Polymerase Chain Reaction (PCR) is a thermal process invented by Kary Mullis (1983). It is used for amplifying the gene of interest in vivo. PCR requires a machine called a thermocycler, which sets the required temperatures for each step (Denaturation, Annealing, Extension). It produces amplicons for various diagnostic and molecular biology uses. Traditional bulk analysis gives only average values for cell populations, which can lead to inaccurate information about individual cells. Single-cell PCR allows analysis of individual cells and yields more precise results.
Single-Cell PCR is defined as the amplification of the gene of interest or a sequence using the DNA from a single cell. This is one to study mutations, genetic disorders, gene therapy, etc.
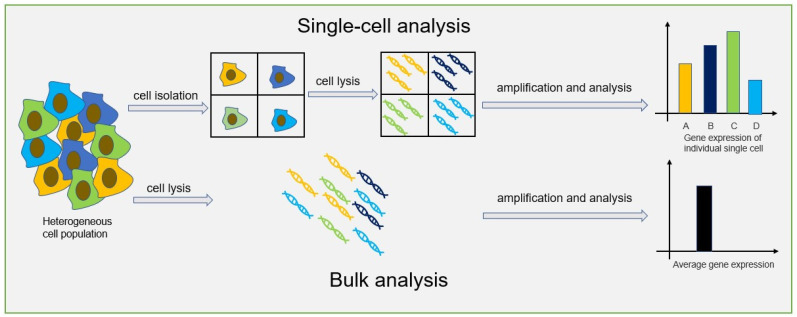
In some cases, it is more practical to study a whole cell instead of a bulk analysis. This is important, for example, in diagnosing genetic disorders from preimplantation embryos, paternity testing, and gene therapy. Single-cell analysis helps us study biochemical reactions and the substances within a single cell. Learning this gives insights into cell functions and their chemical makeup. It also helps explain heterogeneity, biological functions, and disease mechanisms. This type of analysis is useful in cancer, immunology, neurology, and in vitro fertilization.
Objectives of Single-cell PCR (SC-PCR)
The objectives of this PCR method include:
- To study the cells at the individual level
- To isolate and amplify the gene of interest in a single cell for diagnostic and disease research purposes.
- To study the heterogeneity, biochemical, and biological functions of the cell.
- To analyze single cells to understand what happens to organisms at a specific point in time, to understand for disease research.
Requirements for Single-cell PCR
Single-cell PCR requires:
- Isolated cell sample with gene of interest
- Primers
- dNTP’s
- Lysis buffer
- MgCl2
- DNA Polymerase
- Buffers
- Thermocyler
Principle of Single-cell PCR
The principle is similar to that of the traditional PCR method. When provided with optimal conditions in terms of temperature and environment, the gene of interest can be amplified, and multiple copies of the same can be obtained. In Single-Cell PCR, an individual cell is isolated using chemical or physical methods, the most often used method being the monoclonal culture method, where the cell suspensions are serially diluted, resulting in individual cells of interest. Upon isolation, the cells should be lysed to obtain the required DNA for the PCR reaction. The lysates are then subjected to traditional PCR conditions, and amplicons are obtained.
Steps/protocol of Single-cell PCR
The steps of this method include:
- Isolation of the cell
- Preparation of cell lysate
- Reaction mixture preparation
- Amplification
- Post amplification analysis
1. Isolation of the cell
The first step in single-cell PR is the isolation of the cell of interest. Care should be ensured during the isolation, since the cell sample should be free of contaminants and free. Isolation can be done by physical or chemical methods. Some of the methods are:
- Limited serial dilution
This is the most commonly used method for the isolation of individual cells. It is a monoclonal culture method, where the cell suspension is made to undergo a series of dilutions resulting in proper distribution and concentration within the suspension. The final dilution contains only an individual cell. This is a comparatively simple method, and cost-effective; however, it is not considered efficient since achieving a single cell is based on the Poisson distribution (It is a probability distribution to calculate the likelihood of a certain number of events occurring in a fixed interval of time, assuming the events occur independently.)
- Fluorescence-activated cell sorting (FACS)
A method based on flow cytometry, FACS, is a high-throughput cell-sorting technology utilized for single-cell isolation. The cell suspension is labeled fluorescently and pressed into a flow cell, which is then diluted with a sheath fluid to an appropriate concentration. A target operation driving is performed to break the stem into continuous droplets, some of which carry the cells of interest. These cells are then collected using an electrically charged plate. This is an expensive method despite being highly throughput and requires >10,000 cells as input, which is undesirable for rare samples.
- Laser capture microdissection (LCM)
This method accurately separates single cells from tissue samples. The target area on stained tissue sections is chosen and isolated using a laser. It’s comparatively an effective method as the isolation doesn’t damage the tissue structure and also allows for direct isolation of target cells from frozen or paraffin-embedded tissue sections. Since this method relies on a laser capture microdissection platform, it is considered to be a complex and expensive method.
- Manual micromanipulation
Here, manual selection of the individual cells using a microscope is done. In this method, cells of interest can be selected from a prepared cell suspension through microscopic observation. The selected cells are sucked out using a pipette. This process is quite complex and requires highly skilled operators since human interference results in low throughput.
- Microfluidic devices
One of the most widely utilized methods for single-cell isolation is due to its isolation and manipulation abilities for a single cell with microscale and integrated flow channels. The two mainstream microfluidic-based single-cell isolation methods are: Microwell array chip-based methods, and droplet-based methods. This is an advantageous method owing to its high separation throughput and low cost, also because of the fact that it’s easy to operate using commercial instruments. These advantages have made this method the best candidate for single-cell isolation in the future.
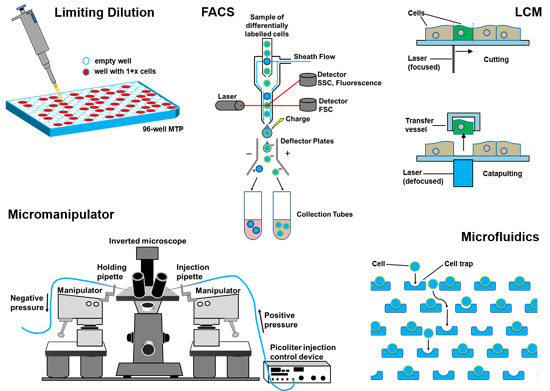
“Different methods of single-cell isolation” Gross, A., Schoendube, J., Zimmermann, S., Steeb, M., Zengerle, R., & Koltay, P. (2015). Technologies for single-cell isolation. International journal of molecular sciences, 16(8), 16897-16919.
| Method | Sample Type | Throughput | Cost | Isolation Efficiency | Automation |
| Limited serial dilution | Dissociated cell suspension | Low | Low | Low | No |
| FACS | Dissociated cell suspension | High | Very high | Low | Yes |
| Micromanipulation | Dissociated cell suspension | Low | Moderate | High | No |
| LCM | Tissue | Low | Moderate | High | No |
| Microfluidic | Dissociated cell suspension | Very high | High | Moderate | Yes |
2. Preparation of cell lysate
Once the single cells have been isolated, they are lysed. This is done to obtain the gene of interest from the individual cell. An adequate cell lysis method is required to obtain accurate PCR results. Lysis can be done through either chemical or mechanical means, with the chemical being relatively mild and the former being more intense, which may lead to DNA breakage in cells. Thus, a suitable method should be chosen, taking cell type, nucleic acid stability, and compatibility with downstream reaction into consideration, with chemical lysis being the best method.
One of the chemical methods involves the use of a lysis solution. The cells are drawn into a Pasteur pipette through capillary force (it should be ensured that only a single cell has been drawn into the pipette, which can be done through a microscope), and the sample is then submerged in the lysis solution and expelled. The solution with the cell is then briefly centrifuged to ensure the cell is at the bottom.
The cell lysates were added to PCR tubes containing a lysis buffer. The buffer may be an SDS-containing lysis buffer (0.005% SDS with 400 ng μl−1 Proteinase K), TritonX-100-containing lysis buffer (1.0% TritonX-100 with 400 ng μl−1 Proteinase K). This was then frozen in liquid nitrogen for at least 10minutes, followed by incubation at a high temperature of 60 °C for 30 minutes. Then a denaturing step at 95 °C for 10 minutes to inactivate the Protease K, and the lysates were used (If not utilized, they should be stored in -80 °C).
3. Reaction mixture preparation
The reaction mixture (45μl) was prepared by
- 5 μl of 10x PCR buffer
- Taq DNA Polymerase
- Each dNTP at a concentration of 0.2mM
- Primers
- 5μl lysate
After preparation, the mixture can be used for SC-PCR.
4. Amplification
This follows the same steps of traditional PCR; Initial denaturation at 95 °C for 2minutes. For the subsequent 30cycles, denaturation at 95 °C for 30 seconds, Annealing of the primer at 55 °C for 30 seconds, and finally Extension using the dNTPs for 60 seconds. After the 30 cycles, the product is incubated at 72 °C for 7 minutes for a final extension.
5. Post-amplification analysis
The PCR products can be analyzed using agarose gel electrophoresis. 10-20μl ofr the product was loaded onto the agarose gel (1.2%) containing an intercalating agent (Ethidium Bromide) and then electrophorized at 100 V. After the run, the gel was visualized under UV illumination. For further nucleotide sequence analysis, computational software (Eg, CLUSTAL X and BLAST) can be utilised.
Types of Single-cell PCR
Single-Cell PCR is not of different types; rather, they are coupled with other PCR methods. Some of them are:
- Digital Single-Cell PCR: The cell lysates are loaded onto microchambers, each of which will undergo subsequent PCR reactions.
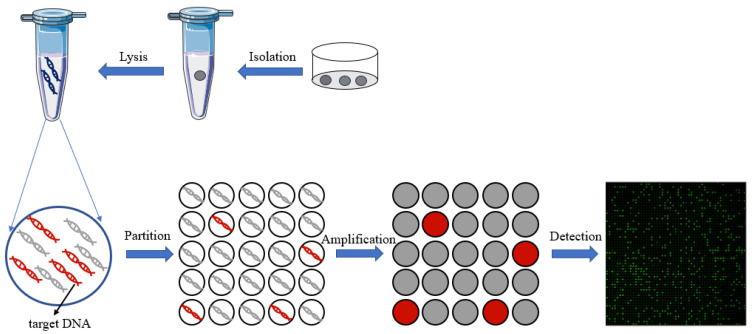
“The principle of digital PCR for single-cell analysis” Fang, W., Liu, X., Maiga, M., Cao, W., Mu, Y., Yan, Q., & Zhu, Q. (2024). Digital PCR for single-cell analysis. Biosensors, 14(2), 64.
- Nested Single-Cell PCR: Here, two primers (external and internal ) are used for the PCR reaction using the cell lysate.
- Reverse Transcriptase Single-cell PCR: For cells of interest, with RNA as the template genome, RT-PCR is utilized.
Examples of Single-cell PCR Kits
Single Cell-to-CT™ qRT-PCR Kit (ThermoFisher Scientific)
- The kit (AmbionTM Single Cell-to-CT) contains a validated workflow for the analysis of gene expression for samples with 1-10 cells.
- It is a real-time RT-PCR kit for the analysis of single cells.
- Provides maximum sensitivity for single-cell analysis, along with high reproducibility and performance.
- A single kit can be used for 50 reactions.
Single Cell RT-PCR Assay Kit (Singnosis)
- To study the gene expression of individual cells, such as stem cells, neurons, and developmental tissues, using laser capture microsections
- It is a sensitive single-cell RT-PCR using cell lysates as samples. Preparation of RNA is not required.
- Contains the necessary buffer, including the lysis buffer.
- Real-time analysis can be done.
Cell Lysis RT-qPCR Kits (BIO-RAD)
- Used to perform RT-qPCR of cultured cells without the RNA purification step.
- Both SYBR Green and Probes are available for real-time analysis.
- Cell yields from 10 to 100,000 cells can be prepared within 20 minutes.
- Genomic DNA is removed, ensuring that the RNA integrity is preserved.
Single-cell DNA-seq (TAKARA)
- Allows for the whole-genome amplification (WGA) using PicoPLEX technology
- Highly reproducible and uniform.
- Provides three kits: Integrated WGA and library preparations, standalone WGA preparation, and direct-to-library workflow utilizing next-generation sequencing.
Applications of Single-cell PCR
Single-cell PCR can be utilized for both diagnostic and non-diagnostic purposes. They are:
Diagnostic purposes
- Preimplantation genetic diagnosis: Utilized for the determination of the sex of single cells obtained from human preimplantation embryos for X-linked genetic disorders. It is used for the determination of single-gene disorders, including cystic fibrosis, hemoglobinopathies, and Duchenne’s muscular dystrophy.
Non-diagnostic purposes
- Immunological and oncology study: Since this method can be used for studying the genetic disparities between cells, it has found wide applications in immunology and oncology fields. It is especially used to examine the changes in the B- or T-cell receptors (BCR and TCR, respectively) or for determining the clonal origin of tumors.
Utilization of next-generation sequencing (NGS) of single cells, or single-cell proteomics, has made this method a suitable candidate for diagnostic and experimental methods in the future.
Advantages of Single-cell PCR
Some of the advantages of this method are:
- A study of gene expression can be accurately done.
- Allows for mutation and tumor studies.
- Whole-genome analysis of a single cell can be done.
- Very useful in studying genetic disorders.
Limitations of Single-cell PCR
The limitations of the method include:
- Chances of contamination and damage to cells during isolation
- Specific technologies should be used depending on the nature of the cell to be isolated, making it an expensive process.
- Early degradation of the DNA/RNA may occur due to the lysis buffer.
- Chances of loss of cell viability.
Conclusion
Single-cell PCR is a type of PCR method for the study of individual cells of interest. It has found wide applications in diagnostic and non-diagnostic purposes such as genetic disorder studies, parental studies, and forensic studies. Various methods are available for the isolation of cells, including Limited Serial Dilution, Micromanipulator, Microfluidics, and Flow Cytometry. With the use of Next-generation sequencing, this method can be very useful in the future for diagnostic as well as research purposes for Whole Genome Analysis.
References
- Fang, W., Liu, X., Maiga, M., Cao, W., Mu, Y., Yan, Q., & Zhu, Q. (2024). Digital PCR for single-cell analysis. Biosensors, 14(2), 64.
- Kai, A. K. L., Cheung, Y. K., Yeung, P. K. K., & Wong, J. T. Y. (2006). Development of single-cell PCR methods for the Raphidophyceae. Harmful Algae, 5(6), 649-657.
- Gross, A., Schoendube, J., Zimmermann, S., Steeb, M., Zengerle, R., & Koltay, P. (2015). Technologies for single-cell isolation. International journal of molecular sciences, 16(8), 16897-16919.
- Hahn*, S., Zhong, X. Y., Troeger, C., Burgemeister, R., Gloning, K., & Holzgreve, W. (2000). Current applications of single-cell PCR. Cellular and Molecular Life Sciences CMLS, 57(1), 96-105.
- Stumpf, F., Schoendube, J., Gross, A., Rath, C., Niekrawietz, S., Koltay, P., & Roth, G. (2015). Single-cell PCR of genomic DNA is enabled by automated single-cell printing for cell isolation. Biosensors and Bioelectronics, 69, 301-306.
- Wang, X., & Stollar, B. D. (2000). Human immunoglobulin variable region gene analysis by single cell RT-PCR. Journal of immunological methods, 244(1-2), 217-225.
- Findlay, I. (1998). Single Cell PCR: Theory, Practice, and Clinical Applications. In Clinical Applications of PCR (pp. 233-263). Totowa, NJ: Humana Press.
- https://www.thermofisher.com/order/catalog/product/4458237
- https://www.signosisinc.com/product-page/single-cell-real-time-reverse-transcriptase-pcr-assay-kit
- https://www.takarabio.com/products/next-generation-sequencing/dna-seq/single-cell-dna-seq
- https://www.bio-rad.com/en-in/product/cell-lysis-rt-qpcr-kits
- https://www.geeksforgeeks.org/maths/poisson-distribution/
