Competent cells are cells (bacteria, yeast, etc.) that have been altered to take up exogenous foreign DNA from the environment. While some bacteria can undergo transformation naturally, common laboratory strains like Escherichia coli cannot transform naturally and must have competency artificially induced.
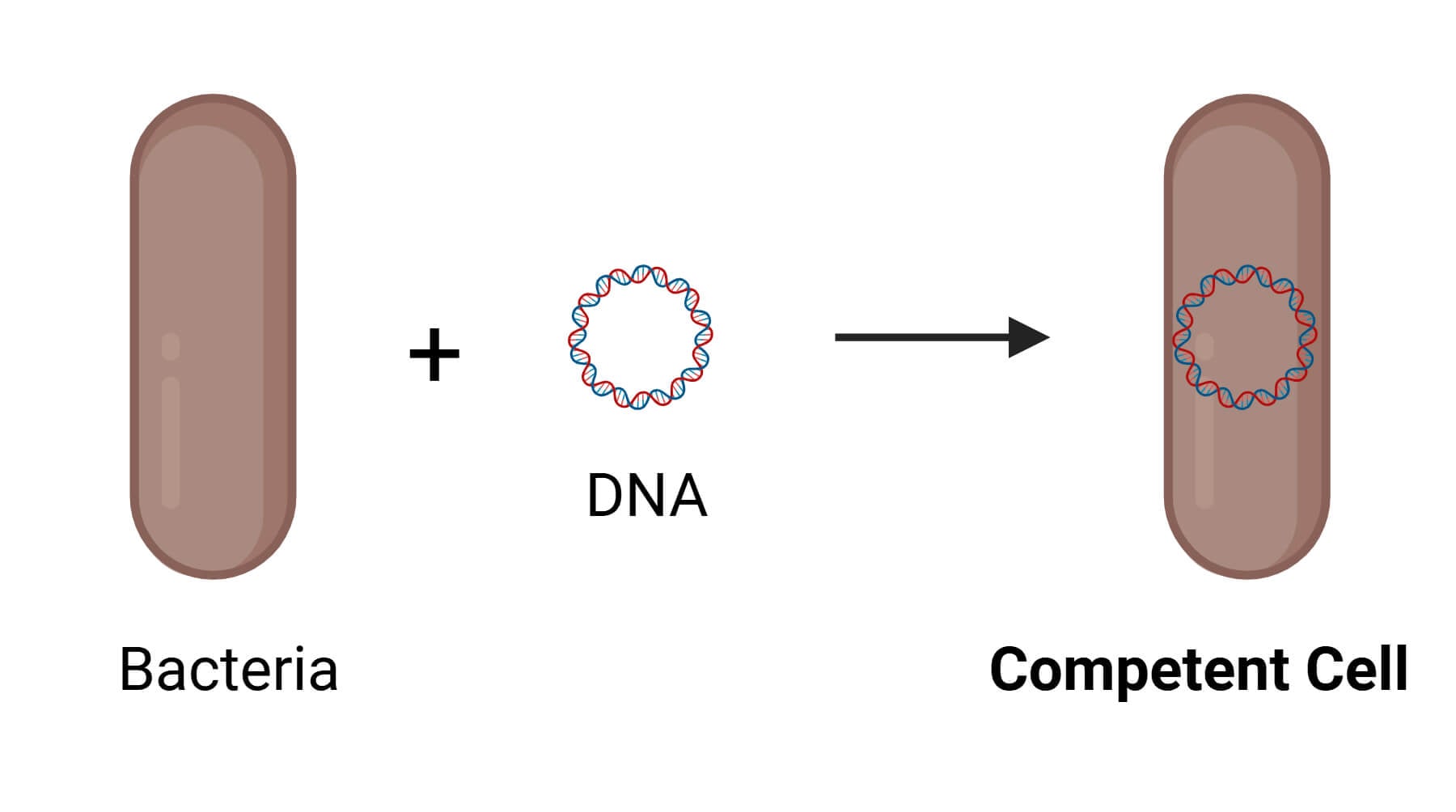
Applications of Competent Cells
- Used to clone and amplify fragments of genes or entire genes into plasmid vectors.
- Serve as hosts to replicate and store recombinant plasmids as bacterial stocks for future use.
- Essential for inserting PCR-amplified DNA into specific plasmids for further study.
- Essential for creating genomic and cDNA libraries, which involve storing and propagating large collections of DNA fragments.
- Cloning large inserts (eg, Bacterial Artificial Chromosomes (BAC)).
- Expression of recombinant protein.
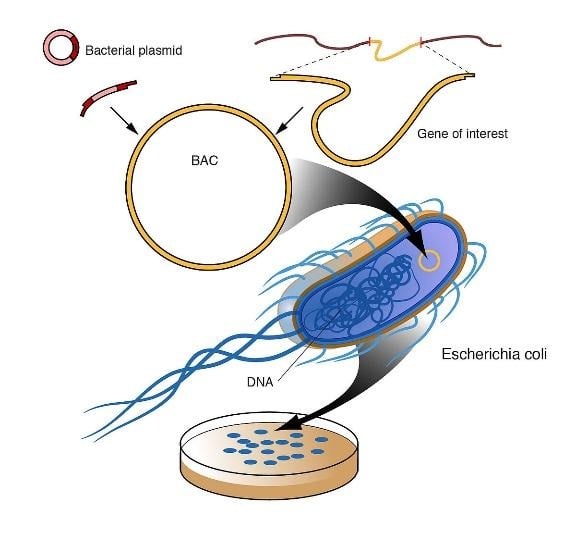
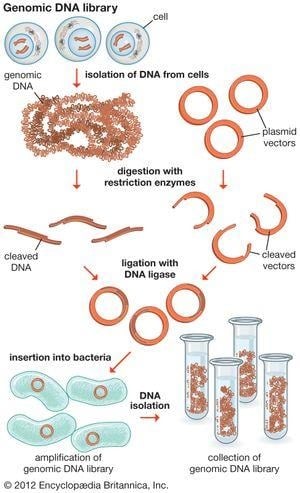
https://cdn.britannica.com/46/117846-050-722729F2/DNA-library-fragments-organism-genome-cells-cloning.jpg
Chemical Competency vs. Electro Competency: Choosing the Right Method
The choice between chemically competent and electrocompetent cells typically depends on the required transformation efficiency, the available equipment, and the specific downstream applications.
| Feature | Chemical competency | Electrocompetency |
| Primary advantage | Low cost: no special equipment required | High transformation efficiency |
| Typical efficiency | 1 x 106 to 5 x 109 CFU/µg | 1 x 1010 to 3 x 1010 CFU/µg |
| Complexity | Laborious preparation steps. | Quick but requires specialized hardware(electroporator) |
| Ideal Application | General cloning and automated screening. | Libraries, large DNA fragments, and recombineering. |
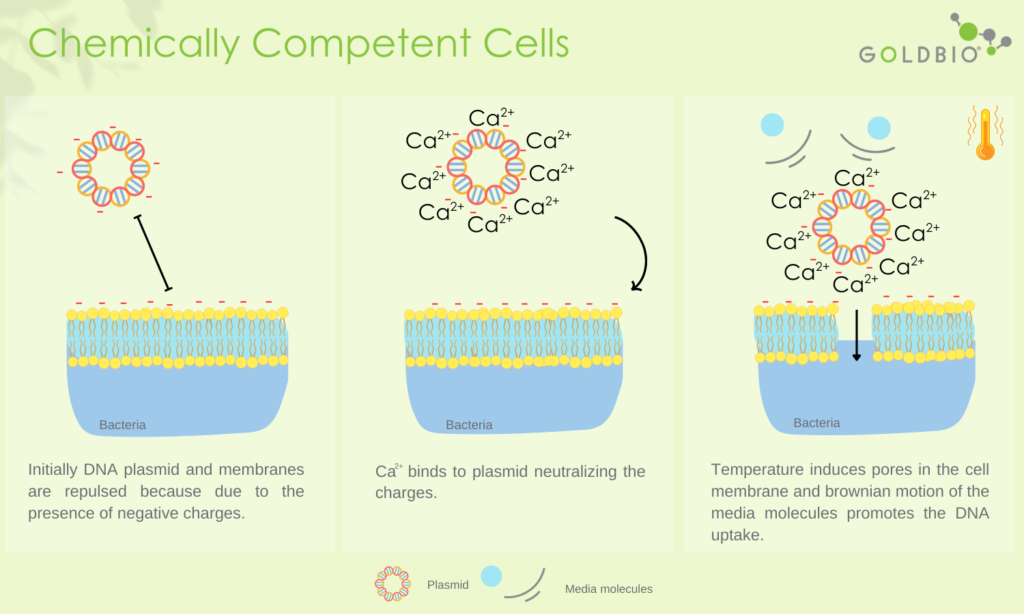
The Science Behind the Calcium Chloride (CaCl2) Method
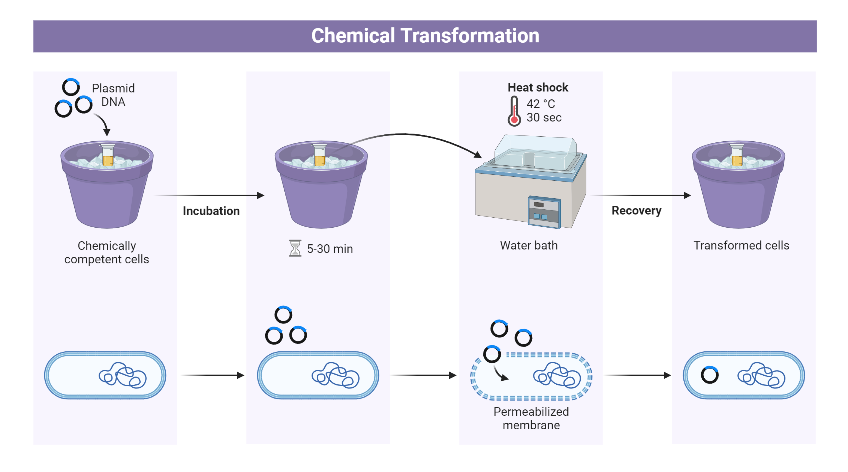
Both DNA and the bacterial cell membrane (particularly Lipopolysaccharide, LPS in Gram-negative bacteria) are negatively charged, resulting in electrostatic repulsion. Divalent cations (Ca2+) bind to the negatively charged phosphate groups as well as other groups ( such as hydroxyl and carboxyl) on the cell membrane, causing a neutralization effect. This neutralization results in the formation of a chemical bridge that promotes contact between exogenous DNA and the cell membrane. These cations at low temperature weaken the membrane fluidity, forming complexes with poly-hydroxybutyrate (PHB) and poly-inorganic phosphate (poly-P) on the membrane, which creates small pores through which DNA can pass. Moreover, the osmotic pressure plays a critical role in the opening of the cell. As chloride ions enter the cell, the influx of water causes temporary swelling of the bacterial cell, making it more permeable to the exogenous DNA. Finally, the rapid shift of temperature from cold to hot (42°C) induces the transient opening of gated membrane channels, allowing DNA to enter the cytoplasm.
The Role of Optical Density: Why Harvesting at OD600 0.4-0.6 Matters
- Harvesting bacterial cells at an optical density (OD600) of 0.4 to 0.6 is a critical step because this range typically represents the early to mid-logarithmic (exponential) growth phase, which is the physiological state where cells are most susceptible to transformation.
- Cells in the early to mid-exponential phase have membranes that are more easily altered by chemical treatments (like calcium chloride) or electrical pulses to allow exogenous DNA to enter.
- During this phase, cells are actively growing, which are naturally more “susceptible” to the induction of competency.
- Once cells pass the mid-log phase and move toward the stationary phase, their physiological state changes, and they lose their capacity to be transformed.
Consequences of the overgrowth
- Harvesting cells too late in the growth curve increases the number of satellite colonies (non-transformed cells growing near real transformants), which reduces the overall quality and reliability of the transformation.
- Overgrown cells in cold-wash protocols can become more fragile and prone to death during the preparation process.
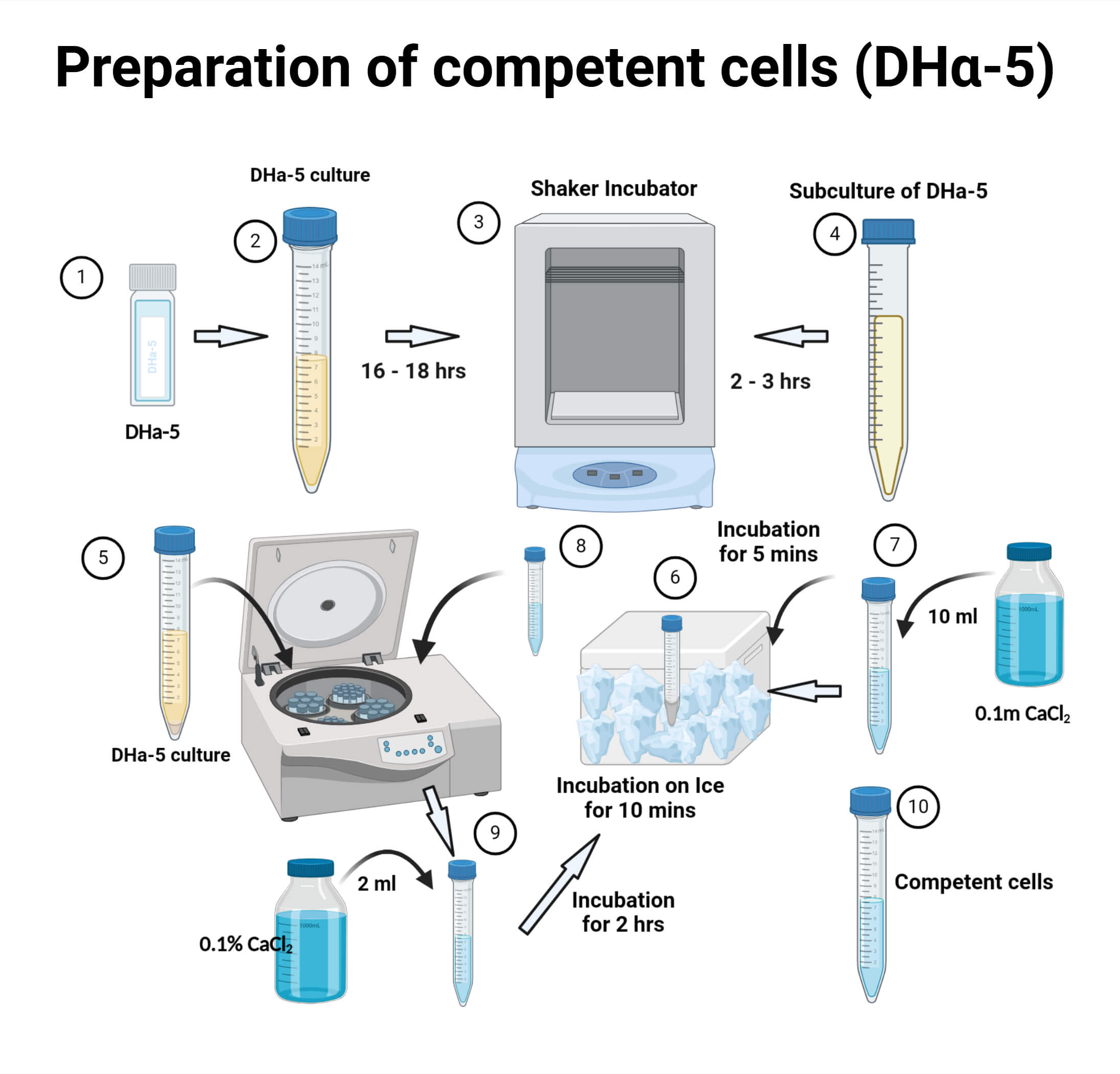
Step-by-Step Protocol: Preparing Chemically Competent E. coli
- Streak DH5α cells from a frozen glycerol stock onto a fresh LB or SOB agar plate and incubate overnight at 37°C (16-18 hours).
- Take a single well-isolated colony from the plate and inoculate it into 2-5 mL of SOB/LB medium and incubate at 37°C overnight with vigorous shaking at 200 rpm.
- Inoculate 50 mL of freshly prepared LB /SOB broth with 500 µL of the overnight starter culture and incubate at 37°C (200 rpm) until the culture reaches an optical density (OD of 0.5. This typically takes about 4 hours.
- Immediately place the culture on ice for 10 minutes to stop growth.
- Spin the cells at 2,000 x g for 10 minutes at 4°C.
- Discard the supernatant. Gently resuspend the pellet in 15 mL of ice-cold 0.1 M MgCl2
- Incubate the mixture on ice for 30 minutes.
- Centrifuge the cells at 4,000 rpm for 8–10 minutes at 4°C.
- Discard the supernatant. Gently resuspend the final pellet in 4 mL of 0.1 M CaCl2 containing 20% (v/v) glycerol and aliquot 100 μL in different epitubes, store at -80°C.
Critical Reagents: The Importance of DMSO and Glycerol
| Reagents | Importances |
| DMSO (Dimethyl sulphoxide) | DMSO helps in altering the permeability of the bacterial cell membrane, making it easier for exogenous DNA to enter the cytoplasm, hence significantly increasing transformation efficiency, and also helps in stabilizing the buffer. |
| Glycerol | Serves as a cryoprotectant for long-term storage at -80 °C, so that it prevents ice crystals from rupturing cell membranes |
Storage and Handling: Flash Freezing with Liquid Nitrogen
Flash freezing with liquid nitrogen is a critical step in the preparation of high-quality competent cells, ensuring that the induced physiological state is halted for future use. After the final resuspension in transformation buffer, cells should be dispensed into small volumes (typically 50 to 100 μl) in sterile microcentrifuge tubes (pre-chilled on ice ). Once aliquoted, the tubes must be immediately immersed in liquid nitrogen (−196 °C), called flash freezing. Immediately after flash freezing, cells should be moved to long-term storage at -80 °C.
Quality Control: Calculating Transformation Efficiency
Measuring transformation efficiency is the primary quality control (QC) test for competent cells. Transformation Efficiency (TrE) is a quantitative measurement used to determine how many bacterial cells successfully took up a plasmid during a transformation protocol. It is expressed as colony-forming units per microgram (cfu/µg) of plasmid DNA. By definition, transformation efficiency is the number of colony-forming units produced by transforming 1 µg of plasmid DNA into a given volume of competent cells.
TrE =Number of colonies (cfu)Amount of DNA used (µg) ×Dilution Factor
Where,
Number of Colonies (cfu): The manual count of bacterial colonies growing on the selective agar plate
Amount of DNA used (µg): The mass of the plasmid added to the competent cells, converted to micrograms
Dilution Factor: This accounts for the fraction of the total recovery mixture that was actually spread on the plate
Example: Calculation of Transformation Efficiency (TrE)
Given:
Amount of plasmid DNA used = 10 ng
Total recovery volume after transformation = 1000µL
Volume plated on LB + antibiotic plate = 100 µL
Number of colonies observed on the plate = 120 colonies
Step 1: Convert the DNA amount to micrograms
10 ng=0.01 µg
Step 2: Calculation of dilution factor
Since 100 µL was plated,
Dilution factor = 1000/100 = 10
Step 3: Apply the transformation efficiency formula
TrE =Number of colonies (cfu)Amount of DNA used (µg) ×Dilution Factor
=1200.01 × 10
= 120,000 cfu/µg
Hence, transformation efficiency = 120,000 cfu/µg
≥ 10⁹ CFU/µg – Excellent (high efficiency, cloning grade)
< 10⁵ CFU/µg – Poor ( not recommended)
Troubleshooting and Common Issues
Common issues encountered during the preparation of competent cells include:
- Harvesting at the Wrong OD: One of the most critical factors is harvesting cells in the early to mid-logarithmic (exponential) growth phase. Harvesting too early results in lower cell density, whereas harvesting cells once they have entered the late log or plateau/stationary phase significantly reduces or eliminates transformation capacity and increases the risk of formation of satellite colonies.
- Incorrect heat shock temperature: Temperatures higher than 42 °C can be lethal to the bacteria, while a heat shock that is too short may fail to facilitate DNA entry.
- Buffer freshness: All the buffers need to be freshly prepared.
- Conductivity and Arcing: During electroporation, if salts/conductive solutes are not thoroughly washed away, arcing (electrical shorting) occurs, which ruins the experiment and kills the cells.
- Slow Freezing: For long-term storage, cells must be flash-frozen using liquid nitrogen. Slow freezing can lead to ice crystal formation that ruptures membranes.
- Defrosting and Refreezing: Competent cells cannot be defrosted more than once; a single thaw-refreeze cycle causes a loss in competency.
- DNA Size: Transformation efficiency naturally decreases as the size of the DNA molecule increases.
- DNA Concentration: Adding an excessive volume or concentration of DNA can actually cause a decrease in transformation efficiency.
Conclusion
Competent cells are a very important component in molecular biology, enabling the uptake of exogenous DNA for routine cloning, DNA library construction, large-insert maintenance, and recombinant protein expression. The success of transformation largely depends on selecting the appropriate competency method, chemical or electrocompetent, based on the required efficiency and experimental objectives. A clear understanding of the mechanism of the role of divalent cations in the CaCl₂ method, proper harvesting at OD₆₀₀ (0.4–0.6), careful buffer preparation, and correct heat-shock or electroporation conditions is essential for achieving high transformation efficiency.
Additionally, critical reagents like DMSO and glycerol, proper flash freezing, and correct storage practices help preserve competent cells for future use. Quality control through transformation efficiency calculation ensures reliability and reproducibility of results. By carefully optimizing each step from culture growth to storage, researchers can consistently prepare high-quality competent cells suitable for routine cloning as well as advanced applications such as genomic library construction and recombinant protein production.
References
- Chan, W., Verma, C. S., David, P., & Gan, S. K. (2013). A comparison and optimization of methods and factors affecting the transformation of Escherichia coli. https://doi.org/10.1042/BSR20130098
- Sharma, A., Girdhar, A., & Srivastava, N. (2011). Development of strategy for competent cell preparation and high efficiency plasmid transformation using different methods. 17–20.
- Chan, W., Verma, C. S., David, P., & Gan, S. K. (2013). A comparison and optimization of methods and factors affecting the transformation of Escherichia coli. https://doi.org/10.1042/BSR20130098
- Sharma, A., Girdhar, A., & Srivastava, N. (2011). Development of strategy for competent cell preparation and high efficiency plasmid transformation using different methods. 17–20
- Chan, W., Verma, C. S., David, P., & Gan, S. K. (2013). A comparison and optimization of methods and factors affecting the transformation of Escherichia coli. https://doi.org/10.1042/BSR20130098
- Sharma, A., Girdhar, A., & Srivastava, N. (2011). Development of strategy for competent cell preparation and high efficiency plasmid transformation using different methods. 17–20.
