Apoptosis, or programmed cell death, is a crucial biological process fundamentally distinct from necrosis, which is cell death resulting from acute injury or trauma. It is a highly controlled and organized process governed by specific molecular pathways within the cell.
This organized nature is evident in the characteristic cellular morphology displayed by dying cells, including cell shrinkage, membrane blebbing, chromatin condensation, and eventual fragmentation into membrane-bound apoptotic bodies. These distinctive modifications facilitate the efficient, non-inflammatory removal of the dying cells by nearby phagocytic cells. Crucially, apoptosis is an active, energy-dependent process that requires ATP to drive a series of biochemical reactions. Apoptosis is a non-inflammatory process that helps preserve tissue integrity by preventing the release of potentially hazardous cellular contents into the surrounding environment.
The term HtrA refers to both a gene and the protein it encodes. HtrA is an abbreviated form for High Temperature requirement A, and structurally all HtrA proteins feature a serine protease domain (for catalytic cutting) and at least one PDZ domain (for binding to target proteins, or substrates). It has a dual function: as a vital component of Mitochondrial Quality Control, promoting cell survival, and as Apoptosis, a critical molecular switch governing cellular fate.
Molecular Structure and Location of HtRA
In humans, there are four known members: HTRA1, HTRA2, HTRA3, and HTRA4. The name, High-temperature requirement A, stems from its initial discovery in E. coli, where the gene htrA was essential for survival under heat stress.
These four proteins are all serine proteases with similar sizes (around 50 kDa), but each is encoded by a gene on a different chromosome and plays slightly different, often location-specific, roles in the body.
| Protein | Location | Key Features |
| HtrA1 | Chromosome 10. The main protein is about 50 kDa (480 amino acids long). | It is mostly a secretory protein (sent outside the cell), but a smaller version (29 kDa) has been found inside the cell’s cytoplasm and nucleus. It was first noticed because its production was reduced in certain cancer-like cells |
| HtrA2 | Chromosome 2. About 49 kDa (458 amino acids long) | crucial for monitoring stress, primarily in the mitochondria. |
| HtrA3 | Chromosome 4. 49 kDa | It is strongly linked to pregnancy and placental development. It is unique because its gene can be read in two different ways (called alternative splicing) to make two different protein versions: HtrA3L (Long) & HtrA3S (Short) |
| HtrA4 | Chromosome 8. About 50 kDa (476 amino acids long) | Similar to HtrA3, it was first identified as a serine protease associated with pregnancy. |
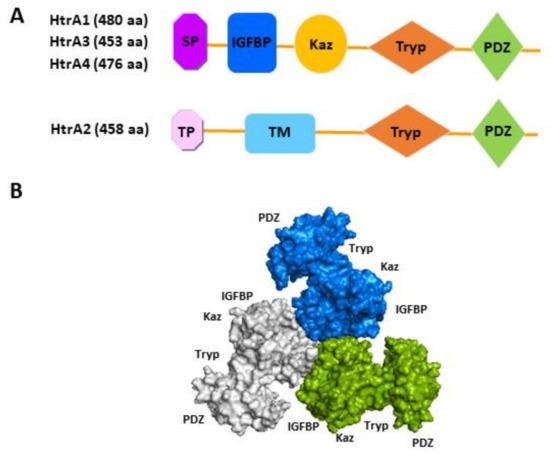
https://www.mdpi.com/1422-0067/22/19/10756
The HtrA proteins are built from several key parts:
- Trypsin-like Serine Protease Domain (PD): This is the catalytic core containing the catalytic triad (His-Asp-Ser) responsible for cutting (protease activity).
- C-terminal PDZ Domain: This domain acts as the sensor or regulatory module. Its primary job is substrate recognition and allosteric regulation of the protease activity.
- Variable N-terminal Regions: These regions dictate where the protein works:
- HtrA1, HtrA3, HtrA4: Have a Signal Peptide (SP) for secretion (extracellular location).
- HtrA2/Omi: Has a Mitochondrial Targeting Sequence (MTS) to target it to the mitochondria.
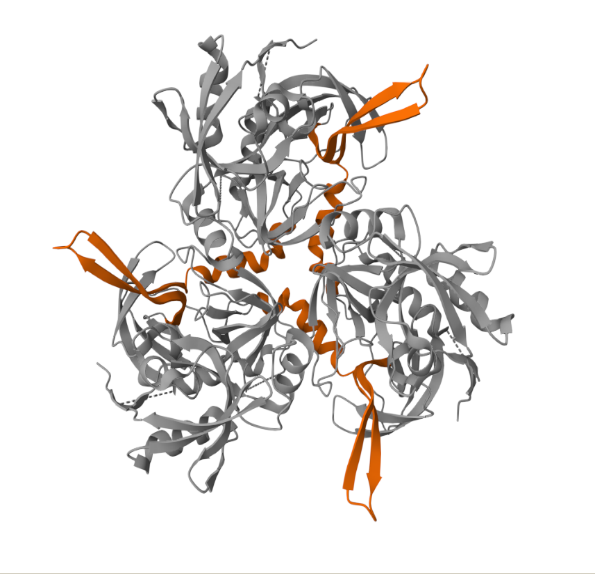
PDB ID: Crystal Structure of the Mitochondrial Serine Protease HtrA2 1LCY (pdb_00001lcy)
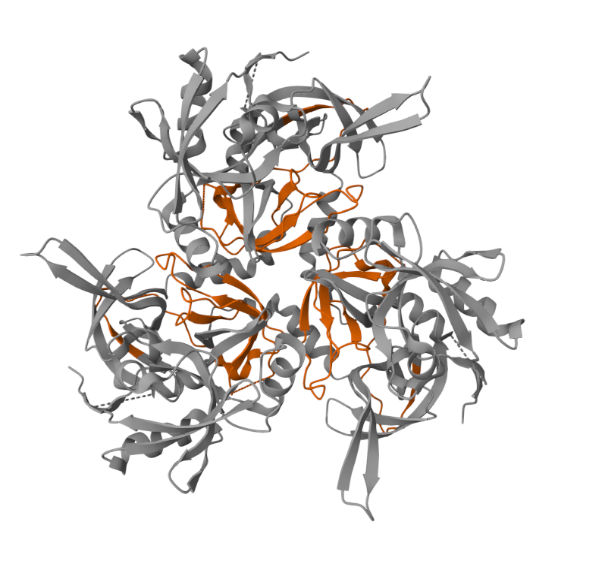
The displayed image shows a 3D structural model of a protein, of the HtrA family member, as viewed using a molecular visualization tool – PDBe. HtrA2 typically represents a homotrimer (three identical subunits) or a hexamer (two stacked trimers). In the first image, the orange segment represents PDZ domains. The image highlights exterior or flexible loops that might be critical for allosteric regulation, substrate binding, or interaction with other proteins (like chaperone complexes). In the second image, the orange-coloured segments are the Serine Protease Domains (PD). This domain is the enzyme’s catalytic “cutter.” It is classified as a trypsin-like peptidase (Peptidase S1, PA clan) and contains the crucial catalytic triad (His, Asp, Ser) that hydrolyses (cleaves) other proteins. The active site is located within a groove in the centre of the domain, facing the central cavity of the trimer.
PDB ID: Crystal Structure of the Mitochondrial Serine Protease HtrA2 1LCY (pdb_00001lcy)
The functional structure and control mechanism ensure the enzyme is only active when needed. The HtrA proteins assemble into a pyramidal homotrimer (three identical units). The three PD domains form the central core, while the three PDZ domains extend outward to scan for damaged proteins. The enzyme’s activity is controlled by the PDZ domain in an on/off switch. When the cell is healthy, the PDZ domain physically inhibits the protease active site, keeping the enzyme inactive. When a misfolded protein binds to the PDZ domain, it causes a conformational change (a structural shift). This shift breaks the inhibitory contact, opening the protease active site and activating the enzyme to degrade the damaged substrate. This regulation mechanism effectively links the protein’s sensing capacity (PDZ) with its degradation capability (Protease Domain), making it a highly specific quality control checkpoint.
The MTS is the critical feature that dictates the localization and final activation of the HtrA2/Omi protein, establishing it as a key mitochondrial stress regulator.
HtrA2’s Dual Functions: Protease & Apoptosis Inducer
Function as a Stress-Sensing Protease
- HtrA2 (Omi) is a vital, stress-responsive mitochondrial protease that safeguards cell health and prevents neurodegeneration by maintaining protein quality control. It works by recognizing and clearing damaged or misfolded proteins, a crucial function that ramps up during cellular stress (like heat shock or oxidative stress). This degradation process prevents toxic protein buildup, thereby sustaining proper mitochondrial function.
- It is an ATP-independent protease/chaperone that governs mitochondrial proteostasis (protein homeostasis).
- The Stringent proteolytic function of HtrA2 ensures it only targets damaged proteins. This precise regulation is managed by its PDZ domain through an allosteric mechanism.
- HtrA2 is typically found as an inactive trimer where the C-terminal PDZ domain acts as a structural lock, blocking access to the central serine protease domain’s catalytic site. The activation process is triggered by the presence of misfolded or damaged proteins
- The PDZ domain acts as a sensor, and it binds directly to specific sequences or exposed hydrophobic patches on the target substrate.
- This binding event generates an allosteric signal (a change in shape) that travels from the PDZ domain to the rest of the protein.
- This signal causes the active site to be unmasked (or “unlocked”) by remodeling the regulatory loops in the serine protease domain.
- This activation mechanism is a sophisticated quality control checkpoint, ensuring that HtrA2 only becomes fully active and degrades proteins when it directly detects the markers of stress and damage (the misfolded substrates) within the mitochondrial space.
Function as a Pro-Apoptotic Factor
While HtrA2/Omi primarily acts as a quality control protease to promote cell survival under mild stress, it switches to a pro-apoptotic role when the cell is confronted with severe, irreversible damage.
Mitochondrial Outer Membrane Permeabilization (MOMP)
- Integrity of the Mitochondrial Outer Membrane (MOM) is compromised when an apoptotic stimulus comes into contact. This permeabilization, often controlled by Bcl-2 family proteins, is an irreversible commitment to the intrinsic death pathway
- As the MOM ruptures, HtrA2 (along with other factors like Cytochrome c) is rapidly released from the mitochondria (IMS) into the cytosol.
Apoptosis Induction
- Once in the cytosol, HtrA2 promotes cell death via two main pathways:
- Caspase Activation (Caspase-Dependent): HtrA2 utilizes an exposed N-terminal motif AVPS to bind and neutralize Inhibitors of Apoptosis Proteins (IAPs), such as XIAP. By blocking the IAPs, HtrA2 effectively removes the inhibitory “brake” on the caspase enzymes, allowing them to activate and execute cell death.
- Direct Cleavage (Caspase-Independent): HtrA2 also employs its native protease activity to directly degrade vital cellular targets, including anti-apoptotic and cytoskeletal proteins, driving cell death independently of the main caspase cascade.
- In essence, the cytosolic release of HtrA2 signals the cell’s definitive transition from a “repair” state to a “self-destruct” state in the face of overwhelming mitochondrial damage.
Clinical Significance and Disease Association
- Parkinson’s Disease (PD): Mutations in the HTRA2 gene (e.g., the G399S mutation) are associated with increased risk of PD. Compromised HtrA2 function impairs mitochondrial quality control, leading to the death of dopaminergic neurons.
- Cerebral Small Vessel Disease (CSVD): HTRA1 mutations are a known cause of several serious cerebral small vessel diseases, such as CARASIL (Cerebral Autosomal Recessive Arteriopathy with Subcortical Infarcts and Leukoencephalopathy) and Autosomal Dominant CSVD
- Age-related Macular Degeneration (AMD): Polymorphisms (variations) in the HTRA1 gene are one of the strongest genetic risk factors for the development of the common “wet” form of AMD
- Alzheimer’s Disease (AD): Both HtrA1 and HtrA2 are implicated in AD pathology, with HtrA1 involved in degrading protein aggregates like APP (Amyloid Precursor Protein) and Tau.
- All four members of the HtrA family have associations with cancer, often acting as tumor suppressors due to their ability to promote apoptosis. Cancer cells frequently downregulate HtrA expression to evade cell death
Conclusion
While it is evident that HtrA2 degrades misfolded proteins, its full range of physiological substrates (the HtrA2″degradome”) in both the mitochondrial and cytosolic compartments remains largely undefined. Identifying these is crucial for understanding their exact role in PD progression and cancer suppression.
Research is urgently needed to develop drugs that can specifically promote HtrA2’s pro-apoptotic release or function to overcome chemotherapy resistance.
Compounds that can boost the neuroprotective quality control function of the mutant HtrA2 in the mitochondria to protect dopaminergic neurons from degeneration.
References
- Alberts, B., Johnson, A., Lewis, J., Raff, M., Roberts, K., & Walter, P. (n.d.). Molecular biology of the cell (4th ed.). Programmed Cell Death (Apoptosis). NCBI Bookshelf Programmed Cell Death (Apoptosis) – Molecular Biology of the Cell – NCBI Bookshelf
- Tzeng, S.-R., Tseng, Y.-C., Lin, C.-C., Hsu, C.-Y., Huang, S.-J., Kuo, Y.-T., & Chang, C.-I. (2021). Molecular insights into substrate recognition and discrimination by the N-terminal domain of Lon $\text{AAA}^+$ protease. eLife, 10, Article e64056.
- Yamaguchi, H., Kanzaki, S., Okui, Y., Tanaka, Y., & Hori, O. (2018). Mitochondrial Lon regulates apoptosis through the association with Hsp60–mtHsp70 complex. BMC Molecular Biology, 19(1), Article 22.Mitochondrial Lon regulates apoptosis through the association with Hsp60–mtHsp70 complex – PMC
- -Casal, R., Ferreira, F. V., Vitiello, P. F., Hoda, F., Pardo, J., & Cañedo, L. (2020). HTRA2/OMI-Mediated Mitochondrial Quality Control Alters Macrophage Polarization Affecting Systemic Chronic Inflammation. The Journal of Immunology, 205(6), 1629–1643.HTRA2/OMI-Mediated Mitochondrial Quality Control Alters Macrophage Polarization Affecting Systemic Chronic Inflammation – PMC
- National Center for Biotechnology Information. (n.d.). HtrA serine peptidase 2 Mus musculus (house mouse) (Gene ID 246524). NCBI Gene.Htra2 HtrA serine peptidase 2 [Mus musculus (house mouse)] – Gene – NCBI
- Cattaneo, P., D’Amico, M., Peverelli, G., Consonni, A., Ghitti, M., Bellinzoni, M., Palleschi, A., & De Mattia, F. (2020). Oligomeric assembly regulating mitochondrial HtrA2 function as examined by methyl-TROSY NMR. Molecules, 25(21), Article 5133.Oligomeric assembly regulating mitochondrial HtrA2 function as examined by methyl-TROSY NMR – PMC
- Structure of HtrA2/Omi.a, Schematic representation of the HtrA2/Omi… | Download Scientific Diagram
- HtrA serine proteases in cancers: A target of interest for cancer therapy – ScienceDirect
- Cieplak, P., & Dmochowska, A. (2018). $\text{HTRA2}$ gene: Function, regulation, and role in disease. Cellular and Molecular Life Sciences, 75, 2235–2247.HTRA2 Gene: Function, Regulation, and Role in Disease
- PDB DOI: https://doi.org/10.2210/pdb1lcy/pdb
- https://www.google.com/url?sa=i&url=https%3A%2F%2Fwww.mdpi.com%2F1422-0067%2F22%2F19%2F10756&psig=AOvVaw1hc6pXHRSYFHySRh6hgn6-&ust=1763398442201000&source=images&cd=vfe&opi=89978449&ved=0CBgQjhxqFwoTCOjP-J6R95ADFQAAAAAdAAAAABAU
