The active site is the region in the enzyme’s (a type of protein) structure, containing amino acid residues, where catalysis occurs. These active sites are 3D crevices or chiral clefts present inside the fully folded protein, comprising 10-20% of the total enzyme structure. There is variation in the active site location within different types of enzymes.
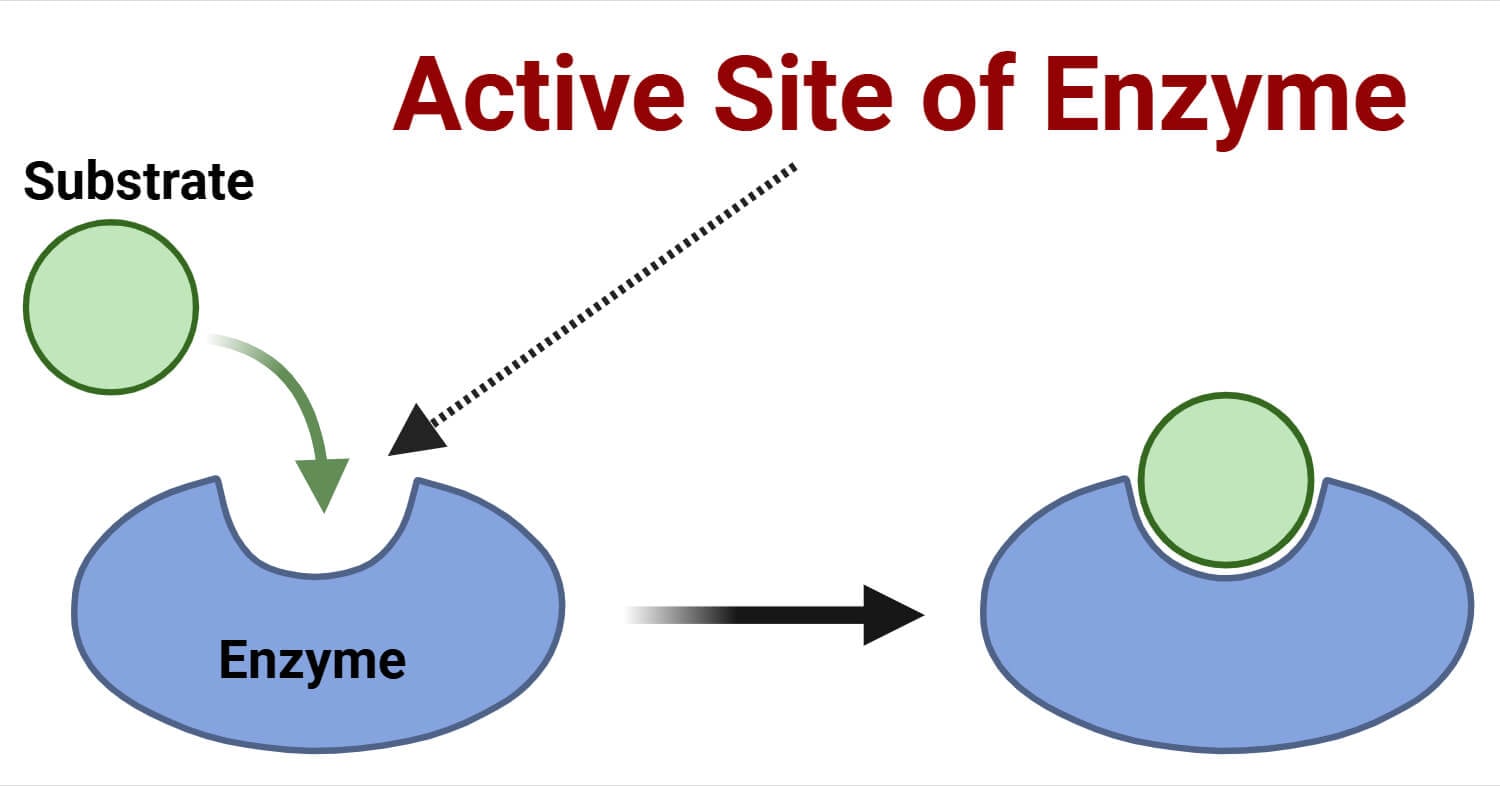
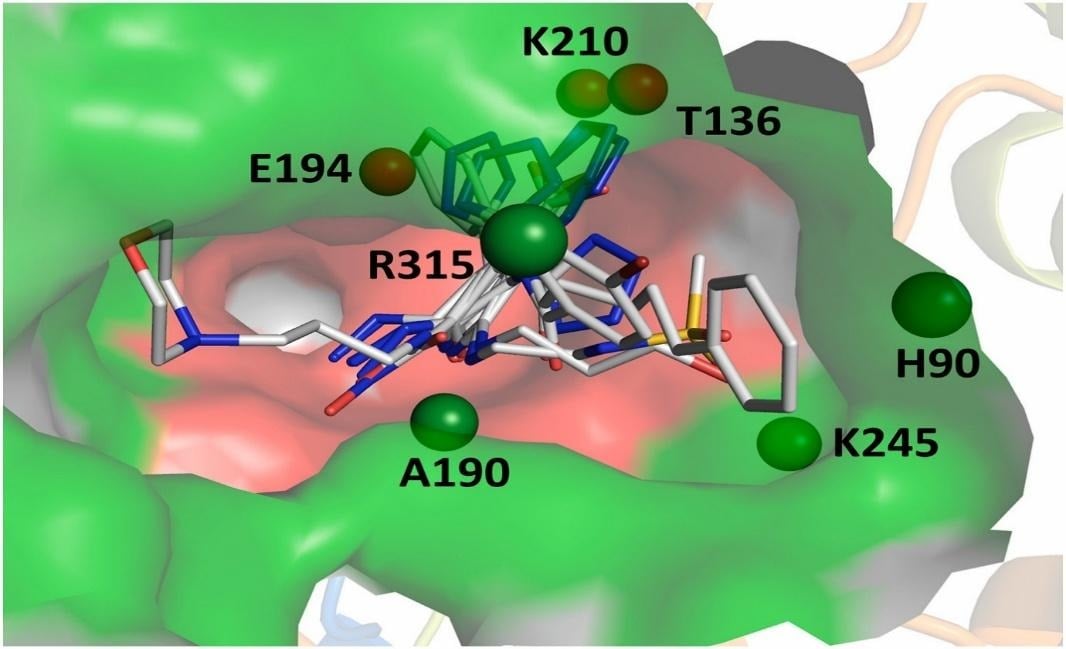
For instance, in the Cytochrome P450 (CYP) enzyme, the active sites are present beside the co-factor, called haem, inside the protein’s core. Similarly, in the C-terminal end of (β/α)8-barrel enzymes, the active sites are present.
Biological Significance of Active Sites
- It is important for catalysis.
- It acts as a protective shield for the attached substrate by desolvation.
- It determines the substrate specificity and the speed of reaction through the arrangement of the amino acid residue.
- It helps in the recognition and binding of the substrate.
- It takes part in chemical bond formation and cleavage.
The 3D Architecture: Clefts, Crevices, and Microenvironments
A deep and large cleft (or crevices) is basically the active site of an enzyme, which is important for the interaction of the protein with other molecules, to perform their biological functions. When a substrate is not bound to the active site, it is filled with water molecules. Usually, we can use geometric criteria, such as identifying the largest cleft in the enzyme’s structure, to locate its active site. However, when there is no single large cleft, we need to analyze the chemical properties of the amino acid residues in all the clefts present.
In natural enzymes, the number of residues, such as polar, hydrophobic, charged or chiral, surrounding the active site provides the microenvironment for enzymatic reactions like transferring electrons, stabilizing transition-state intermediates and binding substrates. When one or more active site residues are mutated, their microenvironment gets altered to achieve the desired enzyme’s properties.
Binding Site vs. Catalytic Site: Understanding the Distinction
The group of 10-20 amino acid residues that interacts with ligands, such as substrates, cofactors, products and inhibitors, related to enzyme function, is known as the binding site. It is also known as a ligand recognition or functional site. It includes any residues that take part in non-bonded interactions. These residues are evolutionarily distinct, altering the enzyme’s substrate specificity. Binding site residues help position the substrate close to the catalytic site correctly, enabling the enzyme to act on a specific substrate.
The catalytic site, a subset of the binding site, is a very specific group of only 3 or 4 amino acid residues that is accountable for the enzyme’s primary biological function, called catalysis. The catalytic site residues remain unchanged throughout evolution. Residues are said to be catalytic if it acts as a nucleophile, alter pKa values of the water molecules or residue involved directly in the catalysis, lower the activation energy of a reaction by stabilizing intermediate or transition state, or activate the substrate by polarizing a bond.
Mechanisms of Specificity: Lock and Key vs. Induced Fit Models
Lock and Key model
In 1894, Emil Fischer proposed an enzyme-substrate binding model known as the Lock and Key model. This model is based on akin to a key fitting into a lock, which means that the shape of the substrate is compatible with the catalytic site of the enzyme. In this model’s mechanism, the existing binding pockets on the free enzymes are completely formed before the substrate is bound to them.
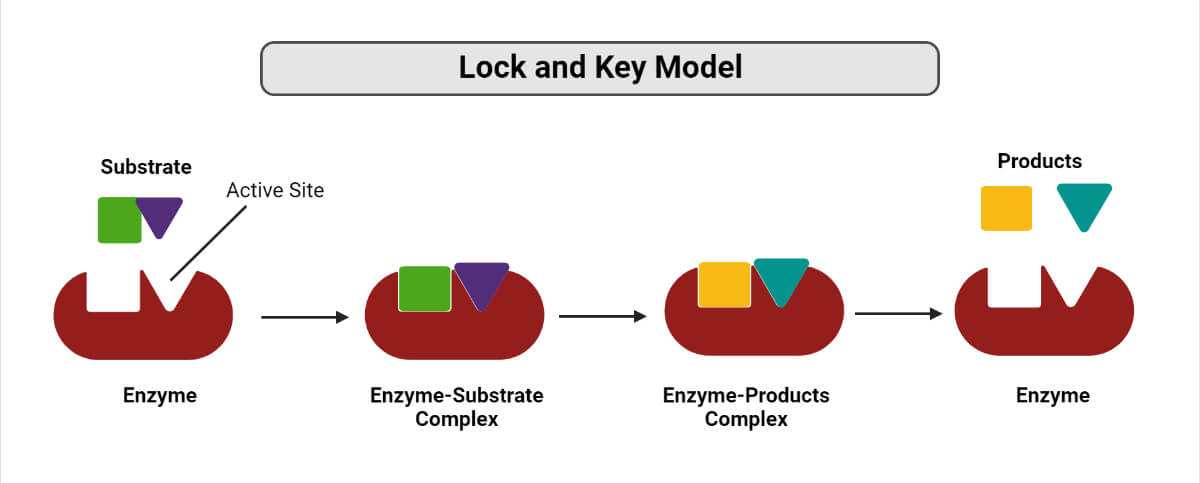
This model is highly efficient as the active sites are already formed to match the transition state of the reaction. Therefore, all the energy is used in stabilizing the transition state. According to this model, both the enzyme and substrate are presented as stiff molecules. However, in biological systems, enzymes have structural flexibility, and depending on the positions and the number of rotatable bonds, various conformations can be adopted by the substrate.
Induced Fit Model
In 1958, Daniel Koshland proposed an enzyme-substrate binding model known as the Induced Fit model. This model is based on akin to putting a hand into a glove. Even when the structure of the substrate is not compatible with the binding site initially, the enzyme undergoes a shape adjustment, allowing it to achieve a stable and tight fit with the substrate. Before the binding of the substrate, the active site of the enzyme is not complementary to the final transition state of the substrate.
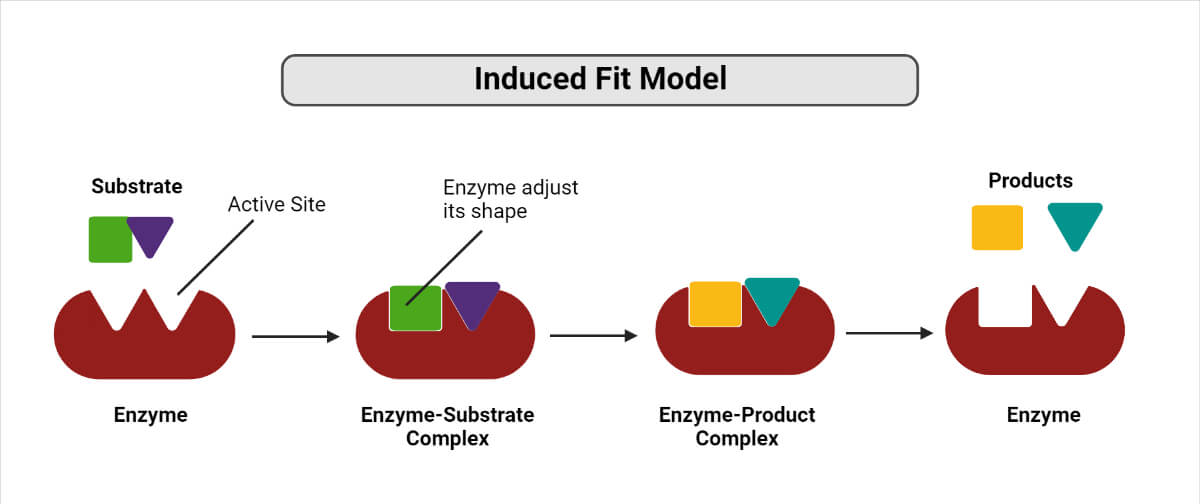
A change in the shape of an enzyme functions as a molecular switch in this model. For instance, in the case of DNA polymerases, when the correct bases are bound to the template DNA strand, it triggers a structural arrangement of catalytic residues that promotes catalysis. However, the mismatched bases will be unable to create the same type of structural arrangement.
The mechanism of the induced fit model is not highly efficient. It is because some of the energy is required to change the enzyme’s structure to perfectly fit the bound substrate. Only a small amount of substrate binding energy is left for the substrate to achieve the transition state.
The Chemistry of Catalysis: Key Amino Acid Residues (The Catalytic Triad)
The catalytic triad consists of the three amino acid residues (Ser-His-Asp) that are present in the enzyme’s active site. Enzymes such as hydrolases and transferases use this catalytic triad to perform chemical reactions. This catalytic triad is a classic design and is present in 2% of all existing proteins. Although these proteins have their own evolutionary origins, they have formed the same catalytic triad. This is the classic example of convergent evolution.
In 1960, David Blow and his colleagues reported the order of α-chymotrypsin by using X-ray crystallography. The amino acid residues, Serine (ser)- Histidine (his)- Aspartate (Asp), plays its specific role in a chemical reaction. Serine, histidine and aspartate act as a nucleophile, a basic residue and an acidic residue, respectively.
Five sequential steps of the catalytic mechanism of Ser-His-Asp: –
- The substrate’s carbonyl group undergoes nucleophilic attack by serine, creating the first tetrahedral intermediate stabilized by the oxyanion hole.
- Breakdown of the tetrahedral intermediate leads to the formation of the acyl-enzyme intermediate between the carbonyl carbon of the substrate and the enzyme.
- Ammonia gets released, leading to the formation of an acyl-enzyme covalent bond.
- A second tetrahedral intermediate is formed by the attack of a water molecule (nucleophile) on the acyl-enzyme.
- Carboxylic acid is released by the breakdown of the second tetrahedral intermediate, and the enzyme is renewed for another cycle.
To function efficiently in various biochemical conditions, some enzymes exhibit variations in the arrangement of catalytic residues. For instance, some LexA repressor proteins and β-lactamases have a lysine (Lys) residue as a base. The threonine (Thr) residue acts as a nucleophile in proteasome catalytic subunits and asparaginases. In arylamine N-acetyltransferases (NATs), cysteine (Cys) acts as a nucleophile.
Stabilizing the Transition State: How Enzymes Lower Activation Energy
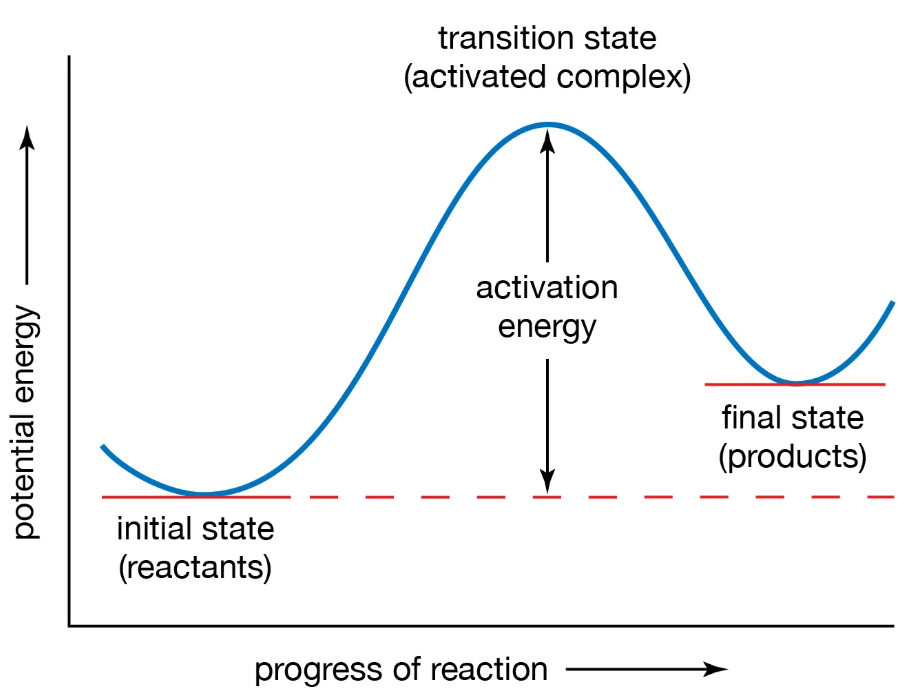
The high-energy, unstable state with a partially bonded structure that lasts for a very short time (femtoseconds) is known as a transition state. The formation and stabilization of this state facilitate the enzymatic reaction move faster.
Enzymes lower activation energy by stabilizing the transition state. It creates a multi-step pathway, thereby avoiding a single large energy barrier. The transition state has electrostatic and geometrical complementarity with the catalytic site of the enzyme. The rapid, small movements of the enzyme help the reactant molecules to position themselves correctly to reach the transition state. Compared to the actual substrate, the transition state binds more tightly due to the favourable enthalpic interactions, such as the strong stabilization of electrostatic forces between the transition state and the enzyme.
Cofactors and Coenzymes: The “Helper Molecules” in the Active Site
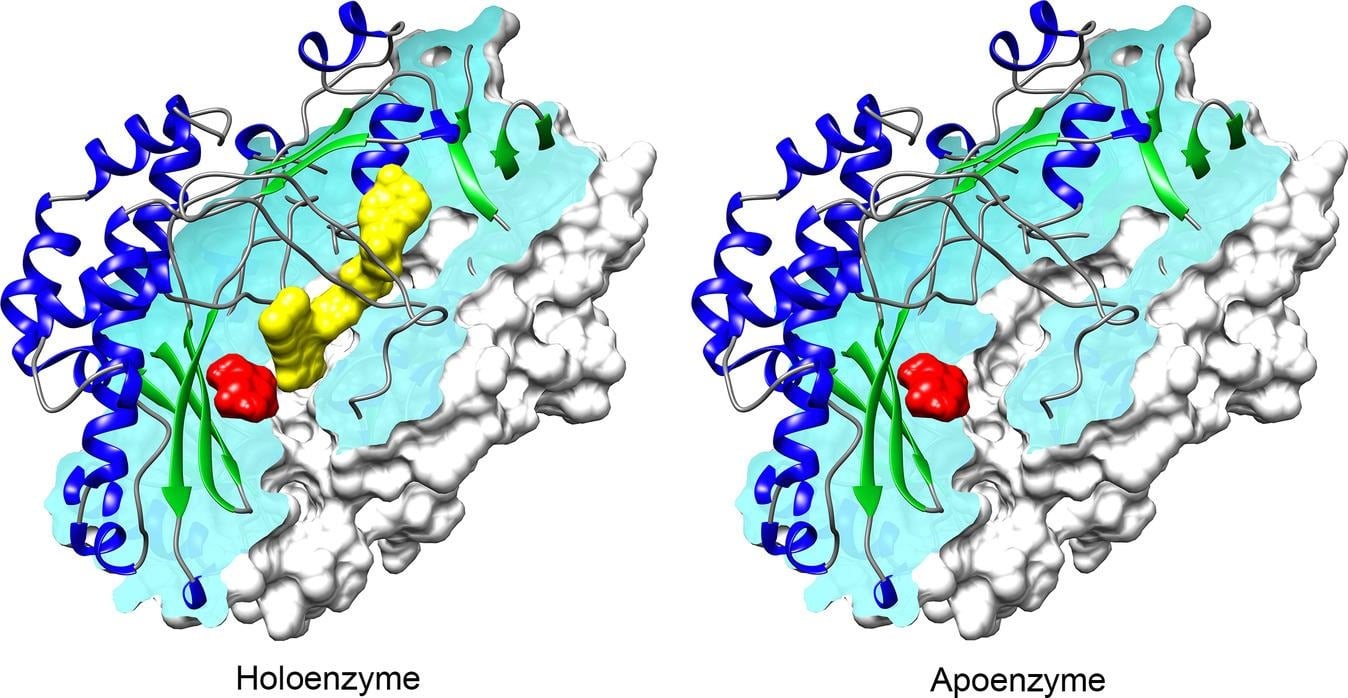
The metal ions or small organic molecules that bind to promote the biological activity of an enzyme are known as cofactors. They are essential for the enzymatic reaction of half of all the enzymes. Cofactors can either be loosely bound coenzymes or tightly bound prosthetic groups. One of the special prosthetic groups, called polypeptide-derived cofactors, is formed by autocatalysis of two or more standard amino acids in the apoenzyme, or from post-translational modifications of standard amino acids [e.g., topaquinone (TPQ)]. Cofactors have regulatory and structural roles by binding away from the enzyme’s active site. They take part in catalysis by binding in the active site.
The simple and small non-protein organic compounds that bind to the enzyme for catalytic reaction are known as coenzymes. At the start of the catalytic cycle, the coenzymes are loosely associated with the enzyme, but they become detached once the reaction is completed. When the coenzyme binds to the inactive protein apoenzyme, it is changed to the functionally active holoenzyme. They enable the transfer of hydrogen species and electron pairs.
Some roles of the cofactors are: –
- They behave as allosteric modulators.
- Within different active sites of the multienzyme complex, the intermediates or substrates are passed due to the flexibility of the cofactor’s structure.
- Organic cofactors such as FMN, SAM and TPP bind to riboswitches to regulate gene expression.
- They help in the proper folding and stability of the protein’s tertiary and quaternary structures.
- They facilitate the formation and cleavage of bonds between heavy atoms.
Environmental Influences: How pH and Temperature Alter Active Site Shape
Temperature alters the active site’s shape by changing its domain position and the flexibility of the molecule. At cryogenic temperature, the substrate-binding cleft is more compact, lacking optimal active structural conformation. For specific proteins such as hen egg white lysozymes (HEWL), high temperature leads to a wide substrate-binding cleft between the α- and β- domains for substrate recognition and catalysis. At an optimal temperature, the active site residues achieve a suitable conformation and position for catalysis. Water molecules in active sites need to be arranged properly for the proper binding of the substrate. At room temperature, the water molecules are flexible and mobile, as compared to the lower temperature conditions where water molecules are rigid.
The structural rearrangement of the enzyme’s substrate-binding pocket is caused by the altered ionization state of its amino acid residues. These types of residues are not found on the active sites, but they are present on the surface of the enzyme, creating repulsive or attractive forces. Water-bridging interaction to stabilize substrate in the active site is hindered by the shift in pH. Hydrogen bonding between acidic/basic Glu residues, and the substrate gets distorted by the enzyme’s pH shift from the optimal pH range. Non-catalytic residues disrupt catalysis by interfering with the substrate position at unfavourable pH.
Enzyme Inhibition: Competitive, Non-Competitive, and Allosteric Effects
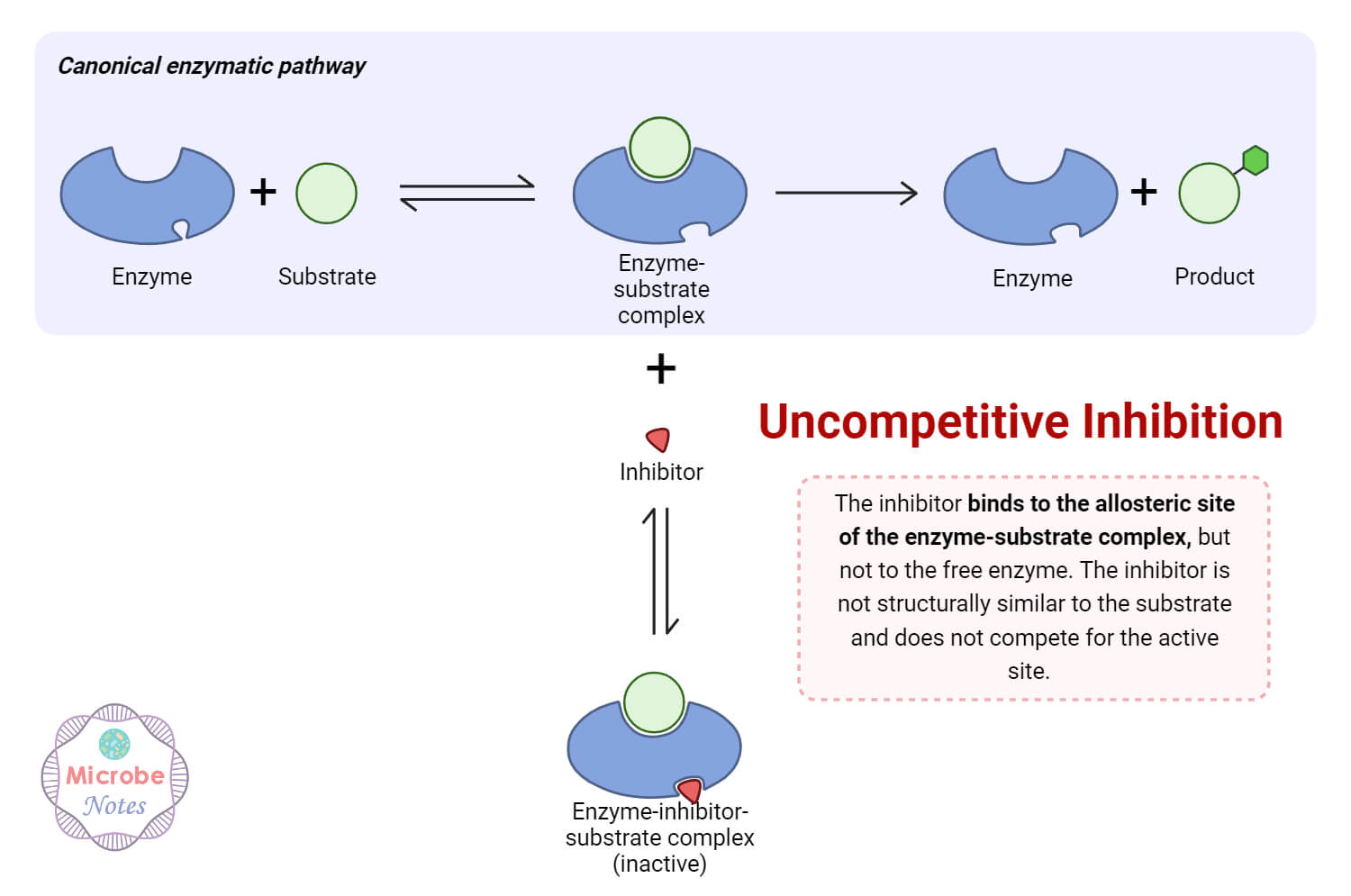
The process in which the inhibitor either binds reversibly or irreversibly to the enzyme to inhibit its activity is known as enzyme inhibition. Approximately 47% of all the current drugs work by inhibiting the enzyme target, even when modern drug design targets cell surface receptors. In irreversible inhibition, a permanent covalent bond is formed by the binding of the ligand to the enzyme’s active site. However, non-covalent interaction exists in reversible inhibition. The types of reversible inhibition are: competitive, uncompetitive, non-competitive and mixed inhibition.
In competitive inhibition, the inhibitor competes with the substrate to bind to the same active site of the enzyme. Only one of them can bind at a time. As the concentration of substrate increases, the degree of inhibition decreases. The binding of the inhibitor does not change maximum velocity (Vmax), but it increases the Michaelis constant (Km).
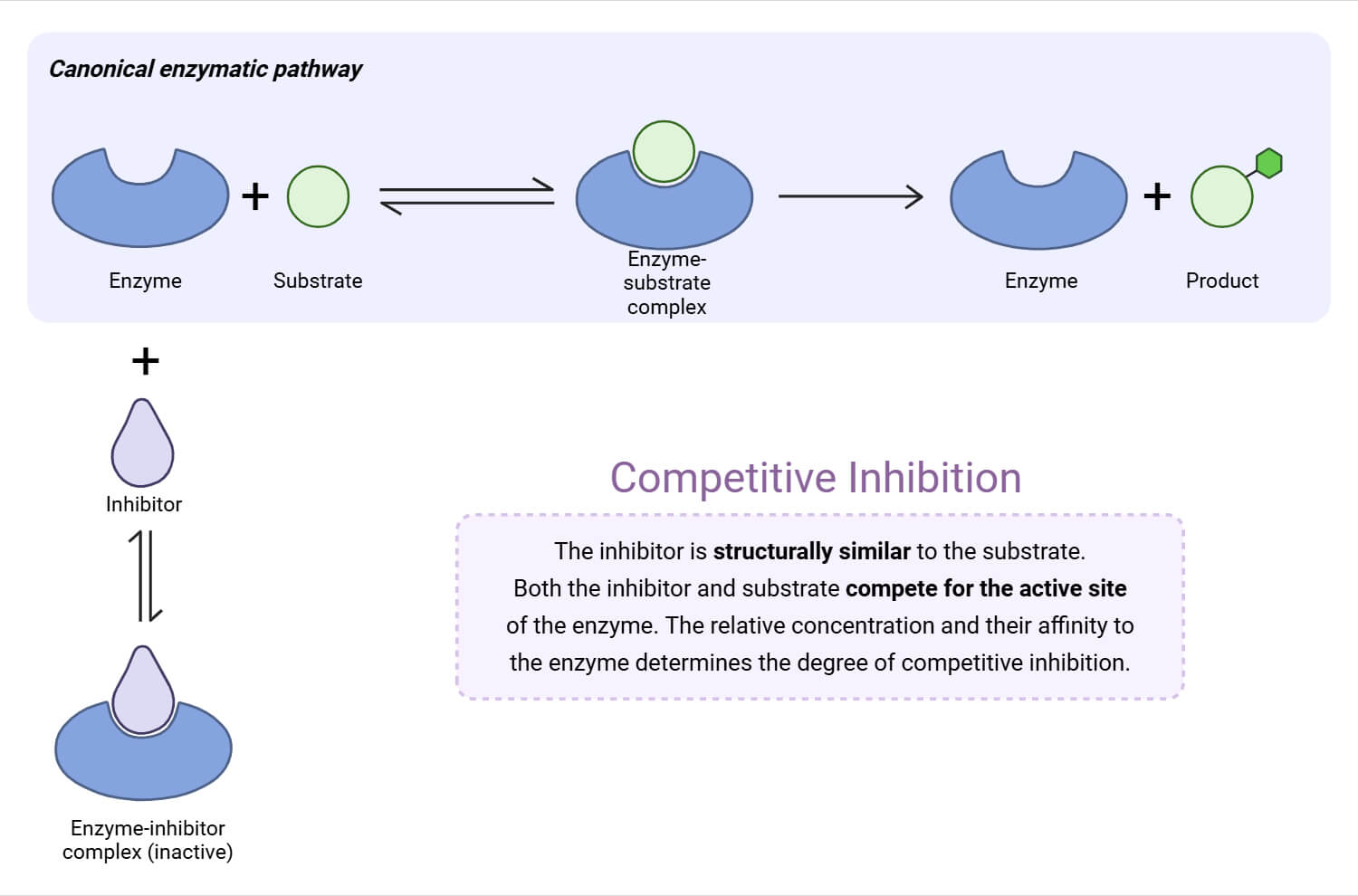
In non-competitive inhibition, instead of binding to the active site of the enzyme, the inhibitor binds to the other site, known as the allosteric site, causing alteration of the enzyme’s shape. Here, Km remains unchanged, but Vmax decreases.
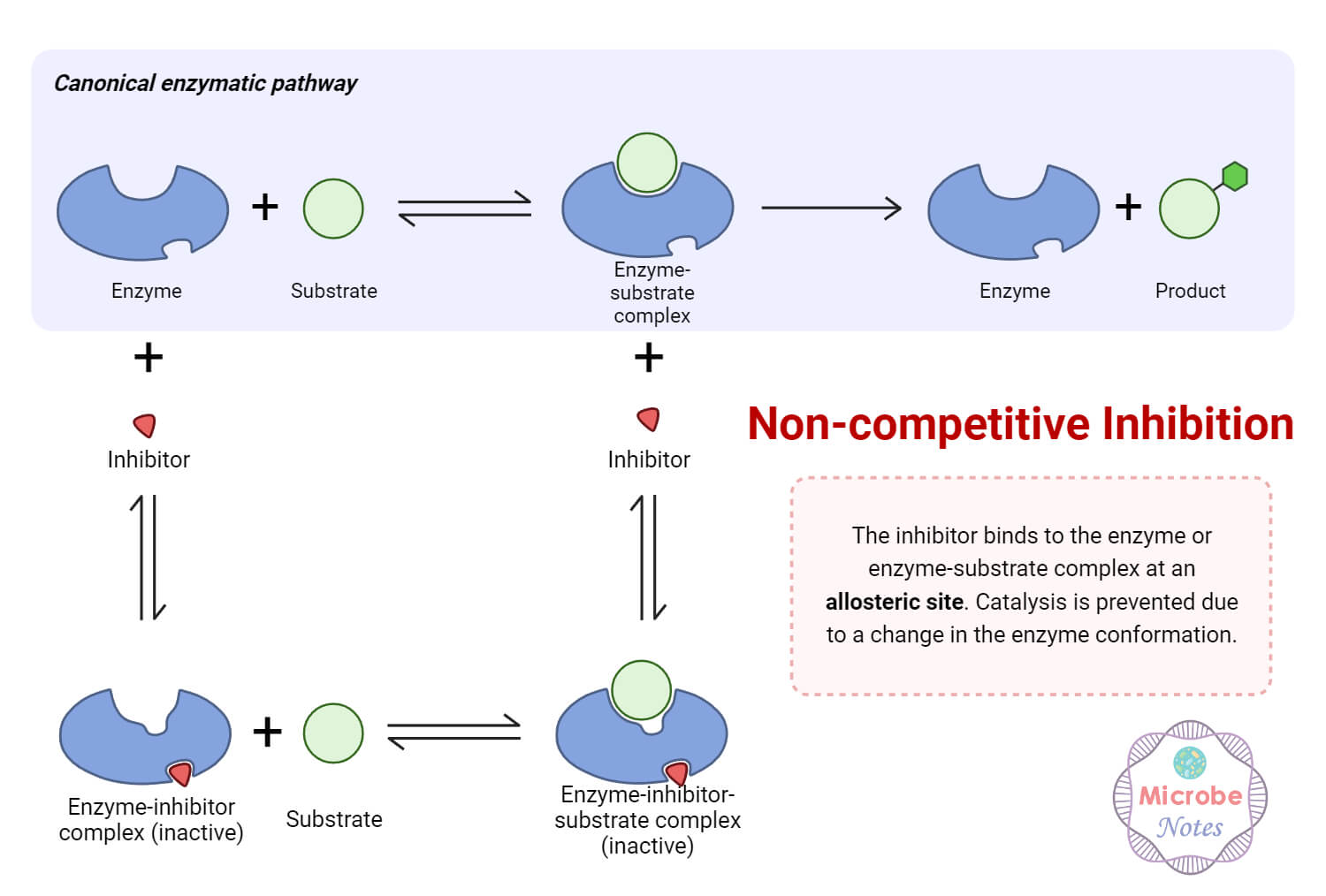
Allosteric inhibition is closely related to non-competitive inhibition. One example of an allosteric effect due to allosteric inhibition is the binding of an inhibitor to the peripheral anionic site (PAS), which is present near the active site of Acetylcholinesterase (AChE), leading to enzyme inhibition through an allosteric mechanism.
Drug Design: Targeting Active Sites for Therapeutic Intervention
The process of developing new therapeutic agents with the help of an understanding of their biological target, such as a protein that has the potential to either inhibit or activate the function, is known as drug design.
Drug design is broadly classified into the following types: –
- Structure-based drug design
- Ligand-based drug design
Other specialized rational drug design approaches are: –
- Covalent drug design
- Fragment-based drug design
- De novo drug design
- Allosteric drug design
The active site contains non-polar groups, hydrogen acceptors and donors that carry out the biological function of the enzyme. By targeting this site, we can design drugs that can bind more strongly than the native ligand. The protein’s 3D structure, determined by X-ray crystallography, provides details about the binding pocket. Among many lead compounds, the promising lead can be identified by using a computational technique known as molecular docking, where the potential lead compounds are docked in the enzyme’s active site and then scored based on their charge distribution and interaction.
Some of the challenges in drug design are as follows: –
- A set of guidelines, called the Lipinski rule of five, must be followed by the lead compounds to evaluate how well the drug can be absorbed and distributed inside the body.
- The accuracy of structure-based drug designing gets lowered if the flexibility of the protein is not considered while the ligand is bound to the bioactive pockets.
- The availability of water molecules in the protein’s binding pocket will either help in stronger or weaker binding of the drug with the protein.
Conclusion
Active sites are regions of an enzyme, formed by clefts and microenvironments, where the substrate binds for catalytic reaction. The active site is composed of both the catalytic site and the binding site. The lock and key and the induced fit models explain how the enzyme binds to its substrate. For the catalytic reaction, the catalytic triad and transition state stabilization are crucial. To enhance the activity of the enzyme, cofactors and coenzymes are needed. The enzyme’s activity is influenced by the alteration of pH and temperature within the active site. For rational drug design, it is crucial to understand the enzyme’s inhibition mechanism and the structure of the active site.
References
- Antunes dos Santos, E., Silva de Oliveira, A., Maria Araújo Rabêlo, L., Ferreira Uchôa, A., & Heloneida Araújo Morais, A. (n.d.). 2 Affinity Chromatography as a Key Tool to Purify Protein Protease Inhibitors from Plants. www.intechopen.com
- Cerqueira, N. M. F. S. A., Moorthy, H., Fernandes, P. A., & Ramos, M. J. (2017). The mechanism of the Ser-(: Cis)Ser-Lys catalytic triad of peptide amidases. Physical Chemistry Chemical Physics, 19(19), 12343–12354. https://doi.org/10.1039/c7cp00277g
- Cristina Oliveira de Lima, V., Piuvezam, G., Leal Lima Maciel, B., & Heloneida de Araújo Morais, A. (2019). Trypsin inhibitors: promising candidate satietogenic proteins as complementary treatment for obesity and metabolic disorders? In Journal of Enzyme
- Dodson, G., & Wlodawer, A. (1998). Catalytic triads and their relatives. Trends in Biochemical Sciences, 23(9), 347–352. https://doi.org/10.1016/S0968-0004(98)01254-7
- Flannelly, D. F., Aoki, T. G., & Aristilde, L. (2015). Short-time dynamics of pH-dependent conformation and substrate binding in the active site of beta-glucosidases: A computational study. Journal of Structural Biology, 191(3), 352–364. https://doi.org/10.1016/j.jsb.2015.07.002
- Ghanem, E., & Raushel, F. M. (2012). Enzymes: The Active Site. In Encyclopedia of Life Sciences. Wiley. https://doi.org/10.1002/9780470015902.a0000714.pub2
- Ghosh, A. K., Samanta, I., Mondal, A., & Liu, W. R. (2019). Covalent Inhibition in Drug Discovery. In ChemMedChem (Vol. 14, Issue 9, pp. 889–906). John Wiley and Sons Ltd. https://doi.org/10.1002/cmdc.201900107
- Gouseti, O., Larsen, M. E., Amin, A., Bakalis, S., Petersen, I. L., Lametsch, R., & Jensen, P. E. (2023). Applications of Enzyme Technology to Enhance Transition to Plant Proteins: A Review. In Foods (Vol. 12, Issue 13). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/foods12132518
- Johnson, K. A. (2008). Role of induced fit in enzyme specificity: A molecular forward/reverse switch. In Journal of Biological Chemistry (Vol. 283, Issue 39, pp. 26297–26301). https://doi.org/10.1074/jbc.R800034200
- Inhibition and Medicinal Chemistry (Vol. 34, Issue 1, pp. 405–419). Taylor and Francis Ltd. https://doi.org/10.1080/14756366.2018.1542387
- Kirschning, A. (2021). Coenzymes and Their Role in the Evolution of Life. In Angewandte Chemie – International Edition (Vol. 60, Issue 12, pp. 6242–6269). John Wiley and Sons Inc. https://doi.org/10.1002/anie.201914786
- Laidler, K.J. (2019, September 23). transition-state theory. Encyclopedia Britannica. https://www.britannica.com/science/transition-state-theory
- Laskowski,’, R. A., Luscombe, N. M., Swindells, M. B., And, ?, Thornton’, J. M., & Thornton, J. M. (1996). Protein clefts in molecular recognition and function. In Protein Science (Vol. 5). Cambridge University Press.
- Małecki, P. H., Fassauer, G. M., Rüger, N., Schulig, L., Link, A., Krylova, O., Heinemann, U., & Weiss, M. S. (2024). Structure-based mapping of the histone-binding pocket of KDM4D using functionalized tetrazole and pyridine core compounds. European Journal of Medicinal Chemistry, 276, 116642. https://doi.org/10.1016/J.EJMECH.2024.116642
- Nam, K. H. (2025). Temperature-Dependent Structural Changes of the Active Site and Substrate-Binding Cleft in Hen Egg White Lysozyme. Crystals, 15(2). https://doi.org/10.3390/cryst15020111
- Polgár, L. (2005). The catalytic triad of serine peptidases. In Cellular and Molecular Life Sciences (Vol. 62, Issues 19–20, pp. 2161–2172). https://doi.org/10.1007/s00018-005-5160-x
- Porter, C. T., Bartlett, G. J., & Thornton, J. M. (2004). The Catalytic Site Atlas: A resource of catalytic sites and residues identified in enzymes using structural data. Nucleic Acids Research, 32(DATABASE ISS.). https://doi.org/10.1093/nar/gkh028
- Ramsay, R. R., & Tipton, K. F. (2017). Assessment of enzyme inhibition: A review with examples from the development of monoamine oxidase and cholinesterase inhibitory drugs. In Molecules (Vol. 22, Issue 7). MDPI AG. https://doi.org/10.3390/molecules22071192
- Rauwerdink, A., & Kazlauskas, R. J. (2015). How the Same Core Catalytic Machinery Catalyzes 17 Different Reactions: The Serine-Histidine-Aspartate Catalytic Triad of α/β-Hydrolase Fold Enzymes. In ACS Catalysis (Vol. 5, Issue 10, pp. 6153–6176). American Chemical Society. https://doi.org/10.1021/acscatal.5b01539
- Sandy, J., Mushtaq, A., Holton, S. J., Schartau, P., Noble, M. E. M., & Sim, E. (2005). Investigation of the catalytic triad of arylamine N-acetyltransferases: Essential residues required for acetyl transfer to arylamines. Biochemical Journal, 390(1), 115–123. https://doi.org/10.1042/BJ20050277
- Schleinkofer, K., Sudarko, Winn, P. J., Lüdemann, S. K., & Wade, R. C. (2005). Do mammalian cytochrome P450s show multiple ligand access pathways and ligand channelling? The EMBO Reports, 6(6), 584–589. https://doi.org/10.1038/sj.embor.7400420
- Schmitt, E., Tanrikulu, I. C., Yoo, T. H., Panvert, M., Tirrell, D. A., & Mechulam, Y. (2009). Switching from an Induced-Fit to a Lock-and-Key Mechanism in an Aminoacyl-tRNA Synthetase with Modified Specificity. Journal of Molecular Biology, 394(5), 843–851. https://doi.org/10.1016/j.jmb.2009.10.016
- Schramm, V. L. (2011). Enzymatic transition states, transition-state analogs, dynamics, thermodynamics, and lifetimes. Annual Review of Biochemistry, 80, 703–732. https://doi.org/10.1146/annurev-biochem-061809-100742
- Sheu, S. H., Lancia, D. R., Clodfelter, K. H., Landon, M. R., & Vajda, S. (2005). PRECISE: A database of Predicted and Consensus Interaction Sites in Enzymes. Nucleic Acids Research, 33(DATABASE ISS.). https://doi.org/10.1093/nar/gki091
- Spassov, D. S. (2024). Binding Affinity Determination in Drug Design: Insights from Lock and Key, Induced Fit, Conformational Selection, and Inhibitor Trapping Models. In International Journal of Molecular Sciences (Vol. 25, Issue 13). Multidisciplinary Digital Publishing Institute (MDPI). https://doi.org/10.3390/ijms25137124
- Staiano, M., Pennacchio, A., Varriale, A., Capo, A., Majoli, A., Capacchione, C., & D’Auria, S. (2017). Enzymes as Sensors. Methods in Enzymology, 589, 115–131. https://doi.org/10.1016/BS.MIE.2017.01.015
- Sun, Y., Yin, S., Feng, Y., Li, J., Zhou, J., Liu, C., Zhu, G., & Guo, Z. (2014). Molecular basis of the general base catalysis of an α/β- hydrolase catalytic triad. Journal of Biological Chemistry, 289(22), 15867–15879. https://doi.org/10.1074/jbc.M113.535641
- Tripathi, A., & Bankaitis, V. A. (n.d.). Molecular Docking: From Lock and Key to Combination Lock.
- Wang, Z., Zhang, H., Gao, Z., Sang, Z., De Clercq, E., Pannecouque, C., Kang, D., Zhan, P., & Liu, X. (2024). Structure-based design and optimization lead to the identification of novel dihydrothiopyrano[3,2-d]pyrimidine derivatives as potent HIV-1 inhibitors against drug-resistant variants. Acta Pharmaceutica Sinica B, 14(3), 1257–1282. https://doi.org/10.1016/j.apsb.2023.11.023
- Wilkinson, I. V. L., Pfanzelt, M., & Sieber, S. A. (2022). Functionalised Cofactor Mimics for Interactome Discovery and Beyond. In Angewandte Chemie – International Edition (Vol. 61, Issue 29). John Wiley and Sons Inc. https://doi.org/10.1002/anie.202201136Yang, L., Zhang, J., Wang, M., Wang, Y., Qi, W., & He, Z. (2023). Probing the effect of microenvironment on the enzyme-like behaviour of catalytic peptide assemblies. Journal of Colloid and Interface Science, 629, 683–693. https://doi.org/10.1016/J.JCIS.2022.09.110
