DH5α is a genetically engineered Escherichia coli K-12 strain widely used in molecular biology to amplify exogenous plasmid DNA.
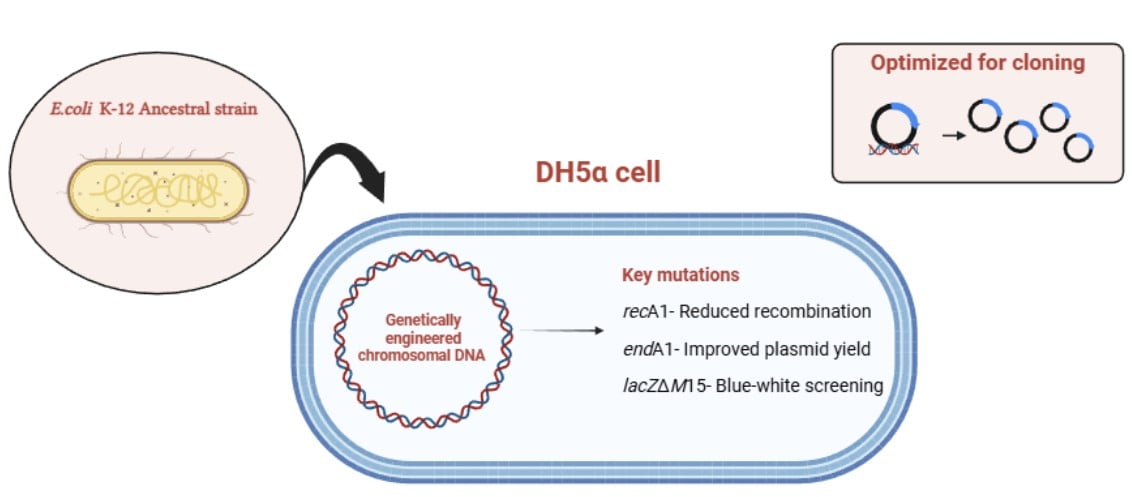
The strain was originally constructed by Douglas Hanahan from the parental strain DH1 and the foundational documentation of its use in plasmid transformation was published by Hanahan and colleagues in 1991. The complete genomic size of DH5 alpha is 4,630,707 bases with specific structural variations accounting for ~96 kb difference from its parent strain DH1.
Genotype of DH5 alpha
F- ϕ80dlacZΔM15, Δ(argF−lac)169, U169, deoR, recA1, endA1, hsdR17(rK-mK+), phoA, supE44, λ-, thi-1, gyrA96, relA1
Understanding the key mutations
DH5α features several classic mutations that make it an ideal host for recombinant DNA;
- recA1 – This mutation inactivates recA protein which results in the reduction of homologous recombination ensuring the stability of foreign plasmids without rearrangement or degradation.
- endA1– This mutation inactivates a non-specific endonuclease that leads to significantly higher quality DNA minipreps. In wild type, the enzyme often degrades plasmid DNA during isolation process.
- gyrA96 – This is the mutation in DNA gyrase which is an enzyme responsible for unwinding and relieving tension in the bacterial DNA during processess like replication, transcription and recombination. This mutation confers resistance to nalidixic acid which is a drug that blocks DNA gyrase. This resistance to nalidixic acid is useful as a selectable marker so only the bacteria with the mutation can grow.
- hsdR17(rK−,mK+) – This is the mutation of restriction-modification system of E. coli. With this mutation, the cells don’t digest the foreign DNA (restriction negative/ rK−) while it can still be able to methylate its own DNA protecting it from accidental digestion (modification positive/ mK+).
- Δ(argF−lac)169 – This is an 85,086 bp deletion (stretching from mmuP to cynS) which simplifies the genome for easier and stable cloning.
- ϕ80dlacZΔM15 – Phage DNA with modified lacZ, which involves a specific modification where the lacZ gene is 94 bases shorter at the N-terminus than in the parental strain, enabling α-complementation for blue-white screening to easily identify bacteria carrying a plasmid.
- phoQ – An amino acid change (Val306 to Ile) in this sensor protein increases the tolerance to multivalent ions (like Mn2+ or Ca2+) found in transformation buffers.
Table 1: Key mutations in DH5α and their roles in cloning
| S.N. | Key mutations | Gene affected /system | Function | Benefit in cloning |
| 1 | recA1 | recA(recombination) protein inactivated | Disables homologous recombination | Prevents plasmid rearrangent, esuring genetic stability. |
| 2 | endA1 | endA1 (endonuclease I) | Inactivates non-specific endonuclease | Produces high quality plasmid minipreps for sequencing and restriction analysis. |
| 3 | gyrA96 | DNA gyrase | Alters DNA supercoiling | Enhances plasmid stability and confers resistance to nalidixic acid, enabling selection of mutant cells. |
| 4 | hsdR17(rK−,mK+) | Restriction/Modification system | Prevents restriction digestion of foreign DNA while being able to methylate its own DNA | Prevents degradation of DNA during transformation. |
| 5 | Δ(argF−lac)169 | lac operon (lacZYA region) | An 85,086 bp deletion (stretching from mmuP to cynS) | Simplifies genome for easy cloning. |
| 6 | ϕ80dlacZΔM15 | lacZ (β-galactosidase α-fragment) | N-terminal deletion of lacZ enabling α-complementation and blue-white screening. | Blue-white screening to easily identify bacteria carrying a plasmid. |
| 7 | phoQ | phoQ sensor protein | An amino acid change (Val306 to Ile) in this sensor protein | Increases the tolerance to multivalent ions (like Mn2+ or Ca2+) found in transformation buffers. |
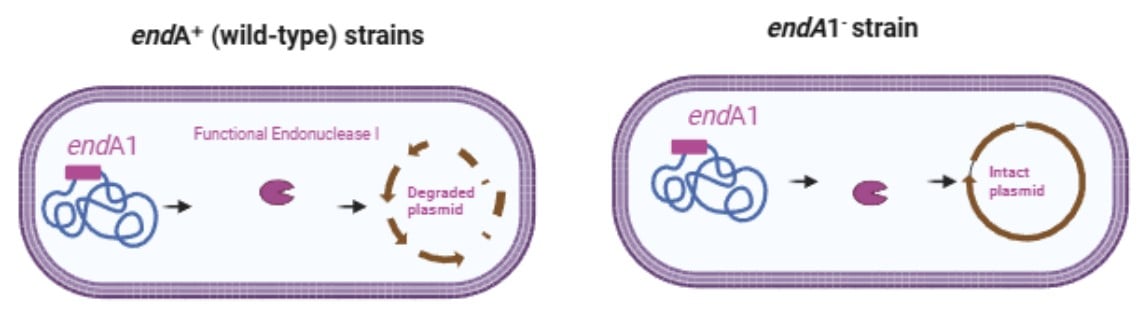
Role of the endA1Mutation: Ensuring High-Quality Plasmid Isolation
The endA gene in E.coli encodes for endonuclease I which is a non-sequence-specific, highly abundant enzyme responsible for the double-strand breaks in duplex DNA. This enzyme plays important role in the DNA turnover and genomic integrity under normal cellular conditions. However, endonuclease I degrades both chromosomal and plasmid DNA during cell lysis during plasmid isolation. In endA⁺ (wild-type) strains, this nuclease activity causes plasmid DNA to lose its supercoiled form and appear as degraded smears on agarose gel, reducing DNA quality.
The endA mutation present in cloning strains like DH5α, significantly reduces endonuclease activity which results in:
- Prevention of plasmid DNA degradation during isolation.
- Stability of supercoiled plasmid conformation.
- Improved purity and integrity of isolated DNA.
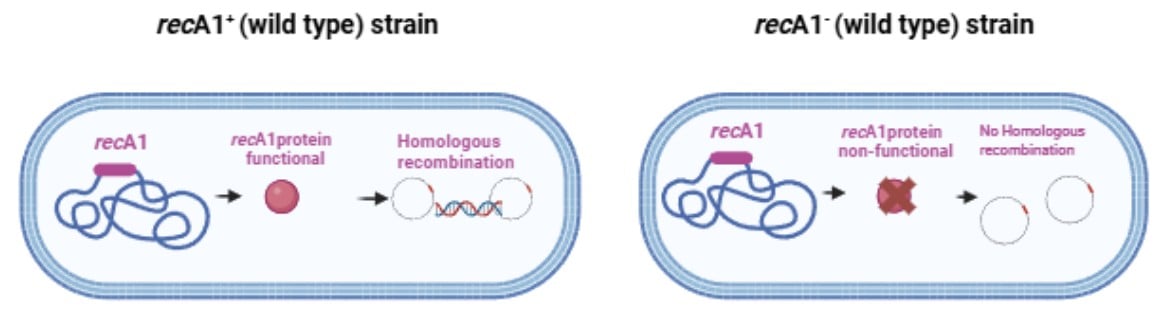
recA1 Deficiency: Preventing Homologous Recombination and DNA Instability
The recA1 mutation is a specific genetic alteration frequently incorporated in E. coli strain to enhance DNA stability by inactivating the cell’s primary homologous recombination pathway.
The recA allele posesses a point mutation that changes glycine 160 to Aspartic acid (G160D) which results in defective recA protein that can still bind to single-stranded DNA (ssDNA), but cannot perform the ATP-dependent strand exchange reaction necessary for homologous recombination. This mutation blocks the pairing and strand exchange between homologous sequences to form heteroduplex DNA.
The recA1 mutation present in cloning strains like DH5α, prevents homologous recombination which results in:
- Prevention of plasmid multimerization due to intermolecular recombination between plasmids that can generate plasmid dimers and multimers which are unstable and can be lost during cell division in the absence of selection pressure. High-copy plasmids are particularly prone to forming these multimers.
- Prevention of intramolecular recombination due to homologous recombination within a single plasmid, thereby preventing deletion of specific DNA sequences.
- Prevention of plasmid-chromosome integration which lowers the risk of recombination between cloned plasmid DNA and the host cell’s chromosome.
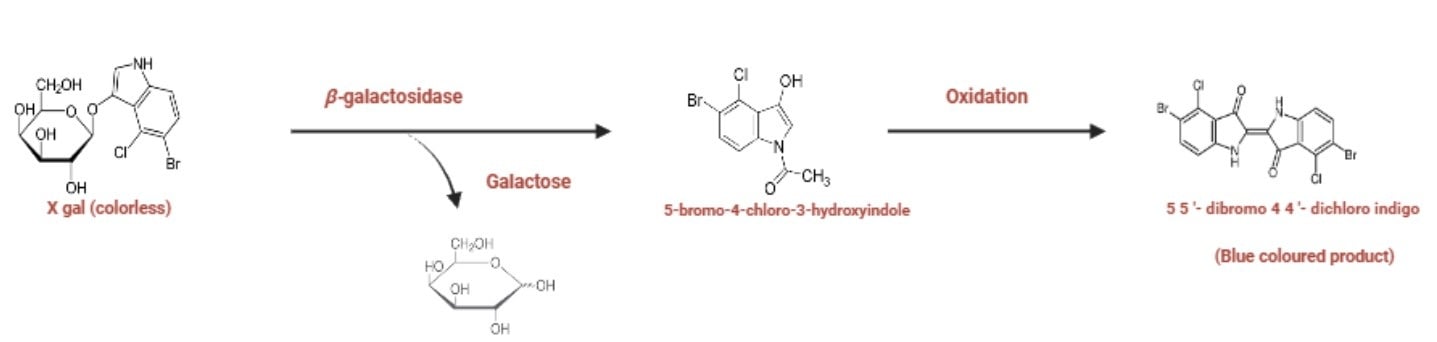
Blue White Screening: The Importance of the lacZΔM15 marker
Blue-white screening is a molecular biology technique used to distinguish between bacterial colonies that have successfully incorporated a recombinant plasmid from those that have not, by exploiting the α-complementation of the lacZΔM15 gene.
The lacZΔM15 marker is a specific deletion mutation within the lacZ gene of DH5α, which encodes the enzyme β-galactosidase. This enzyme allows bacteria to utilize lactose as an energy source in the absence of glucose, under normal conditions.
The lacZΔM15 mutation involves a small deletion in the N-terminal region of the lacZ gene specifically removing 11-41 or 23-31 amino acid residues.
The normal β-galactosidase enzyme is a tetramer with four identical subunits which need to fit together properly for the enzyme to become active. However, the M15 mutation removes amino acids critical for stabilizing the activating interface where the four subunits interact with eachother. Without these amino acids, the resulting protein known as α-acceptor or ω-fragment can only form dimers which are enzymatically inactive.
The Mechanism of Blue-White Screening
The blue white screening process utilizes the requirement of these missing N-terminal amino acid residues to restore enzymatic activity of β-galactosidase.
The host E. coli strain ( such as DH5α or DH10B) carries lacZΔM15 mutation on its chromosome whereas the cloning vector (plasmid) carries the DNA sequence encoding the missing N-terminal fragment, known as α-peptide or α-donor.
When the plasmid gets into the host cell, the α-peptide interact with the M15 dimers (α-acceptor), providing structural necessary for the assembly into active tetramers. If the plasmid produces α-peptide, the active β-galactosidase hydrolyzes the chromogenic substrate X-gal, creating an intensely blue product 5,5′-dibromo-4,4′-dichloro-indigo. However, if the foreign DNA is successfully inserted into the plasmid’s Multiple Cloning Site (MCS), which is located within the α-peptide sequence, it disrupts the reading frame. This prevents the production of active β-galactosidase as a result, complementation fails and colonies remain white.
Why DH5α is the Gold Standard for Cloning?
DH5α is the Gold Standard for Cloning because of its high transformation efficiency. Transformation efficiency refers to the number of colony-forming units (CFU) produced by 1 micorgram (μg) of plasmid DNA. Escherichia coli DH5α strain is genetically optimized for recombinant DNA work and has high transformation efficiency. Highly competent commercial DH5α cells typically achieve efficiencies of 109 colony-forming units (CFU) per microgram (μg) of plasmid DNA. Key genetic features responsible for high transformation efficiency include:
- recA1 – This mutation inactivates recA protein which results in the reduction of homologous recombination ensuring the stability of foreign plasmids without rearrangement or degradation.
- endA1– This mutation inactivates a non-specific endonuclease that leads to significantly higher quality DNA minipreps. In wild type, the enzyme often degrades plasmid DNA during isolation process.
- hsdR17(rK−,mK+) – This is the mutation of restriction-modification system of E. coli. With this mutation, the cells doesnot digest the foreign DNA (restriction negative/ rK−) while it can still be able to methylate its own DNA protecting it from accidental digestion (modification positive/ mK+).
- ϕ80dlacZΔM15 – Phage DNA with modified lacZ, which involves a specific modification where the lacZ gene is 94 bases shorter at the N-terminus than in the parental strain, enabling α-complementation for blue-white screening to easily identify bacteria carrying a plasmid.
Common Applications
- Used to clone a DNA fragment of interest into plasmid and amplify the plasmid DNA and transfer the plasmid to expression vector (such as BL21) for protein expression.
- Blue white screening for the visual identification of the recombinant clones.
- Used to create large collections of plasmid libraries (genomic libraries, mutagenesis libraries, cDNA libraries).
DH5α vs. BL21 Cells
| Properties | DH5α | BL21 |
| Type | Cloning strain | Expression strain |
| Strain origin | K-12 strain of E.coli | B strain of E.coli |
| Primary use | Cloning and plasmid maintainance | Expression of recombinant proteins |
| Key mutations | recA1, endA1, hsdR17(rK−,mK+) | Deficient in Lon and OmpT proteases, which prevents the degradation of produced proteins and often contains T7 RNA polymerase gene (as in BL21(DE3)) for strong, inducible expression of the gene. |
| recA1 | Non-functional | Functional allowing normal recombination |
| endA1 | Non-functional | Active endonuclease hence not ideal for plasmid preparation |
| Blue-white screening | Yes ( ϕ80dlacZΔM15) | No |
| Uses | Routine subcloning, plasmid amplification, site directed mutagenesis, plasmid libraries | Recombinant protein production, enzyme purification |
When to use cloning vs expression strains?
The choice of the strain depends on whether the primary goal is the stable maintainance and amplification of DNA or the high-level production of recombinant protein. So, for the production of recombinant proteins, expression strains are used whereas, cloning strains are used for plasmid maintainance and amplification.
Preparation of Competent Cells: Chemical vs. Electroporation Methods
Competent cells are those cells that have the ability to uptake exogenous DNA from the environment. Preparation of competent cells and their subsequent transformation is an important technique in molecular biology, which invlolves the isolation of a target DNA fragment or plasmid and making multiple copies of it inside the host cell (such as DH5α). There are two most commonly used transformation techniques; physical and chemical. Both the methods aim to temporarily permealize the bacterial cell membrane to allow the exogenous DNA to enter inside the cell.
Chemical
Chemical method is very convenient in molecular cloning experiments because it is simple and inexpensive. The classical chemical method for transformation was first published by Mandel and Higa (1970), which is still a major choice in most laboratories and it yields transformation efficiencies of 105 to 107 cfu/µg of plasmid DNA.
Classic chemical method generally comprises of three major steps:
- Stimulate or induce recipient cells using calcium chloride to make them “competent’
- Transform competent cells with DNA by applying heat shock
- Screening transformants on selective medium
Mechanism
Both DNA and the bacterial cell membrane (particularly Lipopolysaccharide, LPS in gram-negative bacteria) are negatively charged, which results in elecrostatic repulsion. Divalent cations (Ca2+) bind to the negatively charged phosphate groups as well as other groups ( such as hydroxyl and carboxyl) on the cell membrane causing neutralization effect. This neutralization results in the formation of a chemical bridge that promotes the contact between exogenous DNA and the cell membrane. These cations at low temperature weakens the membrane fluidity, forming complex with poly-hydroxybutyrate (PHB) and poly-inorganic phosphate (poly-P) on the membrane, that creates small pores through which DNA can pass. Moreover, the osmotic pressure plays a critical role in opening of the cell. As chloride ions enter the cell, the influx of water causes temporary swelling of the bacterial cell, making it more permeable to the exogenous DNA. Finally, the rapid shift of temperatue from cold to hot (42°C) induces the transient opening of gated membrane channels, allowing DNA to enter the cytoplasm.
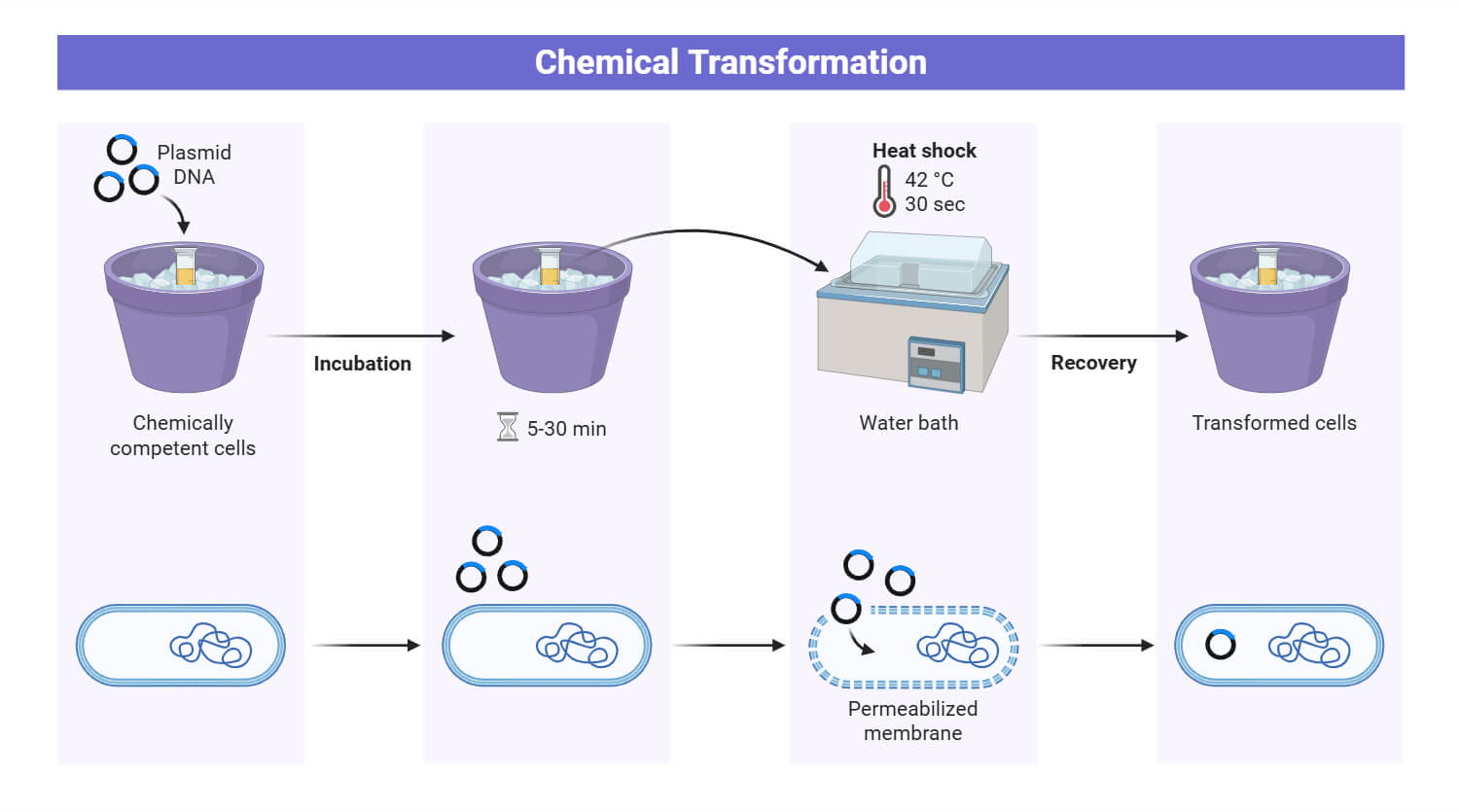
Physical
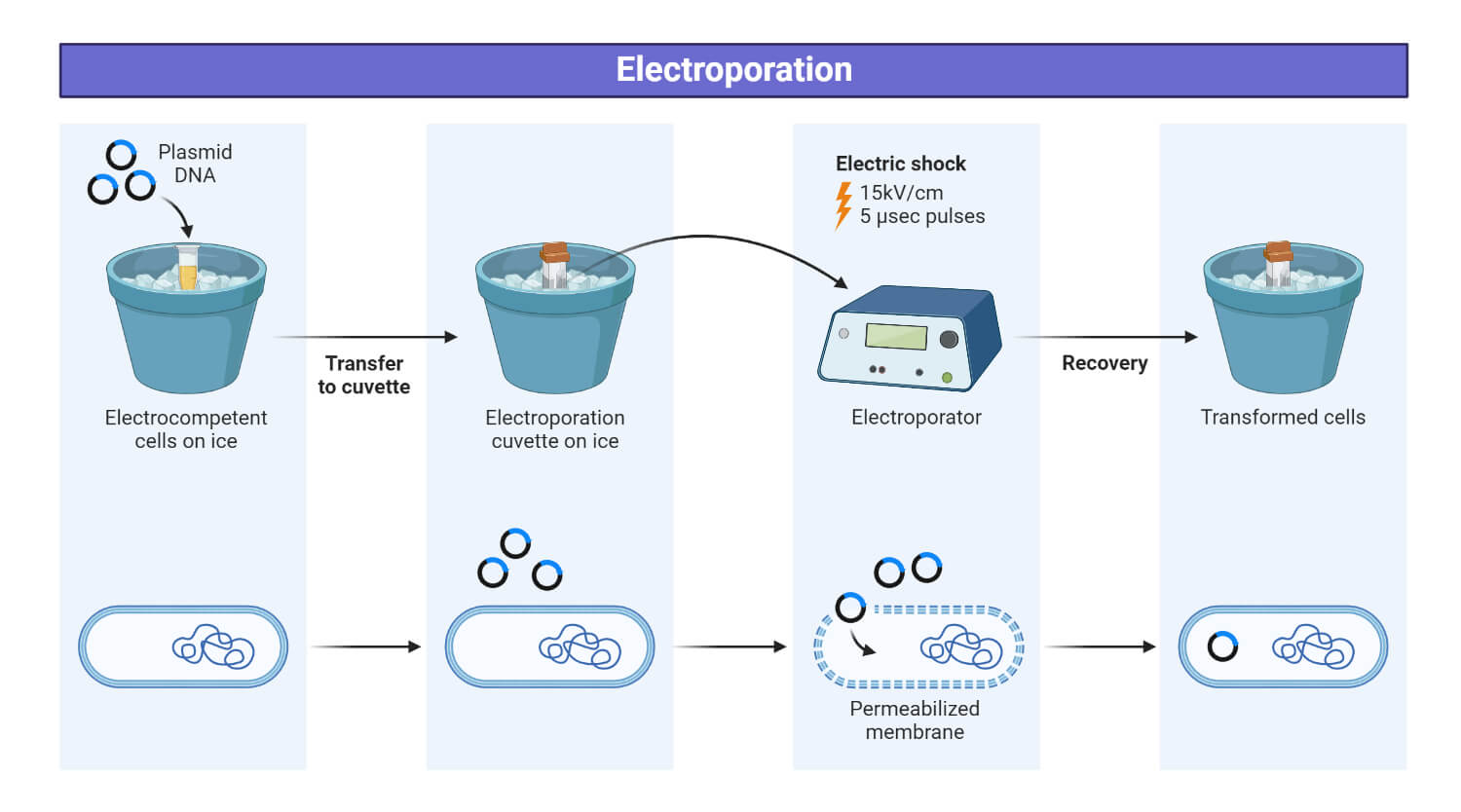
Electroporation is the most common physical method of transformation. It is also known as electro-transformation. This method utilizes high intensity electrical fields to introduce exogenous DNA into the receptor cell. It has exceptional transformation efficiency typically ranging from 109 to 1010 transformants/ µg DNA. Electroporation requires a very high cell density, at least 2×104 to 4 × 1010 cells/mL.
Mechanism
The intense electric field causes transient permealization of the cellular membrane, forming pores as a result the exogenous DNA is driven through pores by electrophoretic force.
Chemical vs. Electroporation
| Properties | Chemical transformation | Physical method (Electroporation) |
| Mechanism | Uses divalent cations (Ca2+) to create an ionic bridge and heat shock to open gated channels. | Uses high intensity electrical fields to introduce exogenous DNA into the receptor cell. |
| Transformation efficiency | 105 to 107 cfu/µg of plasmid DNA | 109 to 1010 cfu/µg of plasmid DNA |
| Requirement | Water bath, ice bath | Electroporator |
| Best for | Routine cloning and small plasmid vectors | High-complexity libraries and Large DNA fragments (BACs) |
| Salt tolerance | High: not affected by salts in DNA preparation | Very low: salts cause electrical sparking which can kill the cells |
| Cost | Inexpensive | Expensive |
Troubleshooting: Common Challenges with DH5-Alpha Growth and Transformation
Competent cells with low transformation efficiencies can cause few or no growth on the plates. To solve this, the transformation efficiency of an uncut plasmid of known concentration such as pUC19 can be calculated. If the transformation efficiency is lower than <104 cfu/μg DNA, the competent cells need to be remade or commercially available high efficiency competent cells are recommended.
The most common problem during electroporation is the electrical sparkling due to high salt contamination from the DNA or the growth media. In order to solve this, the cells must be washed extensively with 10% glycerol or water.
High concentrations of plasmid DNA can have a toxic effect on cells, decreasing overall efficiency. The optimal amount for DH5α is often around 5–10 ng.
The plasmid size can also affect the transformation efficiency. Tranformation using small plasmid has higher efficiency as compared to larger plasmid. To transform larger plasmid, right competent cells need to be used and also electroporation method can be employed for higher efficiencies.
Temprary pores are formed in the cell membrane of transformed cells due to heat-shock step or electric pulse ,which ultimately leads to stress. To recover the cells from this stress, a nutrient-rich medium, such as SOC medium (Super Optimal Broth with Catabolite Repression Medium) is required. SOC medium is preferred over LB because, it has richer nutrients, magnesium ions, glucose that promotes faster cell repair, plasmid establishment and antibiotic resistance gene expression, resulting in higher transformation efficiency.
Temperature also plays a key role during the bacterial transformation especially while working with chemically competent cells. Any deviation from the standard heat-shock protocol can negatively affect the transformation efficiency. Following transformation, the cells need to be incubated at an optimal growth temperature (37 °C), in order to recover the cells and antibiotic resistance gene. This recovery can be perfomed in a shaking incubator so that the nutirents get evenly distributed in the growth medium.
Incorrect antibiotic or antibiotic concentration can affect the number of colonies on the plate after transformation. Using the wrong antibiotic causes no colonies on the plate whereas, too low concentration of antibiotic cause satellite colonies (non-transformed bacterial colonies on the plate forming a uniform layer). Selectable marker in the plasmid needs to be checked properly before perfroming the transformation. Appropriate concentration of antibiotic needs to be used while performing the transformation.
Conclusion
DH5α remains a fundamental tool in molecular biology due to its useful genetic manipulations including recA1, endA1,hsdR17(rK−,mK+),ϕ80dlacZΔM15, that results in high transformation efficiency, genetic stability of the plasmid for routine cloning works. Use of DH5α cells allow researchers to amplify plasmid with minimal degradation or unwanted recombination while, facilitating blue-white screening for the visual identification of the recombinant clones. Even though, DH5α cells cannot be used for protein expression, it serves as an ideal host for constructing as well as maintaining recombinant DNA and later transferred to expression strain like BL21, making DH5α a gold standard for plasmid propagation and a foundation of modern molecular cloning strategies.
References
- Alves, L. D. F., Borelli, T. C., Westmann, C. A., & Silva-rocha, R. (2020). Boundaries in metagenomic screenings using lacZ a -based vectors. 20180252, 1–9.
- Borja, G. M., Mora, E. M., Barrón, B., Gosset, G., Ramírez, O. T., & Lara, A. R. (2012). Engineering Escherichia coli to increase plasmid DNA production in high cell-density cultivations in batch mode. 40, 1–9.
- Cha, D. O., & Rubens, C. E. (1998). Blue / white screening of recombinant plasmids in Gram-positive bacteria by interruption of alkaline phosphatase gene ( phoZ ) expression. 219, 91–99.
- Chen, X., Guo, P., Xie, Z., & Shen, P. (2001). A convenient and rapid method for genetic transformation of E . coli with plasmids. 297–300.
- Fathi-roudsari, M., Akhavian-tehrani, A., & Maghsoudi, N. (2016). Comparison of Three Escherichia coli Strains in Recombinant Production of Reteplase. 8(1), 16–22.
- Gonzales, M. F., Brooks, T., Pukatzki, S. U., & Provenzano, D. (2013). Rapid Protocol for Preparation of Electrocompetent Escherichia coli and Vibrio cholerae. October, 6–11. https://doi.org/10.3791/50684
- Griffith, M., Gietz, R. D., Griffith, M., & Gietz, R. D. (2018). Escherichia coli endA deletion strain for use in two-hybrid shuttle vector selection Escherichia coli endA deletion strain for use in two-hybrid shuttle vector selection. 6205. https://doi.org/10.2144/03352bm05
- Group, M. B., & Medical, S. (1988). Nucleic Acids Research. V(13), 6127–6145.
- Itaya, M., Kawata, Y., Sato, M., Tomita, M., & Nakahigashi, K. (2012). Rapid communication A simple method to provide a shuttling plasmid for delivery to other host ascertained by prolonged stability of extracellular plasmid DNA released from. 152(6), 501–504. https://doi.org/10.1093/jb/mvs111
- Juers, D. H., Matthews, B. W., & Huber, R. E. (2012). LacZ b -galactosidase : Structure and function of an enzyme of historical and molecular biological importance. 21. https://doi.org/10.1002/pro.2165
- Kostylev, M., Otwell, A. E., Richardson, R. E., & Suzuki, Y. (2015). Cloning Should Be Simple : Escherichia coli DH5 α -Mediated Assembly of Multiple DNA Fragments with Short End Homologies. 1–15. https://doi.org/10.1371/journal.pone.0137466
- Liu, Z., Tyo, K. E. J., Martínez, J. L., Petranovic, D., & Nielsen, J. (2012). Different expression systems for production of recombinant proteins in Saccharomyces cerevisiae. Biotechnology and Bioengineering, 109(5), 1259–1268. https://doi.org/10.1002/bit.24409
- Monk, J. M., Koza, A., Campodonico, M., Machado, D., Miguel, J., Palsson, B. O., Herrgård, M. J., & Feist, A. M. (2017). HHS Public Access. 3(3), 238–251. https://doi.org/10.1016/j.cels.2016.08.013.Multi-omics
- Nakata, Y., Tang, X., & Yokoyama, K. K. (n.d.). Preparation of Competent Cells for High-Efficiency. 69.
- Nishimura, A., Moritai, M., Nishimura, Y., & Sugino, Y. (1990). A rapid and highly efficient method for preparation of competent Escherichia coli cells. 18(20), 6169.
- Nováková, J., Izsáková, A., & Grivalský, T. (2013). Improved method for high-efficiency electrotransformation of Escherichia coli with the large BAC plasmids. 1988. https://doi.org/10.1007/s12223-013-0267-1
- Phue, J., Lee, S. J., Trinh, L., & Ã, J. S. (2008). Modified Escherichia coli B ( BL21 ), a Superior Producer of Plasmid DNA Compared With Escherichia coli K ( DH5 a ). 101(4), 831–836. https://doi.org/10.1002/bit.21973
- Plant, T. E., Co, T., & City, T. (1990). High efficiency transformation of E s c h e r i c h i a coli with plasmids Hiroaki Inoue ’, Hiroshi Nojima b and Hiroto 0kayama b ( c ) Optimization of transformation buffer. 96, 23–28.
- Reddy, M., & Reddy, M. (2018). Positive Selection System for Identification of Recombinants Using α-Complementation Plasmids Positive selection system for identification of recombinants using α -complementation plasmids. 6205. https://doi.org/10.2144/04376ST03
- Sadeghi, S., Ahmadi, N., & Esmaeili, A. (2017). RSC Advances Blue-white screening as a new readout for deoxyribozyme activity in bacterial cells †. RSC Advances, 7, 54835–54843. https://doi.org/10.1039/C7RA09679H
- Schweizer, H. P., & Schweizer, H. P. (2018). Bacterial Genetics : Past Achievements , Present State of the Field , and Future Challenges Bacterial genetics : past achievements , present state of the field , and future challenges. 6205(May). https://doi.org/10.2144/000112807
- Sharma, A., Girdhar, A., & Srivastava, N. (2011). Development of strategy for competent cell preparation and high efficiency plasmid transformation using different methods. 17–20.
- Shuman, H. A., & Silhavy, T. J. (2003). THE ART AND DESIGN OF GENETIC SCREENS : ESCHERICHIA COLI. 4(June).
- Tang, X., Nakata, Y., Li, H., Zhang, M., Gao, H., & Fujita, A. (1994). The optimization of preparations of competent cells for transformation of E . coli. 22(14), 2857–2858.
- Tu, Q., Yin, J., Fu, J., Herrmann, J., Li, Y., & Yin, Y. (2016). Room temperature electrocompetent bacterial cells improve DNA transformation and recombineering efficiency. Nature Publishing Group, April, 1–8. https://doi.org/10.1038/srep24648
- Wang, Y., Wang, X., Yu, L., Tian, Y., Li, S., Leng, F., Ma, J., & Chen, J. (2020). Effects of Sr 2 + on the preparation of Escherchia coli DH5 α competent cells and plasmid transformation. 1970, 1–20. https://doi.org/10.7717/peerj.9480Zamenhof, P. J., & Villarejo, M. (1972). Construction and Properties of Escherichia coli Strains Exhibiting a-Complementation of f- Galactosidase Fragments In Vivo. 110(1), 171–178.
