Molecular chaperones represent a specialized functional class of proteins dedicated to maintaining proteostasis by ensuring the structural integrity of the cellular proteome.

While the primary sequence of a protein dictates its potential native conformation, the intracellular environment is excessively crowded, favouring non-functional kinetic traps and irreversible hydrophobic aggregation. Chaperones mitigate these risks by selectively recognizing and stabilizing unstable, non-native states of polypeptides without becoming part of the final structure.
The introduction of these folding assistants is best framed through their diverse mechanisms: some act as holdases, preventing aggregation during synthesis, while others function as foldases, utilizing ATP hydrolysis to actively drive protein maturation. By providing sequestered environments for folding or facilitating the refolding of heat-denatured proteins, molecular chaperones serve as the cell’s primary defense against proteotoxicity and the various conformational diseases associated with protein misfolding.
Biological Roles of Molecular Chaperones
Molecular chaperones are indispensable mediators of cellular proteostasis, functioning as specialized protein complexes that oversee the structural integrity of the proteome. Their primary biological role begins at the ribosome, where they facilitate the de novo folding of nascent polypeptides by shielding exposed hydrophobic regions, thereby preventing the formation of non-functional kinetic traps or insoluble aggregates. Beyond initial synthesis, chaperones act as critical molecular shields during environmental flux; for instance, Heat Shock Proteins (HSPs) are upregulated during thermal stress to stabilize denatured proteins and catalyze their refolding into active conformations.
Furthermore, they are essential for macromolecular trafficking, maintaining proteins in an unfolded, translocation-competent state to allow their passage through the restrictive membranes of mitochondria and the endoplasmic reticulum. Chaperones also orchestrate the assembly and disassembly of large oligomeric complexes, ensuring that multi-subunit structures like nucleosomes achieve correct quaternary symmetry. In instances where a protein is irreversibly damaged, chaperones perform a triage function, specifically recruiting the E3 ubiquitin-proteasome machinery to facilitate targeted degradation. By balancing these pathways of folding, transport, and disposal, molecular chaperones ensure cellular viability and prevent the onset of conformational diseases.
The Protein Folding Problem: Why Cells Need Helpers
The protein folding problem encapsulates the challenge of how a linear amino acid sequence reliably adopts its unique, functional three-dimensional structure. This process is mathematically daunting; according to Levinthal’s Paradox, a polypeptide chain has so many potential conformations that a random search for the native state would exceed the age of the universe. In the dense, chaotic environment of the cytoplasm, the risk is compounded by macromolecular crowding, where exposed hydrophobic segments of unfolded proteins tend to stick together, leading to irreversible and potentially toxic protein aggregation.
Cells utilize molecular chaperones as essential helpers to navigate this thermodynamic landscape.
Classification of Chaperones: Hsp60, Hsp70, and Hsp90 Families
Most chaperones are classified as Heat Shock Proteins (HSPs) because their expression increases significantly under cellular stress (like heat), which threatens protein stability. While they do not provide the steric information required for a protein to fold, as encoded in the amino acid sequence itself, they provide the protected environment or kinetic assistance necessary for the process to occur efficiently.
| Feature | Hsp70 Family | Hsp60 Family (Chaperonins) | Hsp90 Family |
| Primary Structure | Monomeric 70 kDa | Large double-ring oligomer 60 kDa subunits | Functional homodimer 90 kDa subunits |
| Mechanism of Action | Binds short, hydrophobic peptide segments to prevent premature folding/aggregation. | Provides an isolated Anfinsen cage for a single protein to fold in isolation. | Manages the late-stage folding and conformational activation of specific substrates. |
| ATP Dependency | High, ATP hydrolysis triggers the release of the substrate protein. | High; ATP drives the conformational change of the lid | High; ATP binding/hydrolysis drives a clamp-like closing and opening. |
| Substrate Specificity | Broad; acts on nascent polypeptide chains emerging from the ribosome. | Moderate; restricted by the physical volume of the internal cavity. | Narrow; highly specialized for signal transduction proteins (kinases, steroid receptors). |
| Examples | DnaK (Prokaryotes), BiP, Hsc70 (Eukaryotes). | GroEL/GroES (Prokaryotes), TRiC/CCT (Eukaryotes). | HtpG (Prokaryotes), Hsp90 (Eukaryotes) |
Mechanism of Action: ATP-Dependent Folding Cycles
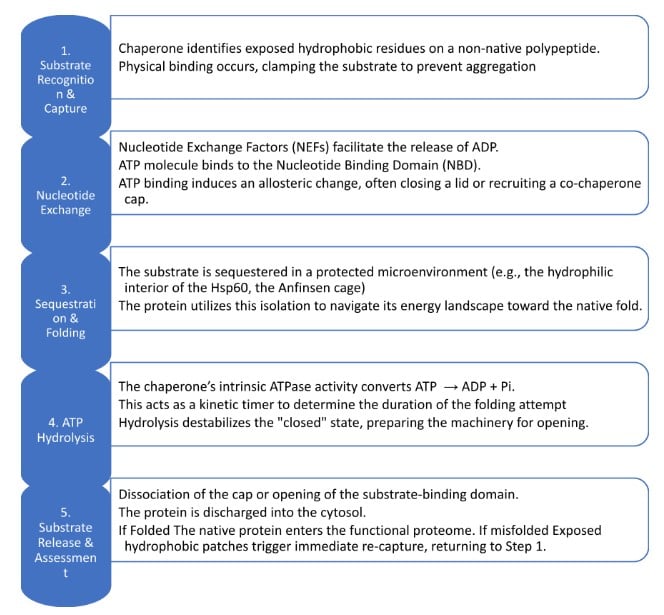
Although Hsp60, Hsp70, and Hsp90 exhibit distinct structural configurations, they operate under a unified thermodynamic principle: the utilization of ATP hydrolysis to drive transition between high-affinity (sticky) and low-affinity (non-sticky) conformational states. This cyclic mechanism effectively suppresses the kinetic trap of protein aggregation by transiently sequestering hydrophobic domains, thereby providing non-native polypeptides with iterative opportunities to achieve their thermodynamically stable native folds.
The Chaperonin System: Structure and Function of GroEL-GroES
The GroEL-GroES chaperonin system of Escherichia coli represents the structural archetype of the Hsp60 family, functioning as an ATP-fueled molecular machine designed to facilitate protein folding through sequestration. Structurally, the complex is a tetradecameric assembly of GroEL subunits arranged into two heptameric rings stacked back-to-back, creating a hollow, double-barrelled architecture.
Each GroEL subunit is divided into three functional domains:
- the equatorial domain, which contains the ATPase catalytic site
- the apical domain, which provides the hydrophobic binding surface for non-native substrates
- intermediate domain, which acts as a flexible hinge to relay allosteric signals between the two.
The system is completed by GroES, a heptameric co-chaperonin that serves as a detachable lid, for the GroEL chambers.
The functional cycle is initiated when a misfolded polypeptide, characterized by exposed hydrophobic patches, binds to the apical rim of an open GroEL ring. This binding event, followed by the recruitment of seven ATP molecules, induces a massive conformational reorganization of the apical domains. These domains rotate and elevate, allowing the GroES cap to bind and seal the chamber. This transformation effectively doubles the internal volume of the cavity and, crucially, alters the chemical environment of the interior walls from hydrophobic to hydrophilic. This transition creates the Anfinsen cage, a protected microenvironment where the sequestered protein is forced to bury its hydrophobic residues, navigating its energy landscape toward the native state in total isolation from the crowded cytoplasm.
The duration of this folding attempt is strictly regulated by the intrinsic ATPase activity of the GroEL ring, which acts as a molecular timer. It takes approximately 10 to 15 seconds for the ATP to be hydrolysed into ADP and inorganic phosphate (Pi). Once hydrolysis is complete in the active ring, a signal is sent to the opposite ring to bind ATP and a new substrate, a phenomenon known as negative cooperativity. This inter-ring communication triggers the dissociation of the GroES lid and the release of the substrate. If the protein has successfully reached its native fold, it remains functional within the cell; if it remains misfolded, its persistent hydrophobic surfaces trigger immediate re-capture by another chaperonin complex for an additional round of folding.
Holdases vs. Foldases: Understanding Different Chaperone Activities
Holdases primarily function as the cell’s first responders to protein instability. Their mechanism is fundamentally passive and ATP-independent, relying on high-affinity binding to exposed hydrophobic residues that would otherwise lead to toxic protein aggregation. By acting as a molecular shield, holdases like sHsps (small heat shock proteins) stabilize unfolded polypeptides during periods of extreme cellular stress, such as thermal fluctuations or oxidative damage. They do not possess the machinery to fix the protein’s shape, instead they maintain the substrate in a folding-competent state, essentially buying time until the cellular environment stabilizes or more specialized machinery becomes available.
Foldases, conversely, are active, ATP-driven machines that catalyze the physical folding of a polypeptide into its functional native state. Unlike holdases, foldases such as the Chaperonins (GroEL/ES) or Hsp70 utilize the energy from ATP hydrolysis to undergo dramatic conformational shifts. These movements allow the chaperone to either sequester the protein within a protected micro environment shielding it from the crowded cytoplasm or to exert mechanical force that unfolds misfolded segments, allowing the protein to try folding again. While holdases are specialized for sequestration and prevention, foldases are the specialized architects that perform the energy-intensive work of structural maturation and refinement.
The Heat Shock Response: Chaperones as Stress Defenders
The Heat Shock Response (HSR) serves as a vital, highly conserved survival mechanism that allows cells to withstand environmental stressors such as extreme heat, oxidative damage, and toxins. When these stressors cause proteins to denature and expose their sticky hydrophobic interiors, the cell prevents lethal aggregation through the rapid up-regulation of Heat Shock Proteins (HSPs), which act as molecular defenders.
This defense strategy follows a sophisticated two-step triage process: first, holdases (such as small HSPs) immediately sequester unfolding proteins to keep them soluble and prevent clumping; subsequently, ATP-dependent foldases like Hsp70 and Hsp90 utilize cellular energy to actively refold these substrates into their native shapes or direct irreparable proteins toward degradation.
This entire protective operation is masterfully regulated by Heat Shock Factor 1 (HSF1), a transcriptional switch that remains tethered and inactive under normal conditions. However, when the concentration of misfolded proteins spikes, chaperones are recruited away from HSF1 to manage the unfolding crisis, thereby releasing the factor to enter the nucleus. Once active, HSF1 triggers a massive production of new chaperones, essentially summoning reinforcements to restore cellular proteostasis and ensure the cell survives the period of instability.
Chaperones in Protein Degradation: The Ubiquitin-Proteasome Link
In the intricate make-up of the cell, molecular chaperones function as quality control inspectors that determine whether a protein should be salvaged or destroyed. This critical transition from folding assistance to disposal, known as chaperone-mediated degradation, serves as the vital link between protein maintenance and the Ubiquitin-Proteasome System (UPS). When a polypeptide remains stubbornly misfolded despite repeated intervention by foldases, chaperones such as Hsp70 and Hsp90 transition from facilitators to reaper intermediaries. They recruit specialized E3 ubiquitin ligases, notably CHIP (Carboxy-terminus of Hsp70-interacting protein), which physically bridges the chaperone-bound substrate to the cell’s ubiquitination machinery.
Once this connection is established, the E3 ligase catalyzes the attachment of a polyubiquitin chain onto the defective protein. This tag provides the necessary signal for the 26S proteasome, a barrel-shaped proteolytic complex, to recognize, unfold, and disassemble the protein into its constituent peptides. By balancing the refold or degrade decision, chaperones act as the ultimate arbiters of proteostasis, ensuring that the cellular environment remains free of toxic aggregates and populated only by functional, correctly folded proteins.
Chaperonopathies: The Role of Chaperones in Neurodegenerative Diseases
Chaperonopathies represent a diverse class of disorders where molecular chaperone dysfunction or deficiency directly contributes to disease pathology, playing a particularly central role in the progression of neurodegenerative diseases like Alzheimer’s, Parkinson’s, and Huntington’s. In these conditions, the balance of proteostasis is disrupted as chaperones fail to effectively neutralize misfolded proteins, which subsequently accumulate into toxic, insoluble aggregates such as amyloid-beta plaques or alpha-synuclein fibrils. This failure can stem from genetic mutations in the chaperones themselves preventing them from recognizing target proteins or from the overwhelming sequestration of available chaperones into existing protein clumps, which depletes the cell’s defensive resources.
Furthermore, as the brain ages, the induction of the Heat Shock Response often becomes sluggish, leading to a diminished capacity for foldases to repair damaged proteins or for the Ubiquitin-Proteasome System to clear them. This chaperone exhaustion creates a vicious cycle where the accumulation of protein further impairs the cell’s folding machinery, eventually triggering neuroinflammation and programmed cell death.
Marinesco-Sjögren Syndrome (MSS), a severe neurodegenerative disorder caused by mutations in the SIL1 gene. SIL1 serves as a crucial co-chaperone (a nucleotide exchange factor) for BiP, the primary Hsp70 foldase in the endoplasmic reticulum. When SIL1 is defective, BiP cannot efficiently swap ADP for ATP, stalling its folding cycle and causing an accumulation of misfolded proteins that eventually triggers the death of cerebellar Purkinje cells. Similarly, in Charcot-Marie-Tooth disease (CMT2F), mutations in the small heat shock protein HSPB1 (Hsp27) disrupt its ability to act as a holdase. These mutant chaperones either lose their ability to sequester misfolded neurofilaments or form hyper-stable complexes that trap proteins, leading to the progressive degeneration of peripheral nerves. In both cases, the chaperone’s inability to balance the folding-degradation equilibrium transforms a vital defense mechanism into a source of cellular toxicity.
Therapeutic Targets: Inhibiting Hsp90 in Cancer Treatment
Hsp90 has emerged as a high-priority therapeutic target because it functions as a critical biochemical buffer that protects the unstable proteome of malignant cells. Cancer cells frequently depend on mutated or overexpressed proteins, termed onco-clients (such as HER2, BCR-ABL, and mutant p53), which are structurally fragile and prone to degradation. By acting as a specialized foldase, Hsp90 stabilizes these volatile proteins, enabling tumor cells to thrive under metabolic stress and bypass apoptotic signals. Consequently, inhibiting Hsp90 provides a unique clinical advantage, it starves the cancer cell of its essential structural support, triggering the simultaneous collapse of multiple oncogenic signaling pathways rather than targeting a single mutation.
The mechanism of Hsp90 inhibition in cancer therapy is a targeted disruption of the chaperone’s ATP-driven cycle, leading to the systemic degradation of oncogenic proteins. At its core, the process involves small-molecule inhibitors competitively occupying the N-terminal ATP-binding pocket of Hsp90. This prevents the chaperone from transitioning into its closed active state, effectively halting the molecular clamp motion required to fold and stabilize onco-clients. Without this protective shielding, these structurally unstable proteins are exposed as terminally misfolded, triggering a shift in the cellular quality control machinery from protein folding to disposal.
The subsequent phase of the mechanism is defined by the recruitment of E3 ubiquitin ligases, such as CHIP, which attach polyubiquitin chains to the destabilized client proteins. This chemical tagging marks the orphaned onco-proteins for immediate recognition by the 26S proteasome, where they are unfolded and proteolytically digested. The final outcome is a combinatorial proteotoxic collapse; by disabling Hsp90, the cell is forced to simultaneously purge multiple signaling kinases and transcription factors essential for tumor survival, proliferation, and stress resistance, ultimately driving the malignant cell toward apoptosis.
Conclusion
Molecular chaperones act as the fundamental guardians of proteostasis, employing a sophisticated mechanical spectrum from passive holdases to ATP-fueled foldases to shield the proteome from terminal aggregation. However, when these systems falter, the result is a chaperonopathy, a state where chaperone deficiency or dysfunction drives the progression of fatal neurodegenerative diseases. In conditions like Alzheimer’s or Parkinson’s, the exhaustion of the chaperone pool leads to the accumulation of toxic fibrils, while specific mutations in co-chaperones, such as SIL1 in Marinesco-Sjögren Syndrome, cause catastrophic cellular collapse. Conversely, in oncology, cancer cells hijack Hsp90 to stabilize unstable onco-clients, effectively using chaperones to bypass natural cell death.
Solving these complex pathologies requires a dual-pronged therapeutic approach: proteome-bolstering or proteome-destabilizing. For neurodegeneration, the solution lies in chaperone induction therapy, utilizing small molecules to hyper-activate HSF1 and replenish the cell’s folding capacity, or employing pharmacological chaperones to stabilize specific mutant proteins. In contrast, cancer treatment involves the strategic inhibition of Hsp90, which starves malignant cells of their structural support and triggers the simultaneous degradation of multiple survival pathways via the Ubiquitin-Proteasome System. By mastering the refold or degraded decision-making process, modern medicine can potentially reverse the structural decay inherent in aging and selectively dismantle the resilience of malignant tumors.
References
- Zhao H, Raines LN, Huang SC. Molecular Chaperones: Molecular Assembly Line Brings Metabolism and Immunity in Shape. Metabolites. 2020 Oct 3;10(10):394. doi: 10.3390/metabo10100394. PMID: 33023034; PMCID: PMC7600384.
- Chatterjee, Bhaskar & Puri, Sarita & Sharma, Ashima & Pastor, Ashutosh & Chaudhuri, Tapan. (2018). Molecular Chaperones: Structure-Function Relationship and their Role in Protein Folding. 10.1007/978-3-319-74715-6_8.
- Dill KA, Ozkan SB, Shell MS, Weikl TR. The protein folding problem. Annu Rev Biophys. 2008;37:289-316. doi: 10.1146/annurev.biophys.37.092707.153558. PMID: 18573083; PMCID: PMC2443096.
- Jee H. Size dependent classification of heat shock proteins: a mini-review. J Exerc Rehabil. 2016 Aug 31;12(4):255-9. doi: 10.12965/jer.1632642.321. PMID: 27656620; PMCID: PMC5031383.
- Castelli M, Magni A, Bonollo G, Pavoni S, Frigerio F, Oliveira ASF, Cinquini F, Serapian SA, Colombo G. Molecular mechanisms of chaperone-directed protein folding: Insights from atomistic simulations. Protein Sci. 2023 Dec 25;33(3):e4880. doi: 10.1002/pro.4880. Epub ahead of print. PMID: 38145386; PMCID: PMC10895457.
- Chaperones GroEL/GroES Accelerate the Refolding of a Multidomain Protein through Modulating On-pathway Intermediates Dahiya, Vinay et al. Journal of Biological Chemistry, Volume 289, Issue 1, 286 – 298
- Mattoo RU, Goloubinoff P. Molecular chaperones are nanomachines that catalytically unfold misfolded and alternatively folded proteins. Cell Mol Life Sci. 2014 Sep;71(17):3311-25. doi: 10.1007/s00018-014-1627-y. Epub 2014 Apr 24. PMID: 24760129; PMCID: PMC4131146.
- Jacob P, Hirt H, Bendahmane A. The heat-shock protein/chaperone network and multiple stress resistance. Plant Biotechnol J. 2017 Apr;15(4):405-414. doi: 10.1111/pbi.12659. Epub 2017 Feb 23. PMID: 27860233; PMCID: PMC5362687.
- Figaj, D. (2025). The Role of Heat Shock Protein (Hsp) Chaperones in Environmental Stress Adaptation and Virulence of Plant Pathogenic Bacteria. International Journal of Molecular Sciences, 26(2), 528. https://doi.org/10.3390/ijms26020528
- Höhfeld, J., Cyr, D. M., & Patterson, C. (2001). From the cradle to the grave: Molecular chaperones that may choose between folding and degradation. EMBO Reports, 2(10), 885–890. https://doi.org/10.1093/embo-reports/kve206
- Scalia, F., Vitale, A. M., Santonocito, R., Conway de Macario, E., Macario, A. J. L., & Cappello, F. (2021). The Neurochaperonopathies: Anomalies of the Chaperone System with Pathogenic Effects in Neurodegenerative and Neuromuscular Disorders. Applied Sciences, 11(3), 898. https://doi.org/10.3390/app11030898
