Protein folding is a precise, cooperative process in which a linear polypeptide chain navigates a complex conformational space to attain its biologically active three-dimensional conformation, known as the native state. This transition is primarily governed by the thermodynamic drive to reach a global energy minimum, largely facilitated by the hydrophobic effect.
Protein Misfolding and Aggregation
As the chain collapses, non-polar side chains are sequestered into a central core to avoid the aqueous cellular environment, while a precise network of hydrogen bonds and electrostatic interactions stabilizes the final structure. When successful, this intricate geometry ensures that functional sites, such as enzymatic pockets or binding domains, are perfectly positioned to execute specific biochemical tasks.
In contrast, misfolding occurs when a protein becomes kinetically trapped in a non-native configuration, often due to genetic mutations, environmental stressors, or molecular crowding. During this failure, the hydrophobic residues that should be buried within the core remain exposed to the solvent. These sticky patches drive abnormal intermolecular associations, leading to the formation of insoluble, highly ordered aggregates or amyloid fibrils.
While folding results in a functional molecular machine, misfolding not only leads to a loss of essential biological activity but also creates proteotoxic species that can disrupt cellular homeostasis. This delicate balance between a functional fold and a pathological aggregate is a defining threshold of molecular health.
Mechanisms of Misfolding: Genetic Mutations and Environmental Stress
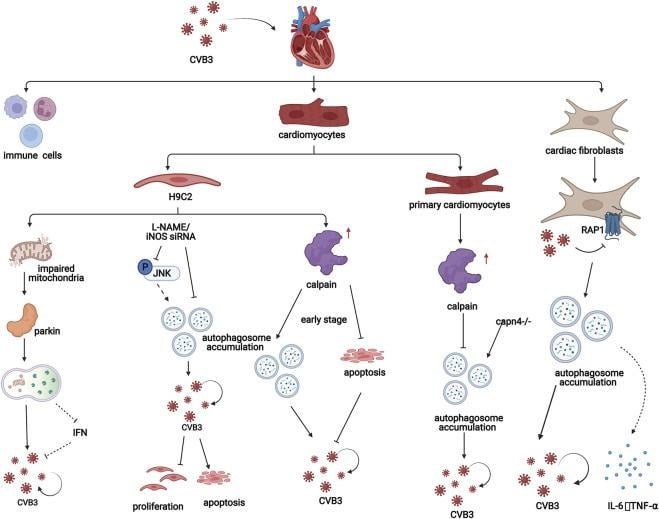
https://www.sciencedirect.com/science/chapter/bookseries/abs/pii/S1876162325000884
Two distinct destabilizing mechanisms typically catalyze the shift from a biologically active protein to a harmful aggregate: genetic mutations and environmental stress. Genetic mutations disrupt the protein’s primary sequence, often replacing critical amino acids with residues that possess incompatible chemical properties. This alteration destabilizes the native state by interfering with the precise network of intramolecular bonds such as salt bridges and van der Waals forces required for correct architecture. Consequently, the protein fails to bury its hydrophobic core, leaving sticky non-polar patches exposed to the aqueous cytosol, which initiates the formation of insoluble oligomers.
The Aggregation Pathway: Nucleation, Oligomers, and Fibrils
Nucleation
Nucleation serves as the rate-limiting initiation of protein aggregation, driven by a thermodynamic transition toward a lower-energy, ordered state. This phenomenon occurs when supersaturated or misfolded monomers overcome a formidable activation energy barrier through stochastic molecular collisions.
The process requires a substantial reduction in configurational entropy to organize disordered proteins into a stable critical nucleus. Once this assembly reaches a threshold size, further monomer addition becomes energetically favourable, terminating the lag phase and triggering rapid, spontaneous fibril propagation.
Oligomers
Oligomers represent metastable, non-fibrillar intermediates that arise during the intermediate stages of protein aggregation. These species are characterized by high structural heterogeneity and are frequently identified as the primary agents of proteotoxicity in neurodegenerative pathologies.
Unlike mature fibrils, oligomers often expose expansive hydrophobic surfaces, facilitating aberrant interactions with lipid bilayers and intracellular components. Their formation reflects a transition toward increased molecular order, yet they lack the rigid, long-range periodicity found in terminal amyloid structures, making them transient but highly reactive. Pathologically, oligomers induce synaptic impairment by disrupting long-term potentiation and stimulate oxidative stress via mitochondrial interference.
Although they lack the rigid periodicity of mature fibrils, their high reactivity and ability to trigger apoptotic cascades make them primary drivers of cellular dysfunction in neurodegenerative diseases.
Fibrils
Amyloid fibrils represent the terminal, thermodynamically stable end-state of the protein aggregation pathway, characterized by a highly ordered cross β sheet structure. In this structural motif, individual beta-strands align perpendicular to the longitudinal fiber axis, stabilized by an extensive network of inter-molecular hydrogen bonds.
These filamentous polymers grow via the templated recruitment of monomers onto their ends, a process known as elongation. Fibrils exhibit remarkable mechanical rigidity and resistance to proteolytic degradation, effectively sequestering misfolded proteins into insoluble macroscopic deposits.
Amyloid Structure: The Cross-Beta Sheet Architecture
The cross β sheet architecture represents the quintessential structural motif of amyloid fibrils, providing a level of thermodynamic stability that renders these aggregates nearly irreversible under physiological conditions. This quaternary structure is defined by an array of β strands oriented perpendicular to the longitudinal axis of the fibril, while the network of stabilizing inter-molecular hydrogen bonds extends parallel to the fiber growth direction.
A critical feature of this arrangement is the steric zipper interface, where the side chains of opposing β sheets interdigitate with high complementarity, effectively excluding water molecules from the fibrillar core. This dry, hydrophobic environment, combined with the dense hydrogen-bonding lattice, confers exceptional mechanical stiffness and significant resistance to both chemical denaturants and proteolytic enzymes.
Beyond its physical robustness, the cross β framework facilitates the process of templated recruitment, allowing mature fibrils to act as scaffolds for the incorporation of incoming misfolded monomers. This repetitive, crystalline-like symmetry is remarkably conserved across diverse, non-homologous protein sequences, suggesting that the amyloid state is a universal energy minimum for polypeptide chains.
Consequently, the formation of this architecture signifies a transition from functional, soluble protein states to insoluble, filamentous polymers, a process fundamentally linked to the progression of various protein misfolding disorders.
Cellular Toxicity: Loss of Function vs. Toxic Gain of Function
Protein aggregation induces cellular toxicity through two distinct yet synergistic pathways: the depletion of functional protein species and the emergence of deleterious molecular interactions. Loss of function involves the deprivation of a protein’s native biological role due to sequestration, while toxic gain of function describes the acquisition of harmful new properties by misfolded assemblies. Together, these mechanisms destabilize cellular homeostasis.
| Feature | Loss of Function (LOF) | Toxic Gain of Function (GOF) |
| Primary Driver | Depletion of the soluble, native protein pool. | Formation of reactive, misfolded intermediates. |
| Mechanism | Failure of essential signaling, catalysis, or transport. | Membrane disruption, pore formation, and ROS production. |
| Species Involved | Absent/Sequestrated monomers. | Metastable oligomers and pre-fibrillar species. |
| Cellular Impact | Diminished homeostatic capacity and metabolic failure. | Active induction of apoptotic and inflammatory cascades. |
| Example | Reduced DNA repair in p53 aggregation. | Calcium dysregulation via Amyloid- β oligomers. |
The Role of Molecular Chaperones and the Unfolded Protein Response (UPR)
Molecular chaperones and the Unfolded Protein Response (UPR) represent a synergistic proteostasis framework essential for preserving proteome stability and preempting the formation of deleterious protein aggregates.
Molecular chaperones, including the Hsp70 and Hsp90 stress-protein families, operate by binding to exposed hydrophobic residues on nascent or unfolded polypeptides. This interaction provides a protected spatial environment that facilitates native folding and prevents non-specific intermolecular associations.
When the accumulation of misfolded proteins overwhelms the biosynthetic capacity of the endoplasmic reticulum (ER), a condition termed ER stress ensues, activating the Unfolded Protein Response. This signal transduction network is orchestrated by three transmembrane sensors: PERK, IRE1, and ATF6.
Initially, the UPR promotes a cytoprotective state by downregulating global protein synthesis thereby easing the ER’s folding burden while concurrently enhancing the expression of chaperones and ER-associated degradation (ERAD) factors. However, under conditions of chronic or unresolved stress, the UPR undergoes a functional switch toward pro-apoptotic signaling.
This transition activates terminal caspase cascades, facilitating the programmed elimination of damaged cells to prevent the systemic dissemination of proteotoxic specie
Prions and Transmissible Spongiform Encephalopathies (TSEs)
Prions are unconventional infectious pathogens consisting exclusively of misfolded proteinaceous material, lacking any genetic material. The pathogenesis of Transmissible Spongiform Encephalopathies (TSEs) centres on the post-translational conformational transition of the endogenous cellular prion protein PrP^C from a predominantly α helical state into the pathological, β sheet-enriched PrP^Sc isoform.
This transition facilitates a self-propagating seeding mechanism, wherein PrP^Sc serves as a structural template that forces native monomers into a protease-resistant, amyloid-like configuration.
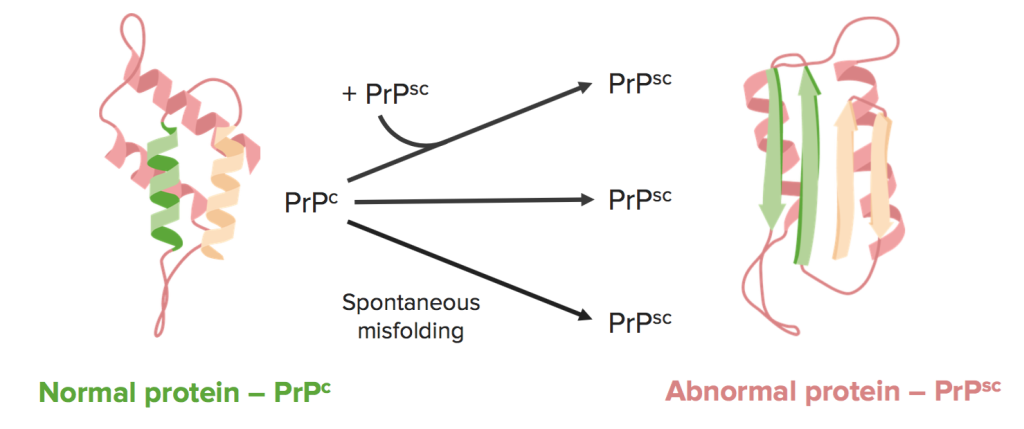
https://www.lecturio.com/concepts/transmissible-spongiform-encephalopathies/
This image illustrates the conversion of a normal cellular prion protein PrP^C, which is rich in alpha-helices, into the pathogenic, beta-sheet-rich abnormal isoform PrP^Sc. This transition can occur through spontaneous misfolding or be induced when an existing abnormal protein interacts with a normal one, triggering a chain reaction of protein misfolding.
Diseases such as Creutzfeldt-Jakob Disease exhibit prolonged latency periods followed by precipitous neurological decline. Histologically, PrP^Sc deposition triggers extensive neuronal vacuolation and reactive astrogliosis, imparting a spongiform morphology to the cerebral cortex. A defining characteristic of prions is their inherent infectivity and remarkable resilience to conventional decontamination protocols, such as autoclaving or enzymatic digestion.
By bypassing typical species barriers and hijacking host proteostasis, prions represent an exceptionally stable and lethal protein fold, distinguishing TSEs from other non-transmissible neurodegenerative proteinopathies.
Neurodegenerative Diseases: Alzheimer’s, Parkinson’s, and Huntington’s
Neurodegenerative pathologies, including Alzheimer’s, Parkinson’s, and Huntington’s diseases, constitute a diverse class of disorders unified by the progressive attrition of specific neuronal subsets due to proteostasis failure.
In Alzheimer’s Disease, the hallmark features include the extracellular accumulation of amyloid-β into senile plaques and the intracellular polymerization of hyperphosphorylated tau into neurofibrillary tangles, which collectively disrupt synaptic signaling and drive cerebral atrophy. In contrast, Parkinson’s Disease is characterized by the misfolding of α-synuclein into insoluble Lewy bodies. These aggregates exert localized toxicity within dopaminergic neurons of the substantia nigra, manifesting as progressive motor dysfunction and cognitive impairment.
Huntington’s Disease is distinguished by its strictly genetic origin, stemming from an autosomal dominant CAG trinucleotide expansion in the HTT gene. This expansion encodes a polyglutamine (polyQ) tract that destabilizes the huntingtin protein, fostering its transition into toxic nuclear and cytoplasmic inclusions.
Despite their varied molecular identities β, α-synuclein, and polyQ-hunting in these disorders converge on shared cytotoxic mechanisms: the induction of oxidative stress, mitochondrial depolarization, and the functional exhaustion of the ubiquitin-proteasome system. Ultimately, the conversion of these proteins into thermodynamically stable, amyloid-like conformations marks a terminal transition that precipitates irreversible neuronal degeneration.
Techniques for Detecting Aggregates: Thioflavin T and Cryo-EM
Analytical detection of protein aggregates relies on complementary biophysical techniques to quantify kinetics and resolve atomic structures. Thioflavin T (ThT) is a benzothiazole dye used for real-time kinetic monitoring; it exhibits enhanced fluorescence upon binding to the periodic β-sheet grooves of amyloid fibrils, serving as a gold-standard extrinsic probe for aggregation assays.
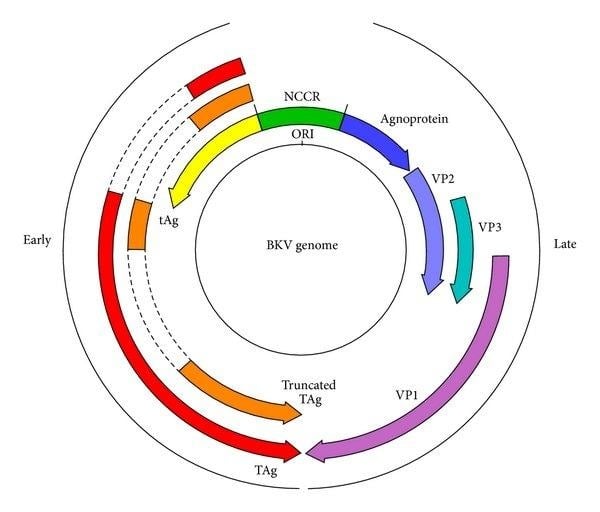
https://www.sciencedirect.com/science/article/abs/pii/S0021979720303738
Thioflavin T is used as a diagnostic tool to detect these misfolded states by binding to the cross-beta sheet structures of amyloid fibrils.
Conversely, Cryo-Electron Microscopy (Cryo-EM) provides high-resolution structural determination by imaging vitrified samples in a near-native state. This technique allows for the atomic-level visualization of the cross β backbone and steric zipper side-chain orientations, enabling the differentiation of distinct clinical polymorphs or strains that fluorescence assays cannot distinguish.
Therapeutic Strategies: Inhibiting Aggregation and Disaggregating Fibrils
Therapeutic intervention in proteinopathies is bifurcated into strategies that preemptively inhibit the formation of new aggregates and those that actively disaggregate existing fibrillar deposits.
Comparative Analysis of Therapeutic Strategies
| Feature | Inhibition of Aggregation | Disaggregation of Fibrils |
| Primary Objective | To prevent the transition from monomeric states to metastable oligomers or fibrils. | To solubilize and clear pre-existing, thermodynamically stable amyloid deposits. |
| Mechanism of Action | Kinetic Stabilization: Binding to the native fold to increase the activation energy barrier. | Active Extraction: Using molecular motors (disaggregases) to pull monomers from the cross-beta lattice. |
| Molecular Targets | Monomers and primary/secondary nucleation sites. | Mature fibrils and high-order insoluble plaques. |
| Examples | Small molecules (e.g., Tafamidis), kinetic stabilizers, and peptides. | Engineered chaperones (e.g., Hsp104, Hsp70 complexes) and monoclonal antibodies. |
| Key Limitation | Generally ineffective once significant plaque burden is established (requires early intervention). | Risk of releasing reactive, toxic oligomeric seeds during the fragmentation process. |
| Therapeutic Logic | Preserves native function by maintaining the soluble protein pool. | Reverses pathology by restoring cellular space and reducing toxic load. |
Conclusion
The transition from functional protein folding to pathological misfolding represents a critical threshold in cellular health. The formation of amyloid fibrils via the nucleation-dependent pathway results in a thermodynamically stable cross β architecture, characterized by interdigitated steric zippers that resist degradation. This structural shift drives cellular toxicity through a synergistic combination of functional depletion (LOF) and the emergence of reactive, pore-forming oligomers (GOF).
While the proteostasis network, orchestrated by molecular chaperones and the Unfolded Protein Response (UPR), serves as a vital defence mechanism, its failure precipitates terminal neurodegenerative disorders like Alzheimer’s, Parkinson’s, and infectious prion diseases. Modern biophysical tools like Cryo-EM and Thioflavin T assays are essential for resolving these structures, providing the atomic insights necessary to develop dual-action therapies.
Ultimately, balancing the inhibition of new aggregates with the safe disaggregation of existing fibrils remains the frontier in reversing the systemic collapse of the cellular proteome.
References
- Dobson, C. M., Šali, A., & Karplus, M. (1998). Protein folding and misfolding: A perspective from theory. Angewandte Chemie International Edition, 37(7), 868–893. https://doi.org/10.1002/(SICI)1521-3773(19980420)37:7<868::AID-ANIE868>3.0.CO;2-
- Moreno-Gonzalez I, Soto C. Misfolded protein aggregates: mechanisms, structures and potential for disease transmission. Semin Cell Dev Biol. 2011 Jul;22(5):482-7. doi: 10.1016/j.semcdb.2011.04.002. Epub 2011 May 5. PMID: 21571086; PMCID: PMC3175247.
- Almeida, Z.; Brito, R. Structure and Aggregation Mechanisms in Amyloids. Encyclopedia. Available online: https://encyclopedia.pub/entry/17529 (accessed on 06 February 2026).
- Nelson R, Sawaya MR, Balbirnie M, Madsen AØ, Riekel C, Grothe R, Eisenberg D. Structure of the cross-beta spine of amyloid-like fibrils. Nature. 2005 Jun 9;435(7043):773-8. doi: 10.1038/nature03680. PMID: 15944695; PMCID: PMC1479801.
- Winklhofer KF, Tatzelt J, Haass C. The two faces of protein misfolding: gain- and loss-of-function in neurodegenerative diseases. EMBO J. 2008 Jan 23;27(2):336-49. doi: 10.1038/sj.emboj.7601930. PMID: 18216876; PMCID: PMC2234348.
- Hetz, C., Zhang, K., & Kaufman, R. J. (2020). Mechanisms, regulation and functions of the unfolded protein response. Nature Reviews Molecular Cell Biology, 21(8), 421–438. https://doi.org/10.1038/s41580-020-0250-z
- https://www.youtube.com/watch?v=ESC3CSApNnk
- Hartl, F. U., Bracher, A., & Hayer-Hartl, M. (2011). Molecular chaperones in protein folding and proteostasis. Nature, 475(7356), 324–332. https://doi.org/10.1038/nature10317
- Reference List: Center for Food Security and Public Health. (2016). Transmissible spongiform encephalopathies. Iowa State University. https://www.cfsph.iastate.edu/Factsheets/pdfs/transmissible_spongiform_encephalopathy.pdf
- Ciurea AV, Mohan AG, Covache-Busuioc RA, Costin HP, Glavan LA, Corlatescu AD, Saceleanu VM. Unraveling Molecular and Genetic Insights into Neurodegenerative
- Diseases: Advances in Understanding Alzheimer’s, Parkinson’s, and Huntington’s Diseases and Amyotrophic Lateral Sclerosis. Int J Mol Sci. 2023 Jun 28;24(13):10809. doi: 10.3390/ijms241310809. PMID: 37445986; PMCID: PMC10341997.
- Min, JH., Sarlus, H., Oasa, S. et al. Thioflavin-T: application as a neuronal body and nucleolar stain and the blue light photo enhancement effect. Sci Rep 14, 24846 (2024). https://doi.org/10.1038/s41598-024-74359-8
- Cao, P., Linden, A. H., & Reif, B. (2020). Cryo-electron microscopy provides mechanistic insights into solution-dependent polymorphism and cross-aggregation phenomena of the human and rat islet amyloid polypeptides. Biochemistry, 59(45), 4307–4310. https://doi.org/10.1021/acs.biochem.0c00713
- Giorgetti S, Greco C, Tortora P, Aprile FA. Targeting Amyloid Aggregation: An Overview of Strategies and Mechanisms. Int J Mol Sci. 2018 Sep 9;19(9):2677. doi: 10.3390/ijms19092677. PMID: 30205618; PMCID: PMC6164555.
