The transition of a protein from a nascent polypeptide chain into a biologically active conformation is an intricate process governed by hierarchical structural levels and thermodynamic principles.
Initially, the primary structure of a linear sequence of amino acids establishes the chemical blueprint for folding. As this chain emerges, local hydrogen bonding between backbone atoms facilitates the emergence of secondary motifs, primarily alpha helices and beta-pleated sheets.
The ultimate three-dimensional geometry, or tertiary structure, is predominantly driven by the hydrophobic effect, where non-polar side chains sequester into a central core to minimize entropic tension with the aqueous environment. This architecture is further stabilized by ionic interactions, van der Waals forces, and covalent disulfide bridges. In complex proteins, multiple subunits may aggregate into a quaternary assembly. Because the native state represents the lowest Gibbs free energy, any deviation or misfolding can result in non-functional aggregates, often implicated in proteopathic diseases. Thus, the fidelity of this transition is essential for maintaining cellular proteostasis and metabolic function.
The Driving Forces: Hydrophobic Effect, Hydrogen Bonds, and Van der Waals
Protein folding is a thermodynamically spontaneous transition driven by the hydrophobic effect, which sequesters nonpolar side chains to maximize solvent entropy. This hydrophobic collapse is stabilized by the directional specificity of hydrogen bonds, facilitating α-helix and β-sheet formation. Furthermore, van der Waals interactions optimize internal packing density. Collectively, these non-covalent forces dictate the protein’s native conformation, balancing structural rigidity with the flexibility required for catalysis.
The Hydrophobic Effect
The hydrophobic effect is the most significant contributor to the thermodynamic stability of the native state of a protein. The hydrophobic effect is less a force and more an entropic imperative. In an unfolded state, water molecules must form highly ordered, cage-like structures called clathrates around nonpolar side chains (like Leucine or Valine) to maintain their hydrogen-bonding network. When the protein folds, these nonpolar groups are buried in the interior. This releases the ordered water molecules back into the bulk solvent, significantly increasing the entropy of the system. This gain in solvent entropy is the primary reason proteins fold spontaneously.
Hydrogen Bonding
Hydrogen bonds act as the structural blueprint of a protein, providing the precise specificity required to define its unique architecture. These interactions arise when a hydrogen atom, covalently bound to an electronegative donor (such as nitrogen or oxygen), experiences an electrostatic attraction toward a nearby electronegative acceptor. This mechanism is the primary stabilizer of secondary structures; for instance, the alpha-helix is maintained by a recurring hydrogen-bonding pattern between the C=O group of one residue and the N-H group of the amino acid four positions downstream. Interestingly, the net energetic contribution of these bonds to overall stability is relatively modest, as the formation of internal bonds requires the prior disruption of existing hydrogen bonds with the surrounding aqueous solvent. Thus, their quintessential role is not merely to drive the collapse of the protein, but to fix it into a specific, functional, and biologically active conformation.
Van der Waals Interactions
Van der Waals interactions serve as the fine-tuning mechanism of protein stability, acting as the final arbiter of structural integrity. These weak, short-range electrostatic attractions arise from transient fluctuations in electron density, which generate the temporary dipoles necessary to stitch atoms together. While individually negligible, their influence is strictly distance-dependent, becoming a dominant stabilizing factor only after the hydrophobic effect has collapsed the polypeptide into a compact state. Within this dense interior where atoms are packed with near-crystalline precision the cumulative effect of these interactions is immense. By maximizing packing density and eliminating unstable gaps or bubbles, these summative sticking forces transform a loose globule into a cohesive, rigid protein core.
The Folding Landscape: Understanding Levinthal’s Paradox
Levinthal’s Paradox delineates the profound incongruity between the astronomical conformational degrees of freedom available to a polypeptide and the millisecond-scale kinetics of biological folding. For a modest 100-residue protein, a random search through an estimated 3^100 conformational states would require a duration exceeding the age of the universe. This mathematical divergence necessitates that folding be viewed not as a stochastic sampling of configuration space, but as a directed transition facilitated by a topographically biased energy landscape.
The resolution of this paradox lies in the Energy Funnel model, which conceptualizes folding as a progressive descent through a hierarchy of increasingly stable intermediates.
As the chain initiates folding, the emergence of local secondary structures and the hydrophobic collapse of the core act as thermodynamic filters, drastically restricting the accessible conformational volume. This creates a steep Gibbs free-energy gradient, funneling the polypeptide toward its unique global minimum (the native state). In this model, the steep loss of conformational entropy is systematically offset by favorable enthalpic gains (via intramolecular bonding) and the entropic liberation of the solvent. Consequently, Levinthal’s Paradox is resolved through a biased search mechanism, a thermodynamic short-cut that ensures rapid, high-fidelity self-assembly within biological timescales.
Thermodynamics of Stability: Gibbs Free Energy (Delta G) and Entropy
The folding of a protein is a complex physical transition governed by the laws of thermodynamics, specifically the relationship defined by the Gibbs Free Energy equation:
Δ G = Δ H − T Δ S
For a polypeptide to achieve its functional native structure, the change in Gibbs Free Energy must be negative, indicating a thermodynamically favourable process. This transition is characterized by a fundamental conflict: the protein undergoes a drastic reduction in conformational entropy as it abandons its flexible, disordered state for a highly constrained three-dimensional architecture. Simultaneously, the formation of non-covalent interactions, such as hydrogen bonding and van der Waals forces, yields a stabilizing enthalpic release. Despite these internal factors, the primary engine of folding is the hydrophobic effect. In an unfolded configuration, non-polar side chains disrupt the solvent structure, forcing water molecules into rigid, ordered clathrate cages. As the protein collapses and buries these hydrophobic residues within its interior, these water molecules are liberated back into the bulk solvent. This release generates a massive surge in solvent entropy that effectively offsets the loss of chain entropy. Notably, the resulting native state is only marginally stable, often by a margin of only 5–15 kcal/mol. This narrow energetic window is biologically essential; it ensures that proteins remain dynamic enough to undergo the conformational shifts required for catalysis, allosteric regulation, and signal transduction without becoming locked in a rigid, immovable state
Assisted Folding: The Role of Molecular Chaperones and Heat Shock Proteins
While the primary amino acid sequence inherently encodes a protein’s native topology, the folding trajectory is significantly impeded by the extreme molecular crowding of the intracellular milieu. In an environment where protein concentrations can reach 300–400 mg/mL, nascent polypeptides face a persistent risk of deleterious intermolecular associations, primarily mediated by exposed hydrophobic moieties. To preserve proteostasis, the cell employs a sophisticated network of molecular chaperones and Heat Shock Proteins (HSPs) that serve as kinetic quality control mediators.
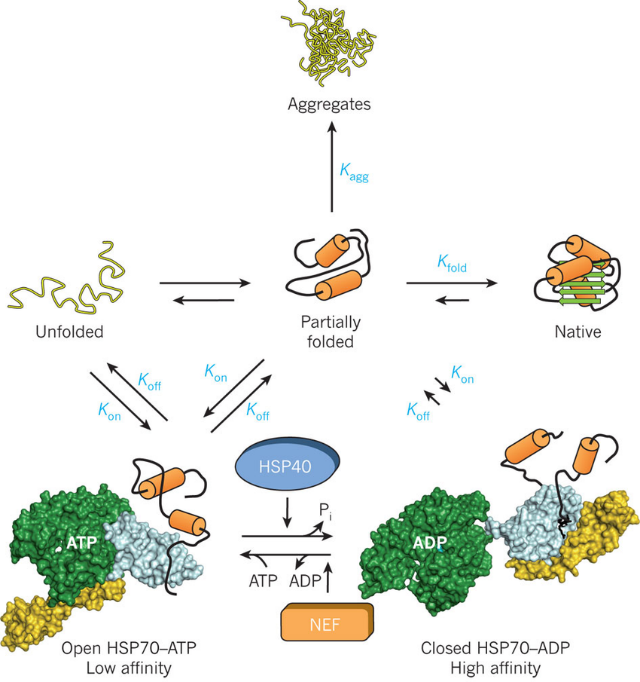
https://www.researchgate.net/figure/The-HSP70-chaperone-cycleHSP70-is-switched-between-high-and-low-affinity-states-for_fig2_51508608
Rather than providing structural templates, these chaperones function as kinetic facilitators that prevent the accumulation of non-functional aggregates and kinetic traps. The HSP70 family operates through a cycle of transient attachment to hydrophobic motifs on emerging chains, effectively shielding them from premature association until the requisite domains are fully synthesized.
In contrast, the HSP60 family (chaperonins) provides a more complex solution by forming a specialized sequestration chamber often referred to as an Anfinsen cage. This cylindrical complex offers a solitary microenvironment that physically isolates a single substrate from the stochastic interference of the cytoplasm.
Driven by the energy of ATP hydrolysis, the chaperonin undergoes orchestrated conformational transitions to capture, encapsulate, and eventually discharge the substrate. This spatial isolation ensures the polypeptide can navigate its energy landscape to reach its global free-energy minimum, safeguarding the proteome against thermal or chemical stressors that would otherwise catalyze widespread misfolding.
Anfinsen’s Dogma: Can Proteins Fold Spontaneously?
Anfinsen’s Dogma, formulated by Christian Anfinsen in the 1960s, serves as the foundational Thermodynamic Hypothesis of molecular biology. Through his seminal work on the reversible denaturation of Ribonuclease A, Anfinsen demonstrated that the three-dimensional architecture of a protein is encoded solely within its primary amino acid sequence. This principle posits that under physiological conditions, the native state represents the unique, kinetically accessible global minimum of Gibbs free energy. In this framework, protein folding is a self-directed process, pre-programmed by the chemical identities and sequence of its residues, requiring no exogenous template to achieve its functional topology.
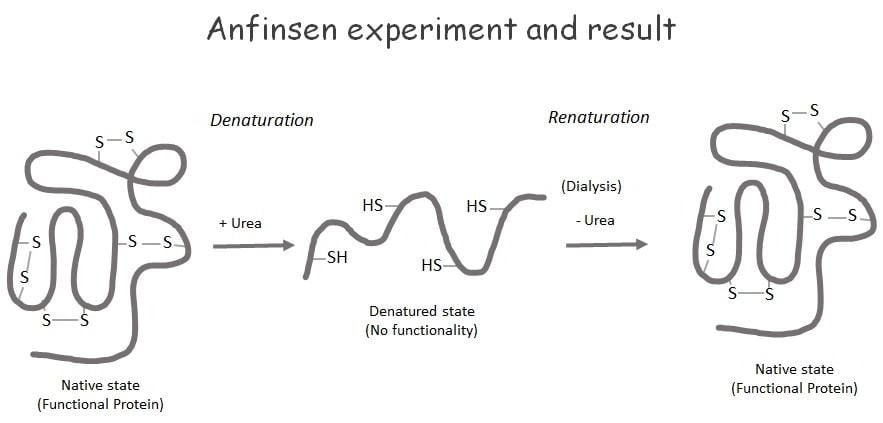
https://www.biologybrain.com/anfinsen-experiment-components-and-conclusion/
Anfinsen’s experiment demonstrated that Ribonuclease A can spontaneously refold after denaturation, proving that a protein’s primary sequence inherently encodes its functional structure. This confirmed the thermodynamic hypothesis, showing that the native state represents the global minimum of free energy at the bottom of the folding funnel.
While Anfinsen’s Thermodynamic Hypothesis suggests a direct, immutable path from sequence to a single stable structure, contemporary biochemistry reveals that the cellular environment is an active participant in the folding process.
| Biophysical Phenomenon | Mechanistic Basis | Deviation from Anfinsen’s Dogma |
| Environmental Determinism | Factors such as pH, ionic strength, and molecular crowding act as external constraints on the folding trajectory. | Suggests that the global minimum is not absolute but is contingent upon the local chemical environment. |
| Post-Translational Modulation | The use of pro-sequences that guide folding before being proteolytically cleaved. | The final functional protein lacks the original sequence information that was required to reach its native state. |
| Metamorphic Proteins | Proteins that adopt two or more distinct, interconvertible folds under physiological conditions | Challenges the unique state rule; one sequence can encode multiple, biologically relevant structures. |
| Prions & Amyloids | Pathological folding where a protein shifts from a functional state to a more stable, aggregated amyloid form. | Demonstrates that the functional native state may only be a metastable local minimum, while the global minimum is a harmful aggregate. |
| Intrinsically Disordered Proteins (IDPs) | Proteins that lack a fixed 3D structure and remain flexible until binding with a partner molecule. | Challenges the idea that a stable structure is even necessary for biological function. |
Factors Affecting Stability: pH, Temperature, and Ionic Strength
Protein stability is a dynamic property determined by the interplay between a polypeptide and its solvent environment. Temperature serves as a primary modulator; rising thermal energy increases atomic vibrations, eventually overcoming the weak non-covalent forces such as hydrogen bonds that maintain the native fold. This typically results in a cooperative melting transition. Interestingly, extreme cold can also induce denaturation as the solvation of hydrophobic groups becomes thermodynamically favoured over their burial in the protein core. Chemical factors like pH and ionic strength further dictate structural integrity. pH fluctuations alter the protonation of ionizable side chains; when the environment deviates from the protein’s isoelectric point (pI), the resulting net charge generates electrostatic repulsion that destabilizes the fold. Similarly, salts affect stability via the Hofmeister series. While low ionic concentrations can stabilize proteins through charge screening, high concentrations may either promote salting out or, in the case of chaotropic agents, disrupt the water-mediated hydrophobic effect to induce unfolding.
Denaturation vs. Degradation: Unfolding Mechanisms Explained
a rigorous distinction must be maintained between the loss of architectural integrity and the chemical destruction of the molecular chain. Denaturation refers to the transition where a protein loses its higher-order quaternary, tertiary, and secondary structures. This process is driven by the disruption of non-covalent interactions such as hydrogen bonds, salt bridges, and van der Waals forces resulting in an unfolded random coil that is biologically inactive. Crucially, denaturation does not break the covalent peptide bonds; the primary amino acid sequence remains preserved. Consequently, this process is often reversible through renaturation, provided the denaturing stress is removed and the environment permits the protein to refold into its global energy minimum.
Conversely, degradation (proteolysis) represents a definitive and irreversible dismantling of the protein. This involves the chemical cleavage of the peptide backbone, typically facilitated by enzymes called proteases or by extreme hydrolysis. Once a protein is degraded, it is reduced to small peptides or individual amino acids, and its original sequence information is lost to the system. While denaturation is frequently a sign of environmental stress, degradation is a high-precision regulatory mechanism. For example, the Ubiquitin-Proteasome System identifies and shreds misfolded or unnecessary proteins, recycling their components to maintain cellular proteostasis and prevent the formation of toxic aggregates.
Protein Misfolding and Aggregation: The Basis of Amyloid Diseases
The transition of proteins from their native, functional states into insoluble aggregates is the pathological hallmark of numerous neurodegenerative conditions, such as Alzheimer’s and Parkinson’s. This process is rooted in proteostasis failure, where environmental stress, aging, or genetic factors cause a protein to lose its specific three-dimensional shape. Once misfolded, the protein exposes hydrophobic regions that trigger a self-assembly process, resulting in the formation of highly organized, beta-sheet rich structures called amyloid fibrils.
These fibrils are defined by a cross-beta architecture, providing extreme stability and resistance to the body’s natural clearance mechanisms. Current research suggests that while mature fibrils are the visible markers of disease, the smaller, soluble oligomeric intermediates are the primary drivers of cellular toxicity, often by disrupting cell membranes. Mapping these aggregation kinetics is vital for designing molecular chaperones or stabilizers that can halt the progression of amyloid diseases at the earliest stage
Measuring Protein Stability: Differential Scanning Calorimetry (DSC) and CD Spectroscopy
Evaluating the structural resilience of a protein is essential for understanding its behavior in biological systems. Differential Scanning Calorimetry (DSC) is a premier thermodynamic tool used to determine a protein’s melting point (Tm). By measuring the heat energy required to unfold a protein, DSC provides a direct readout of thermal stability and the enthalpy of the folding transition. A robust Tm indicates a well-packed, stable protein core, which is less likely to succumb to spontaneous misfolding.
While DSC measures the energetics, Circular Dichroism (CD) Spectroscopy monitors the physical architecture. CD uses circularly polarized light to detect specific secondary structures; for instance, alpha-helices and beta-sheets produce distinct spectral fingerprints in the far-UV range. By tracking changes in these signals across varying temperatures or pH levels, researchers can pinpoint exactly when a protein begins to denature. Used together, DSC and CD offer a high-resolution view of a protein’s stability, providing the data necessary to predict and prevent the onset of aggregation.
Conclusion
The metamorphosis of a polypeptide from a stochastic linear chain into a functional three-dimensional architecture represents a sophisticated orchestration of thermodynamic imperatives and kinetic regulation. As elucidated by Anfinsen’s Dogma, the primary sequence dictates the native state, yet the resolution of Levinthal’s Paradox through the energy funnel model reveals a directed descent toward a global Gibbs free energy minimum. This transition is primarily driven by the hydrophobic effect and refined by the directional specificity of hydrogen bonding and van der Waals interactions.
Within the crowded intracellular milieu, molecular chaperones function as essential kinetic mediators, preventing the formation of pathological amyloids and ensuring high-fidelity folding. The stability of the resulting native state is a precarious equilibrium, sensitive to fluctuations in temperature, pH, and ionic strength. Through analytical frameworks like DSC and CD spectroscopy, the boundaries of this stability are quantified. Ultimately, the synergy between spontaneous self-assembly and assisted folding maintains the cellular proteostasis necessary for metabolic viability and the prevention of proteopathic transition.
References
- Muñoz, V., & Cerminara, M. (2016). When fast is better: Protein folding fundamentals and mechanisms from ultrafast approaches. Biochemical Society Transactions, 44(3), 751–762. https://doi.org/10.1042/BST20160010
- Szilagyi, Andras & Kardos, J. & Osváth, S. & Barna, László & Zavodszky, Peter. (2007). Protein Folding. 10.1007/978-0-387-30379-6_10.
- Muñoz, V., & Cerminara, M. (2016). When fast is better: Protein folding fundamentals and mechanisms from ultrafast approaches. Biochemical Journal, 473(17), 2545–2559. https://doi.org/10.1042/BCJ20160107
- Richard Kullmann, Martina Delbianco, Christian Roth, and Thomas R. Weikl The Journal of Physical Chemistry B 2024 128 (49), 12114-12121 DOI: 10.1021/acs.jpcb.4c06841
- Ivankov DN, Finkelstein AV. Solution of Levinthal’s Paradox and a Physical Theory of Protein Folding Times. Biomolecules. 2020 Feb 6;10(2):250. doi: 10.3390/biom10020250. PMID: 32041303; PMCID: PMC7072185.
- APA 7th Edition: Jiménez, J. S., & Benítez, M. J. (2024). Gibbs free energy and enthalpy–entropy compensation in protein–ligand interactions. Biophysica, 4(2), 298–309. https://doi.org/10.3390/biophysica4020021
- Bhaskar, A. (2021). Heat shock proteins and molecular chaperones: Implications for adaptive responses in the skin. In Heat shock proteins in veterinary medicine and sciences (pp. 145–168). Springer Nature. https://doi.org/10.1007/978-3-030-68112-8_7
- Rose, G. D., Fleming, P. J., Banavar, J. R., & Maritan, A. (2006). A backbone-based theory of protein folding. Proceedings of the National Academy of Sciences, 103(45), 16623–16633. https://doi.org/10.1073/pnas.0606843103
- Masson, P., & Lushchekina, S. (2022). Conformational stability and denaturation processes of proteins investigated by electrophoresis under extreme conditions. Molecules, 27(20), 6861. https://doi.org/10.3390/molecules27206861
- Uddin, M. S., Kabir, M. T., Tewari, D., Mathew, B., & Aleya, L. (2020). Targeting protein misfolding and aggregation as a therapeutic perspective in neurodegenerative disorders. Cells, 9(3), 534. https://doi.org/10.3390/cells9030534
- Characterization of biopharmaceutical stability with DSC [Application Note]. https://www.tainstruments.com/pdf/literature/APN036_Characterisation-of-biopharmaceutical-stability-with-DSC.pdf
