Digital PCR was invented in 1999 by Vogelstein and Kinzler, a method that involves diluting DNA into microtiter plates and utilizing fluorescent probes for detection. Such a method allows for the detection and quantification of the DNA segments, which estimates the number of molecules by statistical means.
Unlike traditional PCR, which measures the amount of accumulated product at the end of each cycle, dPCR measures the fraction of negative microreactions to determine absolute copies of the segment of interest. Compared to the traditional PCR method, dPCR is a more quantitative method as it counts individual molecules for absolute quantification.
Since dPCR works on the use of minimally diluted samples, it has allowed for the detection of rare mutations in a bulk of wild-type sequences. Such an application helped in the tumour heterogeneity analysis in oncology and liquid biopsy applications. It is also used in prenatal diagnosis for inherited mutations.
Objectives of Digital PCR (dPCR)
The main objective of dPCR is to enable a precise detection and highly sensitive quantification of nucleic acid from minimally diluted samples of DNA segments of interest. This is done by partitioning a single PCR reaction into multiple subreactions and allowing these reactions to multiply in their optimal conditions. If the subreaction amplifies, it indicates the presence of the target nucleic acid in that partition. Similarly, if a subreaction does not amplify, that is, it is a negative subreaction, this indicates the absence of such nucleic acid. Digital PCR allows for the precise quantification without its dependence on an external reference curve (a standard curve).
Requirement for Digital PCR (dPCR)
The materials involved in dPCR include:
- Conventional PCR materials such as primers, dNTPs, polymerase, buffers, ions, etc.
- A reporter molecule is used, which can either be a dye or a probe. Fluorescent DNA-binding dye is usually used as it is sequence-specific; hydrolysis probes can also be utilised.
- Thermocycler
- Partition chambers: This is required for the partition of the sample mixture into microchambers. The chamber used can be a microfluidic chip, microarrays, spinning microfluidic discs, or droplet techniques based on oil-water emulsions.
- Fluorescence analyser: This is required for the quantification of the results since a single subreaction may contain more than one target molecule. The results from this are then analysed by using the Poisson model.
Principle of Digital PCR (dPCR)
Digital PCR is a precise and sensitive method that allows for the detection of rare events such as single-nucleotide mutations in a population of wild-type sequences. Here, the competition between the partitions with the targets is reduced, which subjugates the inherent difficulties in amplifying rareequence, thereby allowing for sensitive and precise absolute quantification of nucleic acids. Digital PCR has been widely used in oncology in the detection of rare mutations, the detection and quantification of viruses and pathogens, single-cell analysis, etc.
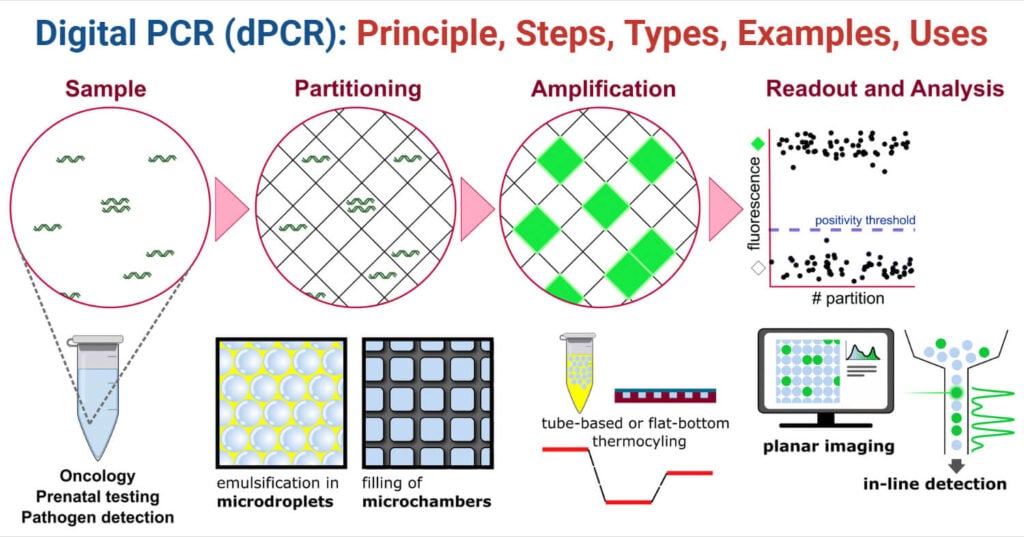
The detection of nucleic acids through this experimental approach is done by; first the DNA to be analyzed is first diluted and evenly partitioned into microtitre plates, with one template per two wells. The amplification is then performed by providing the optimal conditions to multiply and provide copies of the segment of interest. The copies, i.e, the amplicons, are then hybridized with a reporter molecule, here being a fluorescent DNA-binding probe. Such a probe allows detection of sequence-specific products using different fluorophores. Finally, the fluorescent signals are analysed using statistical approaches.
An important step in dPCR is the division of the sample mixture into microwells before its amplification. Each of these partitions may contain zero, one, or more template molecules. The efficiency of the amplification varies from sample to sample, with minor inhibitors or reduced target accessibility having less impact on such variations.
Upon amplification, these amplicons are analysed with the help of fluorescent probes, which allows for detecting the presence of the target DNA. Samples that show a positive reaction (1), i.e, the samples that fluoresce, contain the target molecule, which would not be the case in a negative reaction (0).
The ratio of positives to negatives in each sample is the basis of quantification. The partitioning of the samples, coupled with the quantification done by the Poisson statistic data analysis, is more reliable than traditional PCR methods, as it helps in yielding more precise and reliable results.
Steps of Digital PCR (dPCR)
Based on the aforementioned principle, the steps in performing dPCR can be summarized into five main steps. This includes:
- Sample: The samples are used to detect DNA, RNA, or cDNA from different specimens.
- Partitioning: Partition of the sample mixture into various wells or microtitre plates. Here, the sample is separated into small containers, ranging from thousands to millions in number. The target DNA is randomly distributed in each of these wells.
- Amplification: Upon partitioning, the sample present in each of the wells is provided with optimal PCR conditions and made to amplify using a thermal cycler.
- Hybridization: Once done with the amplification step, the samples are then hybridized with a reporter molecule (Fluorescent probe), which is then used for performing an endpoint analysis.
- Analysis and Quantification: With the result obtained, statistical analysis of the positive and negative subreactions is done using the Poisson statistical model.
1. Sample
The sample contains the specimen with the target sequence, along with the usual components of a traditional PCR such as primers, dNTPs, polymerase, buffers, ions, enzyme, etc, and reporter molecule, which can be a hydrolysis probe or Fluorescent DNA binding dye. The specimen can be taken from various sources such as tissue samples, body fluids, etc.
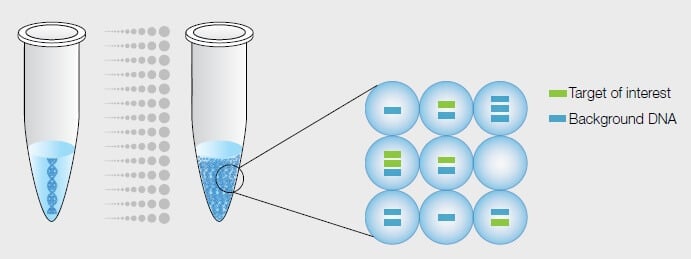
2. Partitioning
Once the sample is thoroughly mixed, it is equally separated into microchambers to undergo the amplification process. Typically, the sample amount comes out to be 20μL, which is then separated into wells, the most common being 96-well microchambers. The partitioning is done in a way that there is, on average, one template molecule per two wells.
Partitioning is of two major types: Water-in-oil droplet emulsification and microchambers. In the former, which is commonly known as droplet digital PCR (ddPCR), the sample mixture is scattered into droplets of size ranging from pL to nL. These monodroplets can be generated at high speed using a microfluidic chip by using forces that can actively break the aqueous/oil interface. During this temperature variation of the PCR, there are chances of coalescence of water-in-oil droplets. Thus, their stabilisation can be done with an appropriate surfactant. ddPCR provides greater scalability and cost-effectiveness, unlike microchamber dPCR.
Microchamber-based dPCR was the first to be invented. Here, an array of thousands of microscopic wells that are embedded in solip chip is used. Unlike ddPCR, which requires precise emulsification of the sample and stability of the monodisperse droplets, the microchamber dPCR provides a higher reproducibility rate and easy automation. However, it is limited to a fixed number of partitions and has a higher cost.
With advances in technology, there are various instruments available in the market that help in fast and precise partitioning into hundreds, thousands, or even millions of subreactions with various degrees of resolution.
3. Amplificaton
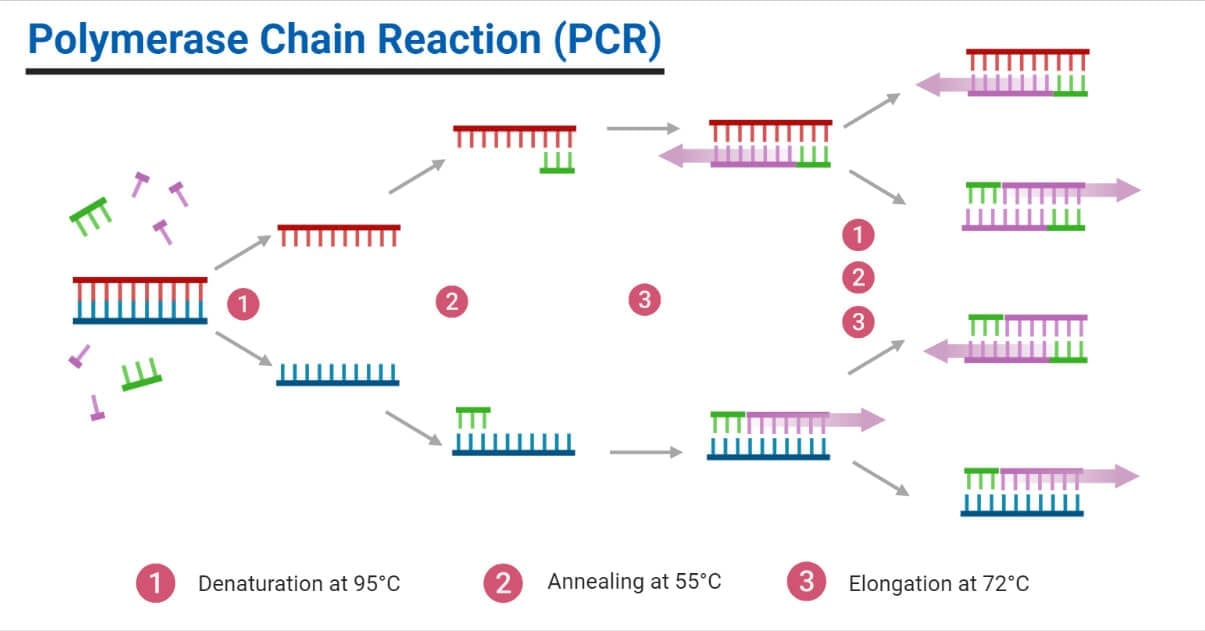
Once the separation/partitioning of the sample has been done, they are provided with optimal conditions to perform the standard PCR reaction. Here, each of the wells undergoes a separate PCR reaction. The wells with the target amplify, providing a positive subreaction. The standard PCR reaction includes three steps:
- Denaturation: Here, the DNA is heated up to 95°, which aids in its disruption of hydrogen bonds between the base pairs.
- Annealing: Right after denaturation, the DNA is cooled to a temp range of 55°C to 72°C. Such a cooling helps in the primers to bind to their complementary sequences. This stage allows for the subsequent synthesis and elongation of the DNA.
- Extension: DNA elongation occurs in this phase. Here, the temperature is raised to 75°C to 80°C, which optimizes the DNA polymerase activity to promote strand elongation.
The efficiency of the amplification process varies with the sample, which can be affected by minor inhibitors. Delayed amplification start due to issues with the target accessibility does not have a bigger impact on the quantitative results than when using other techniques. However, general inhibition does not affect the analysis of the results as long as the positive subreactions can be differentiated from the negative results.
4. Hybridization
After the amplification process, the samples in the wells are hybridized with a fluorescent probe, which helps in differentiating between the positive and negative subreactions. Since the products of the PCR after amplification are homogenous in sequence as the template is a single molecule, fluorescence-based technologies are the best methods for the analysis of the products. Currently, molecular beacons are widely used for analysis in dPCR reactions.
Additionally hydrolysis probe can also be used for the detection and differentiation of the procedures. There are cases when the positive ureactions do not fluoresce; this is due to delayed amplification from primer accessibility issues or a point mutation at the top probe annealing site.
5. Analysis and Quantification
Even though the fluorescent probes help in the detection and differentiation of the positive and negative subreaction, due to random assortment, it is ensured that each positive reaction contains only a single molecule. This happens when the amount of sample taken is above a certain percentage, and there are fewer wells. Thus, for proper quantification of positive reaction, a correction factor to determine the absolute template quantity is applied using the Poisson model.
To determine the total target molecule count, a positive and a negative subreaction count are used. Here, the ratio of positives (p) to the total number is used to determine the average number of targets per subreaction (λ) using the Poisson formula below, and then multiplied by the number of subreactions per μl. Sample digitization coupled with the Poisson statistical model helps in providing precise data. Negative reaction holds the most importance when determining the copies per well. They help in establishing the ratio of positive and negative reactions. The formula used for Poisson modelling is :
Copies per partition, λ = -ln(1-p)
Where p is the number of positive reactions
Types of Digital PCR (dPCR)
Digital PCR can be divided into two types based on the partitioning technique used. This includes:
- Droplet Digital PCR: In ddPCR, the sample is dispersed within an immiscible oil phase as tiny droplets
- Microchamber Digital PCR: Here, the sample is partitioned into small chambers or wells, which are embedded in a solid chip.
Despite the type of partitioning followed, the principle the method of the PCR remain unchanged.
Examples of Digital PCR (dPCR)
1. QX700™ E System (Bio-Rad)
- Droplet Digital PCR is a water-emulsion droplet-based technology.
- The sample is fractionated up to 17,000 droplets and amplified by providing optimal conditions.
- The individual target is amplified, and the positive results are taken to obtain precise, absolute target quantification.
- Up to 5 μL of the sample is partitioned into 192 wells, and the results are obtained at the 5-hour mark.
- Bio-Rad’s QX700™ E System utilises water-emulsion technology coupled with microfluidics.
- Other ddPCR instruments from Bio-Rad are QX Continuum System, QX700 S System, QX700 HT System, QX600 System, QX200 System, QX ONE System, and QXDx AutoDG System.
2. QuantStudio Absolute Q Digital PCR System (ThermoFisher)
- This is a plate-based dPCR system incorporated with Microfluidic array plate (MAP) technology.
- This technology allows the necessary steps, such as compartmentalizing, thermal cycling, and data acquisition, to be performed in a single instrument.
- MAP has 16 distinct dPCR reaction units, and each of these reaction units is made up of 20,480 fixed array microchambers.
- These microchambers are, in turn, connected to a distribution flow system that feeds them with the PCR reagents.
- Upon compartmentalization of these reagents, amplificaionproceeds and the microchambers with successful amplification are counted.
- This instrument analyzes over 95% of the input sample, unlike the other ones in the market, which analyze only 25-60%.

3. Nio®PCR (Stilla)
- Nio® Digital PCR is an all-in-one instrument that is user-friendly and available in 3 different configurations.
- It sis easy to use, fast, and highly automated, and features one of the best hardware and software.
- This technology is also referred to as Crystal Digital PCR. It partitions samples into a large array of thousands of individual droplet crystals.
- Each of these crystals has its own reaction compartment, and they undergo amplification within these compartments.
- Upon amplification, the targets are tagged with fluorophores and read up to seven different fluorescence light channels. Such a method allows for the development of assays that additionally allow for individually quantifying up to 21 targets simultaneously.
4. QIAcuityDx Digital PCR System (Qiagen)
- This is a nano-plate-based technology used for in vitro diagnostic use, using automated multiplex quantification dPCR technology.
- The nanoplates technology allows for the nanoplate partitioning, thermocycling, and image analysis, which is entirely automated.
- Based on the type of plate, up to 96 samples can be analyzed per plate. For example, for diagnostic purposes, the QIAcuityDx Nanoplate 26K is used with 24 wells.
- The wells have a system which allows for continuous loading, making it minimal hand on technology.
- The QIAcuityDx Control Software manages the integrated system, from handling the robotic gripper for nanoplate handling, a partitioning module, a thermocycler, to the fluorescence imaging module.
- Up to 4 plates are processed per run, with 120 minutes required for the first plate and 80 minutes for the subsequent ones.
Applications of Digital PCR (dPCR)
Digital PCR has found itself useful in various sectors; some of the applications are as follows.
- Viral pathogen detection: Due to the low amount of sample requirement for dPCR, it makes a good technology for the detection of viruses, which is often a cumbersome process due to the low availability of the specimen.
- High-resolution gene expression analysis: Since the quantification of the results is done by the Poisson statistical model, the numerical results are independent of other samples. Because of this, when the results are used to reference genes, they produce a more statistically significant result.
- Copy number variation: dPCR allows for the detection and quantification of small percentages of copy number difference with a higher degree of precision.
- Single-cell analysis: Typically, when analyzing single cells, an amplification step is required before proceeding with downstream processing. dPCR can yield highly accurate results with fewer preamplification steps, as dPCR can occur with lower target levels.
- Rare mutation abundance detection: The use of partitioning and enrichment effects in dPCR enables high-resolution detection of somatic mutations from disease-containing samples.
It is also used in environmental pathogen detection, library quantification for next-generation sequencing, GMO and food testing, etc.
Advantages of Digital PCR (dPCR)
There are several advantages of dPCR over traditional PCR methods:
- Partitioning of the sample ensures a more precise and high-resolution result.
- Since the Poisson statistical model is utilised for determining he absolute copy number, there is no need to rely on references or any standard curve.
- The inhibitors that may be present in the sample do not affect the amplification process.
- Helps in detecting rare mutations, i.e, able to analyze complex mixtures
- Even the smallest percentage of change can be detected and is reported linearly to the number of copies.
- It is calibration-free, highly sensitive, and provides absolute quantification.
Limitations of Digital PCR (dPCR)
Some of the disadvantages of dPCR are:
- Partitioning of the sample requires high precision and time.
- dPCR requires a high cost due to the use of the reagents, fluorescent probes, etc.
- In the case of droplet digital PCR, precise emulsification and stability should be ensured, making it a consuming process.
- Microhamber dPCR is limited by the fixed number of partitions and typically has higher costs.
- Care should be taken when calculating using the Poisson model to reduce the chances of errors.
Conclusion
Digital PCR is a third-generation PCR type that allows for obtaining accurate results with low levels of sample. Partitioning of the samples into small microchambers adds to its success in providing high-resolution results with low levels of inaccuracies. Detection of rare mutations and their applications in oncology is due to its outstanding accuracy of quantification of nucleic acids. Despite its limitations in being expensive and time-consuming, various advances are being made to overcome the practical and technical obstacles in the use of dPCR.
References
- Trouchet, A., Gines, G., Benhaim, L., & Taly, V. (2025). Digital PCR: from early developments to its future application in clinics. Lab on a Chip.
- Vogelstein, B., & Kinzler, K. W. (1999). Digital pcr. Proceedings of the National Academy of Sciences, 96(16), 9236-9241.
- https://www.thermofisher.com/blog/behindthebench/what-is-digital-pcr/
- Khehra N, Padda IS, Zubair M. Polymerase Chain Reaction (PCR) [Updated 2025 Jul 7]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan-.
- https://www.genome.gov/genetics-glossary/Polymerase-Chain-Reaction-PCR
- https://www.bio-rad.com/en-us/life-science/learning-center/introduction-to-digital-pcr/what-is-droplet-digital-pcr
- Pohl, G., & Shih, I. M. (2004). Principle and applications of digital PCR. Expert review of molecular diagnostics, 4(1), 41-47.
- Bizouarn, F. (2014). Introduction to digital PCR. In Quantitative real-time PCR: Methods and protocols (pp. 27-41). New York, NY: Springer New York.
- https://www.bio-rad.com/en-in/life-science/droplet-digital-pcr
- https://www.thermofisher.com/in/en/home/life-science/pcr/digital-pcr/quantstudio-absolute-q-system.html
- https://www.stillatechnologies.com/multiplex-pcr/nio-dpcr/
- https://www.qiagen.com/us/products/instruments-and-automation/pcr-instruments/qiacuity-dx-mdx.
