Polymerase Chain Reaction is a thermal method invented by Kary Mullis in 1983. This method involves amplifying the gene of interest in a sequence. It uses all the required components for DNA replication, such as Taq DNA Polymerase, dNTPs, primers, and buffers.
The principle is simple: with optimal conditions and required components, the gene of interest is amplified in three steps: Denaturation, Annealing, and Extension. Sometimes, the gene of interest may be RNA. In such cases, it needs conversion to DNA before amplification. To address this, a modified method called Reverse Transcription Real-Time PCR (RT-qPCR) was developed.

RT-qPCR uses two principles: Reverse Transcription and Quantitative PCR. The sample is RNA. It is reverse-transcribed into complementary DNA (cDNA), which is then amplified. Amplification results can be read in real time with fluorochromes. RT-qPCR enables the analysis of gene expression in model systems with inhibitors, stimulants, siRNAs, or knockout models. It is also used for detecting changes in gene expression before and after RNA-Seq experiments.
Objectives of RT-qPCR
The objectives of this method include:
- To amplify the gene of interest from RNA genome or from messenger RNA by converting it into its complementary DNA form (cDNA).
- To be able to quantify the results in real-time with the use of a fluorescent dye/probe.
- To obtain accurate and rapid results.
- To quantify RNA to study gene expression and regulation
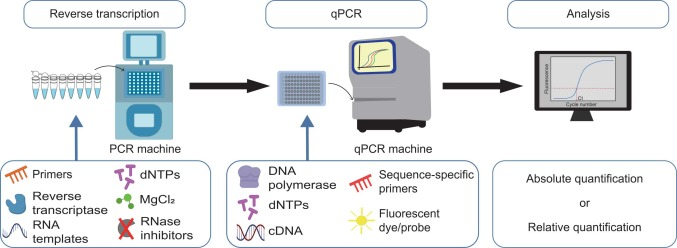
Requirements of RT-qPCR
The components required for RT-qPCR include:
- Primers: They are short stretches of oligonucleotides required for the replication process to begin.
- Deoxynucleoside Triphosphates (dNTPs: dATP, dCTP, dGTP, dTTP)
- Ribonuclease (RNase) inhibitors
- Reverse Transcriptase
- Taq Polymerase
- Fluorescent dyes/probes
- MgCl2 Solution
- Thermocycler
- Template RNA
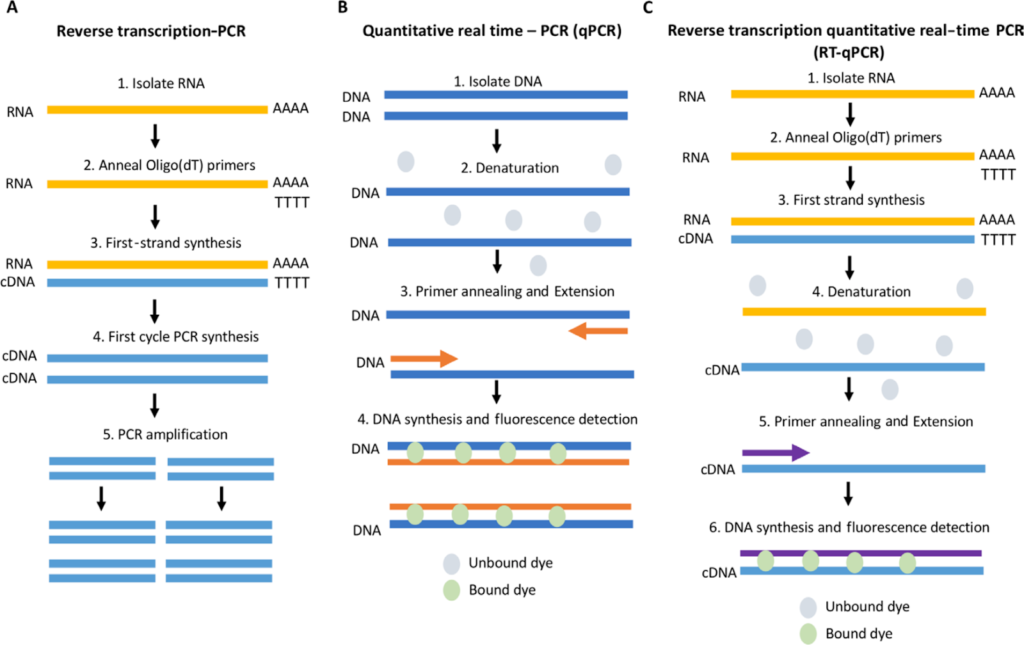
Principle of RT-qPCR
Reverse Transcription- Real Time PCR (RT-qPCR) is a quantitative method that combines the principles of two types of PCR, namely Reverse Transcription PCR (RT-PCR) and Quantitative PCR (qPCR), to analyse and study the RNA of interest. In the former PR, the RNA template is converted into a complementary DNA (cDNA) sequence, which is then amplified using the Taq Polymerase. The reverse transcription of RNA to DNA is done with the help of the enzyme Reverse Transcriptase, thus the name. The cDNA sequence is a single-stranded copy, which is then made into a double-stranded copy with the help of the polymerase. In the latter method (qPCR), the analysis of the PR amplification is done in real time with the help of a fluorescent probe/dye, which is most commonly a hydrolysis-based probe or an intercalating dye. These agents allow the quantification of the PCR product in real time, eliminating the requirement for gel electrophoresis.
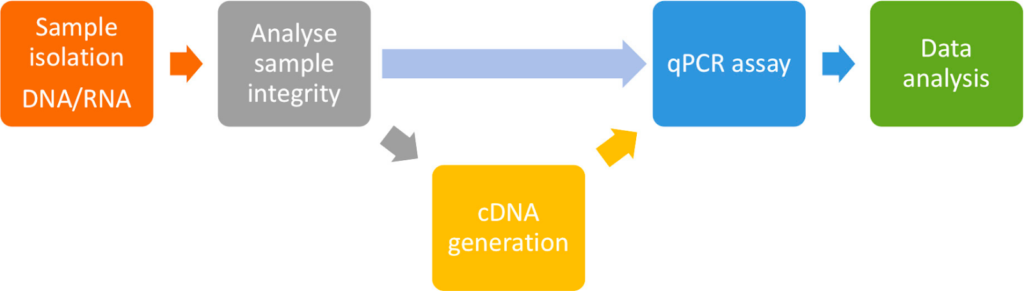
Steps / Protocol of RT-qPCR
The steps for this method include:
- Sample preparation
- Reaction mixture preparation
- Reverse transcription
- Quantitative PCR
- Detection
- Result analysis
1. Sample Preparation
This is the most crucial step of PCR. The quality of the sample isolated should be ensured to be free from contaminants and any chance of degradation. The RNA sample extraction depends on the type to be utilised; extraction kits are widely used for this purpose. The kit can be used for the extraction of mRNA, tRNA, and rRNA; however, it is not useful for the extraction of smaller RNAs such as non-coding RNA (ncRNA: functional RNA transcribed from DNA, but not translated into proteins), and microRNAs (miRNAs: regulate gene expression by inhibiting mRNA translation). The extracted RNA should be free from contaminants, especially DNA, since this cannot be distinguished from ccDNA in the qPCR reaction. Thus, to prevent such contamination, the sample is treated with DNase I, which digests the DNA present. Between mRNA and total RNA, the former provides more sensitivity; however, it is total RNA that has more advantages. It has fewer purification steps, which ensures a quantitative recovery for the template. Unlike mRNA, which requires an enrichment process that may lead to skewed results, Total RNA does not require such a process.
2. Reaction mixture preparation
Here, the components required for the reaction are prepared.
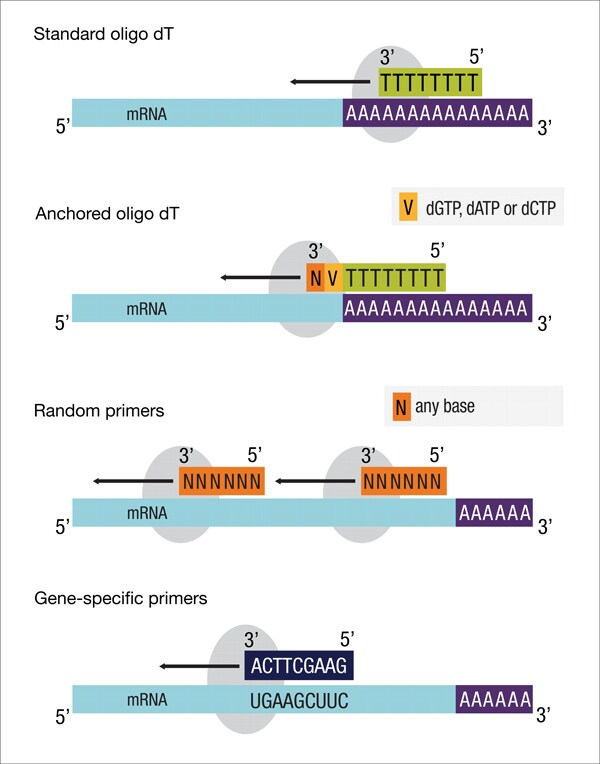
Primers are of three types: Gene-specific primers, oligo (dT) primers, and random primers.
- The gene primers are used to enhance the sensitivity and the specificity of the reverse transcription reaction, as they enhance the sensitivity and specificity of the RT, allowing for targeted gene expression and detection of low-abundance transcripts.
- The oligo dT primers are short sequence stretches of thymidine nucleotides that bind to the poly (A) tail, thereby helping in the transcription of the mRNA.
- The third primers, the random primers, are randomly generated oligonucleotide sequences used to synthesise cDNA.
A combination of the anchored oligo (dT) primers and the random primers can improve the efficiency of the process and the qPCR sensitivity, as it reduces the chances of synthesis of truncated cDNAs.
Reverse transcriptase
The enzyme catalyzes the cDNA synthesis utilising the dNTPs. The ribonuclease H activity of the enzyme is used to remove RNA from the RNA-DNA hybrid during the synthesis of the cDNA. Commonly used enzymes are: Moloney murine leukemia virus reverse transcriptase and Avian myeloblastosis virus reverse transcriptase.
MgCl2+: Provides the Mg2+ for reverse transcription as it is an essential cofactor for the enzyme activity.
RNase Inhibitors: To prevent the template RNA degradation.
DNA Polymerase: Synthesises the new DNA strands in 5’-3’ direction, by addition of dNTPs to the 3’-OH of the annealed primer. Uses the reverse-transcribed cDNA as the template.
Fluorescent dyes/ Probes: SYBR Green (dye) or TaqMan Probes (Probe) are utilised for the quantification of the amplicons.
All these may be added in one tube (one-step) or separately (two-step).
3. Reverse Transcription
This is a 4-step process:
- Denaturation of secondary structures: The isolated RNA samples are incubated at a temperature of 65 °C- 70 °C for a duration of 5-10 minutes. This denatures the secondary structures.
- Primer annealing: Heating of the template and the primer is done, thus allowing primer annealing.
- Incubation: Reverse transcriptase enzyme and the reaction mixture, including the dNTPs, RNase inhibitors, and MgCl2, are incubated for the synthesis of the cDNA. It is done at 37°C to 50°C for 30 to 60 minutes, but may vary depending on the optimal temperature requirements for the reverse transcriptase.
- Inactivation/ Termination: The enzyme activity is inactivated through heating at 70°C to 85°C, thereby terminating the reaction.
4. Quantitative PCR
Here, the cDNA synthesised in the previous step is utilised as the template to produce amplifications. The quantification of the DNA sequence involves two steps:
Reaction setup: The cDNA and the reaction mixture are aliquoted into a qPCR plate. The plates may be 96- and 384-well plates. The mixture contains the DNA polymerase, gene-specific primers, dNTPs, fluorescent dye/probe, and MgCl2.
Thermal cycling: This includes the three steps of a traditional PCR. The cDNA is first denatured through heating at a temperature of 92°C for 15 seconds to 2 minutes, for the strand separation. Next is annealing, and the temperature is reduced to 55-65°C (15–60 seconds)for the gene-specific primers. Finally, the extension is done by DNA polymerase, and the temperature is raised to 72°C (15–60 seconds). In the extension step, the fluorescent dye/probe binds to the newly synthesized DNA, and the fluorescence is measured in real-time. This cycle is repeated for 30-40 cycles, with the fluorescence data collected per cycle.
5. Detection
The detection of the amplicons produced is done in each thermal cycle by collecting the fluorescent data. The detection can be done through two methods:
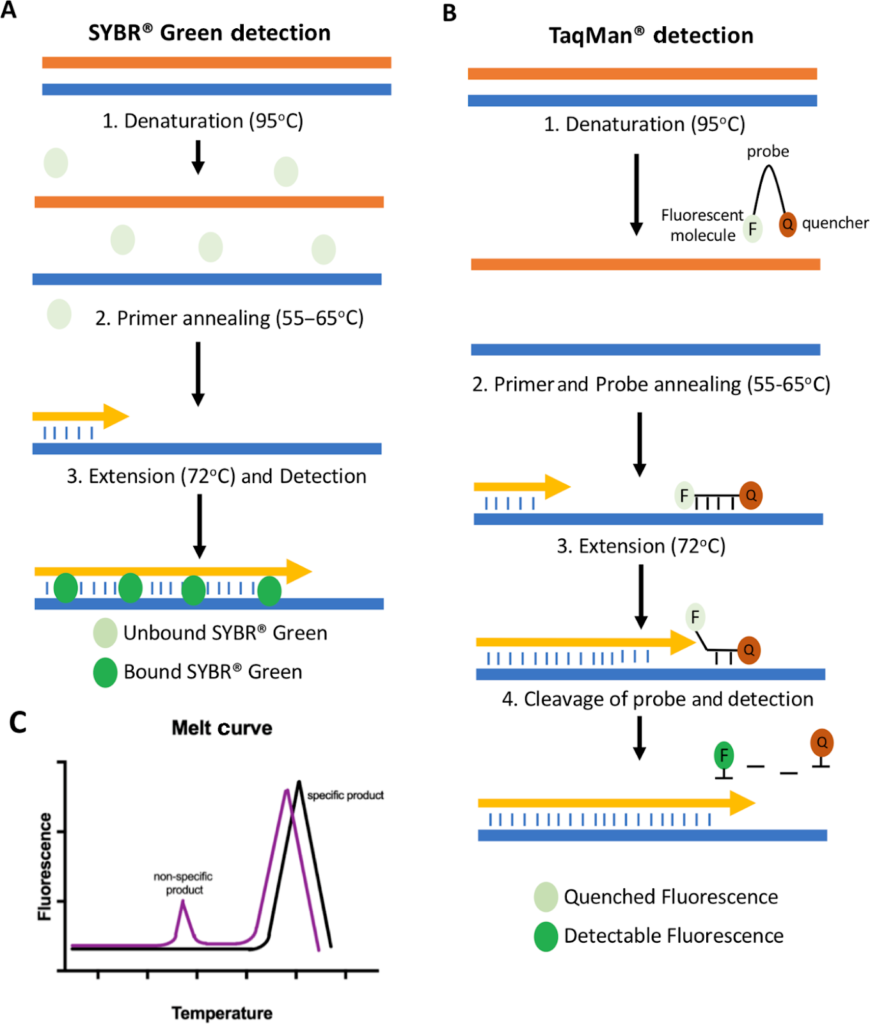
- Dyes: Here, dyes such as SYBR Green are utilised. It is the most common intercalating dye and emits a fluorescent signal upon intercalating with newly synthesised DNA. With increased amplicon production, increased fluorescence is detected. However, this method is non-specific and requires further downstream analysis in the form of a melt curve (A melt curve measures the dissociation of the double-stranded DNA at high temperatures) to ensure the fluorescence detected from the target synthesis. With the use of primers and by validating their specificity, the analysis can be done. These dyes are significantly cheaper to use, and the same dye can be used for multiple different primer pairs. A new generation dye, EvaGreen, allows for melt curve analysis and amplification detection.
- Probes: Most commonly used are hydrolysis probes such as TaqMan probes. They depend on Förster Resonance Energy Transfer (FRET). They hybridize with the target sequences during the PCR reaction. FRET prevents the dye moiety from emitting a signal in the presence of the quencher when the probe is intact. The probe has two components attached to it; to the 5’end, a fluorescent reporter such as the carboxyfluorescein (FAM) moiety is present, whereas at the 3’ end, the quencher molecule, which prevents the signal emission, is present. During the extension step, the 5’ exonuclease activity of the Taq polymerase cleaves the reporter from the quencher, which allows for the fluorescence emission. Probes are more sequence-specific than dyes and thus provide better specificity.
6. Result Analysis
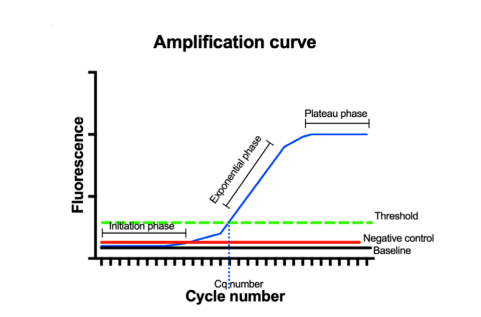
Quantification of the target sequence is done by analyzing the fluorescence emitted by the dye/ probe. The analysis results in an amplification curve with three sections: initiation, exponential, and plateau phases. In the exponential phase, the fluorescence reaches a level higher than that of the baseline, referred to as the threshold level. The cycle threshold value (Ct) represents the number of cycles required for the fluorescent signal to surpass a defined threshold. This indicates the detection of the amplification of the target RNA. Ct value is inversely proportional to the RNA quantities, with a low Ct value indicating high RNA quantities and vice versa.
The RNA quantification is done through two methods:
- Absolute quantification: Here, the exact amount of the RNA is determined using a standard curve with a known concentration of target RNA.
- Relative quantification: This provides a ratio of the expression levels between the samples. Expression levels are obtained by comparing the expression level of a target gene to that of a reference gene.
Types of RT-qPCR
RT-qPCR is of two types;
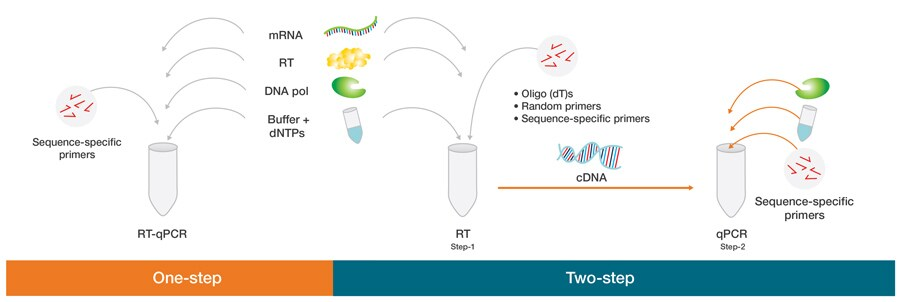
- One-step RT-qPCR: Here, both the reverse transcription and the PCR are done i a single tube and buffer. This is done using a reverse transcriptase along with a DNA polymerase. Additionally, it utilizes sequence-specific primers.
- Two-step RT-qPCR: Here, the reverse transcription and the PCR steps are done in separate tubes, with specific optimized buffers for each, reaction conditions, and priming strategies
| Advantages | Disadvantages | |
| One-step | -Less experimental variation since both reactions take place in the same tube. -Fewer pipetting steps reduce the risk of contamination. -Suitable for high-throughput amplification/screening. -Fast and highly reproducible. | -Impossible to optimize the two reactions separately. -Less sensitive than the two-step because the reaction conditions are a compromise between the two combined reactions. -Detection of fewer targets per sample. |
| Two-step | -A stable cDNA pool is generated that can be stored for long periods of time and used for multiple reactions. -The target and reference genes can be amplified from the same cDNA pool without multiplexing. -Optimized reaction buffers and reaction conditions can be used for each reaction. -Flexible priming options | -The use of several tubes and pipetting steps exposes the reaction to a greater risk of DNA contamination. -Time-consuming. -Requires more optimization than a one-step. |
Examples of RT-qPCR
Some of the different RT-qPCR instrumentation available are:
One Step PrimeScript III RT-qPCR Kit (TakaraBio)
- Synthesis of the template RNA using the PrimerScript Reverse Transcriptase, and the PCR amplification of the cDNA can be done using the Ex Taq Hot Start version. Detection of the amplicons using TB Green-based detection.
- Available in 100 and 500reaction sizes.
- Rapid and cost-effective, and no interruption since it can be done in a single tube.
qScript One-Step RT-qPCR Kit (QuantaBio)
- The reference dye, which has been pre-mixed, is present.
- Cost-effective and efficient method since it follows the one-step RT-qPCR method.
- Utilised for molecular biology requirements and not for diagnosis.
- Contains TaqMan probe for quantification.
qPCR Kits: GoTaq® qPCR and RT-qPCR Systems (Promega)
- Ready-to-use and dye-based kit.
- Compatible with both fast and standard cycling methods.
- Resistant to a wide range of inhibitors.
- Contains both one and two-step RT-qPCR kits.
Applications of RT-qPCR
Some of the applications include:
- Gene expression analysis
- RNAi analysis
- Microarray validation
- Pathogendetection
- Genetic testing
- Disease research
- Detection of viruses causing respiratory infections (eg, Coronavirus)
Advantages of RT-qPCR
- It is a cost-effective method for RNA template analysis.
- Quantitative PCR allows for the detection of the amplicons produced in real time, eliminating the requirement of gel electrophoresis.
- Rapid detection of gene expression changes.
- Analysis can be done for low sample sizes.
Limitations of RT-qPCR
- The presence of inhibitors may hinder the process.
- Quantification of the amplicons requires a standard curve, and thus, there is a chance of error.
- Requires an RNA sample free of contaminants.
- Chances of primer dimer formation
Conclusion
RNA quantification is crucial for understanding gene expression regulation. RT-qPCR is a PCR method that utilizes the principles of both Reverse transcription PCR and Quantitative PCR for the analysis of RNA as the gene of interest and its real-time quantification. It is a rapid and specific method which can be used in various detectiodiagnosisn os, is and molecular biology experiments. Real-time detection and analysis depend on the fluorescent dye/probe, which provides the results in each cycle of PCR. This is a rapid detection method that can be utilized in future public health emergencies.
References
- Adams, G. (2020). A beginner’s guide to RT-PCR, qPCR, and RT-qPCR. The Biochemist, 42(3), 48-53.
- Bong, Dajeong, Jooyeon Sohn, and Seung-Jae V. Lee. “Brief guide to RT-qPCR.” Molecules and Cells 47.12 (2024): 100141.
- https://www.thermofisher.com/in/en/home/brands/thermo-scientific/molecular-biology/molecular-biology-learning-center/molecular-biology-resource-library/spotlight-articles/basic-principles-rt-qpcr.html
- https://www.promega.in/products/pcr/qpcr-and-rt-qpcr/qpcr-kits
- https://www.quantabio.com/product/qscript-one-step-qrt-pcr-kit/
- https://www.takarabio.com/learning-centers/real-time-pcr/overview/one-step-rt-qpcr-kits
