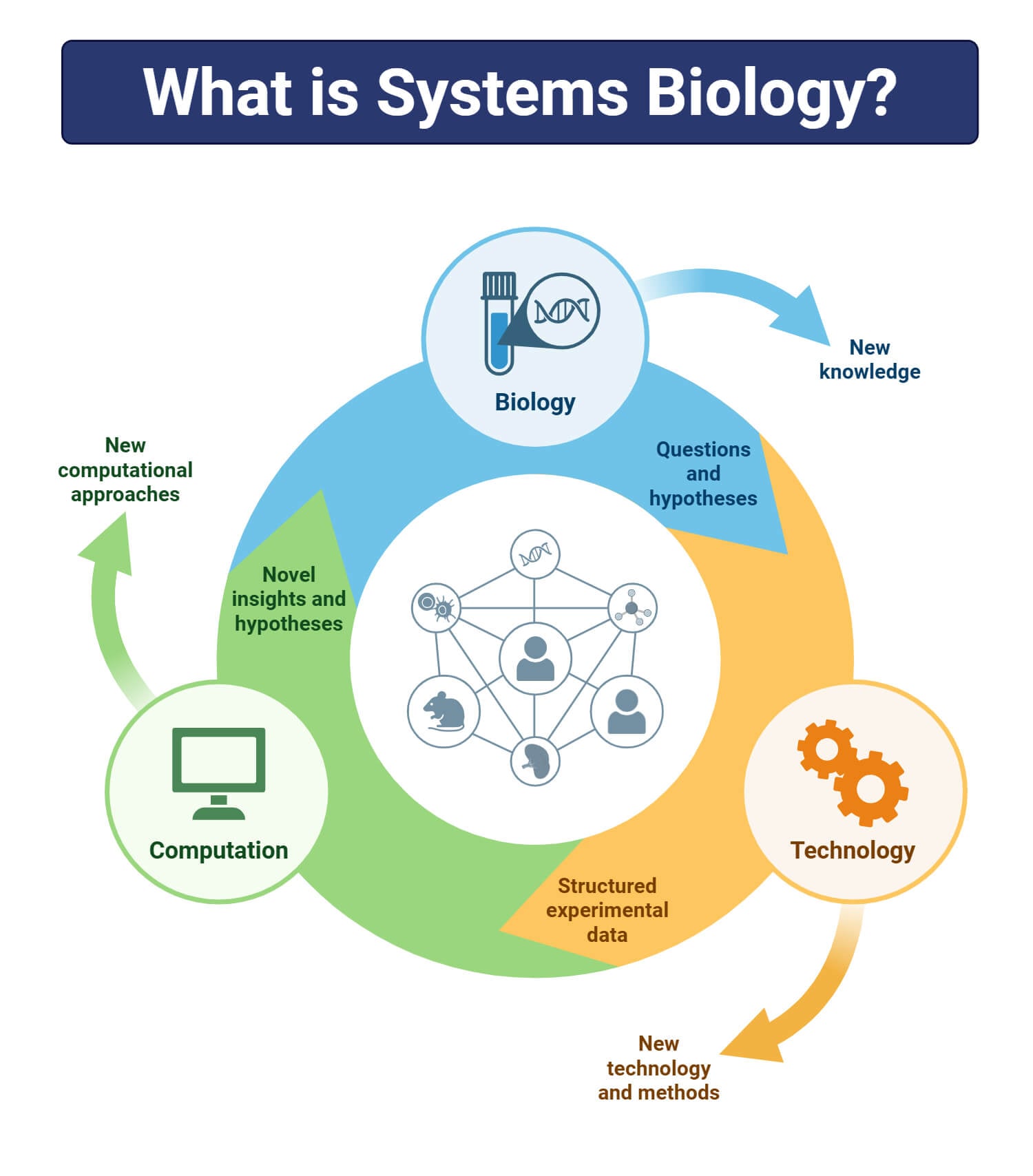
Systems biology enables the flow of information from the processing of large amounts of data generated by genome sequencing, their transcriptional profiling in response to stimuli, interactions between cellular functions, and their significance in application, i.e., drug designing.
Core Principles of Systems Biology
Emergence
Complex systems show a property known as ‘emergent properties’ that are not displayed by individual components and can only be determined by understanding the full picture. Life is an example of an emergent property. Its components, such as DNA, RNA, proteins, carbohydrates, or lipids, do not make it whole, but their interactions and consequences do.
Robustness
Biological systems maintain stability even when external environmental factors, genetic variation, and stochastic changes occur. Robustness becomes apparent through positive and negative feedback loops, and other controls that restrict genetic output. Generally, positive feedback enhances sensitivity, whereas negative feedback dampens noise and rejects changes. Robustness is an inherent quality strongly preferred by evolution.
Modularity
A module is a collection of nodes in a network which has strong interaction and a common function. Modularity functions to sustain the robustness of the entire system by minimizing damage to specific parts.
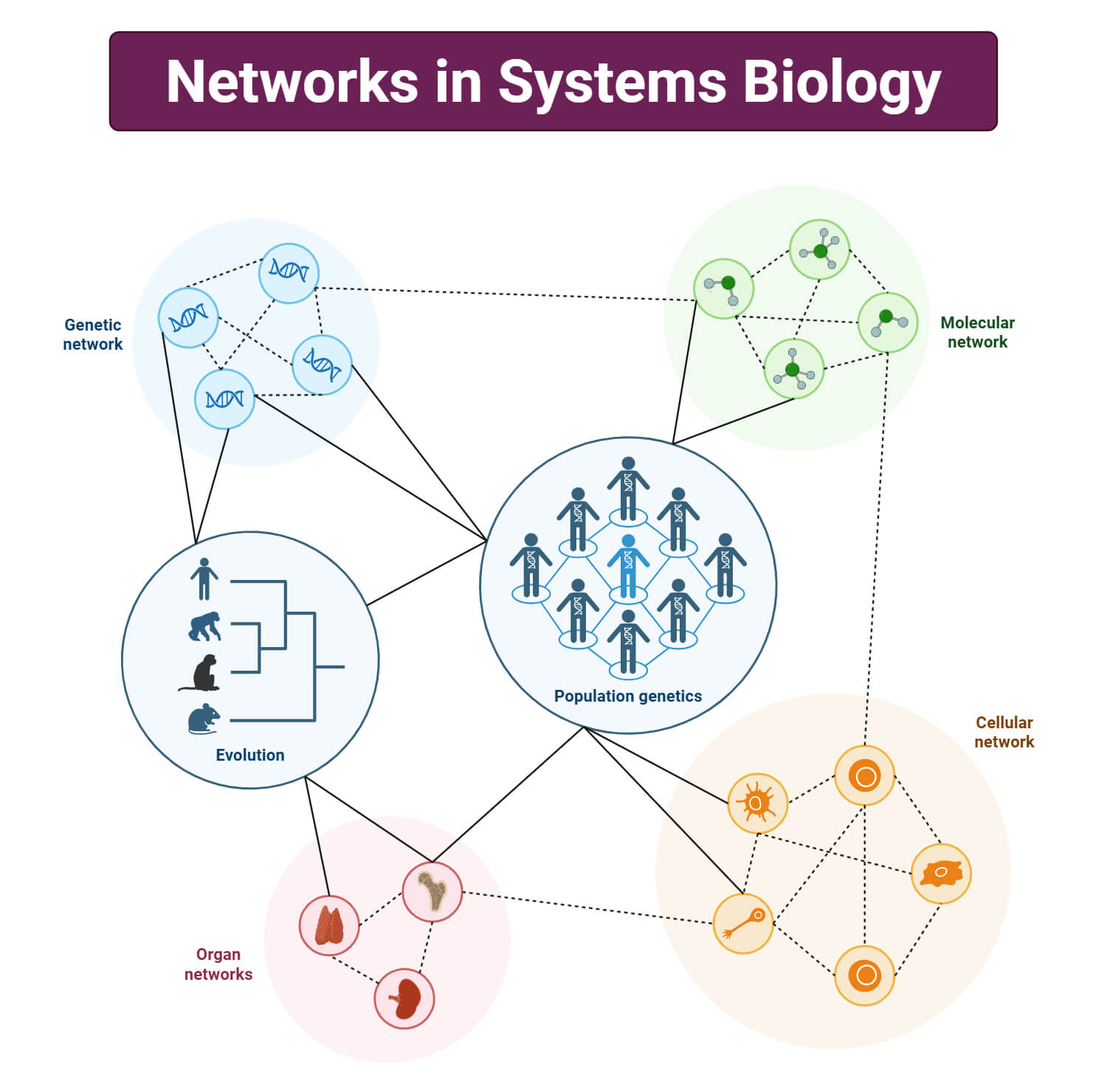
Networks in Systems Biology
Systems biology is made up of networks, filled with nodes consisting of genes, proteins, and their interactions, called edges. The system is built mimicking the machineries of living beings, with networks of molecular and cellular interactions that integrate and communicate across multiple scales. Networks are computable and provide knowledge of how the system is organized. In terms of cellular biology/biochemistry, nodes are cellular components, and edges are interactions, relationships, and reactions between nodes. Often, the edges between the nodes are conditional. To elaborate, in the figure below, the expression of Gene C can be induced by both Gene A and Gene B, with varying probabilities. The incorporation of a computerized network helps us to form hypotheses and provide dynamic biological studies in understanding the complexities of life.
Different Types of Networks
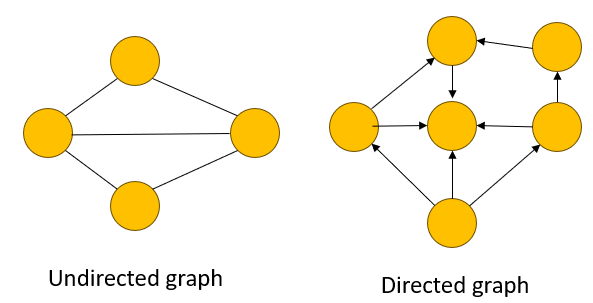
Undirected graphs
In undirected graphs, the direction of the edges is not specified. These kinds of networks are usually constructed through high-throughput data, with only partial knowledge of cell biological systems.
Directed graphs
In directed graphs, the networks are specified, restricting the capability to traverse the graph or direct signal flow. The relationship of the network, however, is not specified.
Sign-specific Directed graphs
Sign-specific directed graphs comprise different kinds of signs, each with a meaning, such as stimulatory relationships and inhibitory relationships. For example, in the figure below, nodes are genes represented as dots, and edges represent the regulation of genes, either by induction or inhibition.
Key Terms in Systems Biology
Node degree (k): The node degree (k) is the number of edges in a network.
Degree Distribution P (k): The number of connections spread across the network is the degree distribution P (k). It is given as:
P k=N (k)N
Where P (k) is the degree distribution,
N (k) is the number of edges.
N is the total number of nodes in the network.
Clique: In a network, a clique is the region of a network where every node is connected to every other node.
Clustering coefficient Ci: Clustering coefficient (Ci) measures the closeness/cohesiveness of nodes to being in a clique. Mathematically, it is given as:
Cv=2ek (k-1)
Where k is the degree of node v,
e is the number of edges between the k neighbors of v, and
k (k – 1) / 2 is the maximum number of edges possible between the neighbors.
When C (v) = 1, a perfect clique, i.e., all neighbors are connected, whereas C (v) = 0 means that none of the neighbors are connected.
Graph distance: The graph distance refers to the shortest path between two nodes in a graph.
Logic Gates in Systems Biology
Logic gates are simple switching circuits that determine if an input signal can pass through to the output. In systems biology, logic gates govern the flow of information between multiple components to reach an output. It can be coordinated with different proteins interacting to change activity, expression levels, or even enzyme concentration through DNA/protein interactions. Systems biology includes a promoter regulated by one activator or by a repressor. Some of the logic gates in systems biology are as follows:
- The amplifier-gate represents a simple activation process.
- The NOT-gate represents transcriptional regression.
- The AND-gate consists of an inducer B that activates the promoter.
- The OR-gate requires one or more inducers to activate the promoter.
Bistability
When a gene is switched ON, it remains locked ON for a relatively longer period; this is known as a bistable switch. Bistable systems can be triggered by an external stimulus, leading to stimulus-dependent variation in cellular machinery.
A prominent example of bistability is the bistable switch of B. subtilis, which controls the competence of the bacterium. Here, the DNA-binding protein ComK forms a positive feedback loop, which activates approximately 100 genes, including its own.
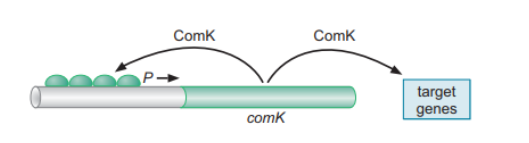
Figure 5: Mechanism of Competence in B. subtilis. Source: (Pederson, 2015)
Conclusion
Systems biology is a growing field with immense potential. Its powerful framework helps in understanding the dynamic network of the cellular and molecular relationships in life. Systems biology can be applied to real-world problems in synthetic biology, drug discovery, and medicine.
References
- Aderem, A. (2005). Systems Biology: Its Practice and Challenges. Cell, 121(4), 511–513. https://doi.org/10.1016/j.cell.2005.04.020
- Kolch, W., Fey, D., & Ryan, C. J. (2018). Systems biology: Old news or new stimulus for biochemistry. Essays in Biochemistry, 62(4), 483–486. https://doi.org/10.1042/EBC20180002
- Pederson, T. (2015). Molecular Biology of the Gene: By James D. Watson: W. A. Benjamin (1965): New York, New York. FASEB Journal: Official Publication of the Federation of American Societies for Experimental Biology, 29(11), 4399–4401. https://doi.org/10.1096/fj.15-1101ufm
- Silva-Rocha, R., & de Lorenzo, V. (2008). Mining logic gates in prokaryotic transcriptional regulation networks. FEBS Letters, 582(8), 1237–1244. https://doi.org/10.1016/j.febslet.2008.01.060
- Trewavas, A. (2006). A Brief History of Systems Biology. The Plant Cell, 18(10), 2420–2430. https://doi.org/10.1105/tpc.106.042267
- What is Systems Biology? (n.d.). Institute for Systems Biology (ISB). Retrieved April 14, 2025, from https://isbscience.org/what-is-systems-biology/
