Structural bioinformatics is the computational study of the three-dimensional structures of biological macromolecules such as proteins, RNA, and DNA. It is based on the principle that a molecule’s physical structure determines its biological function.

The Sequence – Structure – Function framework delineates the phenomena of how a sequence of AGTC strings are folded into complex 3D shapes.
| Component | Specification | Role of Structural Bioinformatics |
| Sequence | The Blueprint | Predictive modelling and alignment of genetic code |
| Structure | The Machine | Atomic coordinate analysis, surface mapping, and simulation. |
| Function | The Task | Inferring biological roles based on shape and chemical pockets. |
By analysing data from various repositories, scientists use AI and physics-based models to predict how molecules fold, move, interact with drugs. This field is essential for rational drug design, understanding genetic diseases, and engineering new enzymes, effectively bridging the gap between raw genetic sequences and the complex machinery of life.
Fundamentals of Protein Structure: Primary, Secondary, Tertiary, and Quaternary
Proteins are large biological molecules composed of 20 different amino acids, categorized into essential and non-essential types. Protein structure is a hierarchical organization where the primary structure (the linear sequence of amino acids held by peptide bonds) directs how the chain will fold. This chain then develops into a secondary structure, forming local patterns like alpha helices or beta-pleated sheets through hydrogen bonding of the backbone. As these segments interact, the protein collapses into its tertiary structure, a complex 3D shape stabilized by various R-group interactions such as disulfide bridges and hydrophobic effects. Consequently, for many functional proteins multiple folded chains come together to form a quaternary structure creating a large multi-subunit machine.
The Protein Data Bank (PDB): Navigating the World’s Structural Archive
The Protein Data Bank (PDB) is the world’s primary open-access archive for the 3D shapes of proteins, DNA, and RNA. Since its start in 1971, it has become an indispensable archive for understanding the molecular machinery of life. It serves as a key repository for modern biology, housing hundreds of thousands of structures determined through experimental methods like X-ray crystallography and cryo-electron microscopy. Each entry is assigned a unique four-character PDB ID, which acts as a primary key for researchers to access atomic coordinates, chemical properties, and experimental metadata.
Navigating this vast archive is simplified by a clear structural hierarchy: an Entry contains the full dataset, an Entity identifies unique molecules, and Assemblies represent the functional biological complexes. This organization is critical for drug discovery, as it allows scientists to visualize precise lock-and-key interactions between proteins and potential therapeutic compounds. Beyond pharmacology, the PDB provides the essential training data for revolutionary artificial intelligence tools like AlphaFold. By transforming raw experimental data into navigable spatial maps, the PDB enables scientists to decode the molecular mechanisms of disease and advance biotechnology.
Experimental Methods: X-ray Crystallography, NMR, and Cryo-EM
- X-ray Crystallography: X-ray crystallography is often called the gold standard because it provides the highest possible resolution, it explicates individual atoms and the chemical bonds between them. The process begins by purifying and concentrating proteins until they form crystals, where many copies of the protein align in symmetrical arrays until a constant lattice has been formed of the crystal. When an X-ray beam strikes these crystals, the rays scatter creating a unique pattern of spots called a diffraction pattern. This method provides the detailed 3D maps essential for understanding biological functions. The technique can reach resolutions below 1.5 Å, revealing the exact placement of water molecules and drug inhibitors. Crystallography works for proteins of almost any size, from small hormones to massive viral capsids. Because it shows the lock (protein) and key (ligand) so clearly it is the primary tool used by pharmaceutical companies to design new drugs. The primary disadvantage of X-ray crystallography is the difficult and unpredictable requirement to grow high-quality crystals, which often fails for flexible or membrane proteins. This results in a static snapshot of the molecule that may not fully represent how it moves and functions in its natural, liquid environment
- NMR: NMR stands for Nuclear Magnetic Resonance and is used as an alternative for X-ray crystallography since it allows the study of molecules in a liquid state, which is much closer to their natural biological environment. The sample is placed in a high-strength magnetic field, causing atomic nuclei (usually 1H, 13C, or 15N) to align. Radiofrequency pulses disturb these nuclei as they return to equilibrium, they emit signals (chemical shifts) that reveal their chemical environment. Using the Nuclear Overhauser Effect (NOE), measurement of how close atoms are to one another through space has become easy. These distances act as physical constraints. Software then calculates a 3D ensemble of structures that best fit the data. Unlike static X-ray images, NMR can track how a protein moves, and folds in real-time.
- Cryo- EM: One of the emerging and revolutionary imaging techniques used to determine the 3D structure of large biomolecules. Cryo-EM excels at visualizing massive molecular machines like viruses, ribosomes, and membrane proteins that are too large for NMR and too difficult to crystallize for X-ray studies. Cryo-EM is an abbreviated form of Cryo-Electron microscopy where Cryo stands for cryogenic conditions maintained for the biological sample mostly at -180 °C in liquid ethane or liquid nitrogen. Proteins are essentially wet molecules that rely on a surrounding hydration shell of water to maintain their complex 3D shapes or they would shrivel up and collapse. To study them in an electron microscope, protein must be kept hydrated, but liquid water would instantly evaporate in the microscope’s vacuum. To solve this problem, cryo-freezing is used to turn the water into vitreous ice. Unlike regular ice, which forms jagged crystals that would crush the protein, this glassy ice forms so fast that it locks the protein and its water preserves in place without damaging them. This effectively hits pause on the molecule, preserving its natural, liquid-state structure in a solid form that can survive the microscope’s harsh environment.
Protein Structure Prediction: Homology Modelling and Threading
Homology modelling: Homology modelling is one of the fundamental methods for predicting protein 3D structure merely based on comparative modelling of known structure (template) and the unknown sequence (target). The foundation of homology modelling is that structural evolution is slower than sequence evolution. Consequently, proteins derived from a common ancestor often retain a nearly identical 3D shape (fold) long after their amino acid sequences have significantly diverged. Protein tertiary structure is significantly more resilient to evolutionary pressure than primary sequence. While amino acid sequences undergo continuous mutational drift, the overarching three-dimensional fold is often preserved as a structural invariant because any significant conformational alteration would compromise biological activity. this phenomenon is often described as the divergence of sequence vs. the conservation of structure.
The process of homology modelling is a sequential, Integrated Protocol designed to transform a raw amino acid sequence into a high-resolution 3D atomic model using an evolutionarily related protein structure as a scaffold. The 5-step homology modelling workflow is as follows:
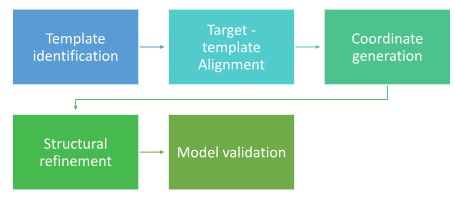
- The process begins by querying the Protein Data Bank (PDB) using tools like BLAST or HHsearch to find a high-resolution structural relative that shares significant sequence identity (ideally >30%) with your target.
- The second stage involves creating a residue-by-residue map to ensure conserved functional motifs are matched. Precision here is vital to ensure that gaps (insertions and deletions) are assigned to flexible loop regions rather than rigid secondary structures.
- In coordinate generation, the software transfers the X, Y, Z coordinates from the template’s backbone atoms to the target sequence. Missing loops are modelled ab initio, and side chains are positioned using rotamer libraries to achieve the most stable chemical configuration.
- To resolve stereochemical clashes where atoms may overlap, the raw model undergoes refinement. Using force fields or Molecular Dynamics, the structure is relaxed into a state of minimum potential energy.
- The final structure is audited for physical and biological plausibility. A Ramachandran Plot is used to verify backbone dihedral angles, while tools like ProSA or ERRAT confirm that the model’s global energy profile matches that of experimentally determined proteins.
Protein Threading: Protein Threading or Fold Recognition is a sophisticated modelling technique designed for the target proteins that lack identifiable relatives in nature. It operates in the structural Twilight Zone, where amino acid sequences have diverged so much (<25% identity) that simple sequence comparisons fail. Threading shifts the focus from evolutionary history to physical compatibility. It is guided by two fundamental observations:
- The Finite Fold Universe: Most new proteins discovered today simply reuse one of these established shapes.
- Thermodynamic Fitness: A protein’s sequence is fit for a structure if it can reside within that 3D shape at a low energy state.
Protein threading is a four-step predictive pipeline that evaluates how well an unknown target sequence fits into a library of known 3D templates.
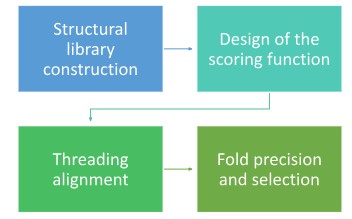
- The process begins by compiling a comprehensive database of unique, experimentally solved protein folds. These templates are sourced from the Protein Data Bank (PDB) and organized into structural classifications using databases like SCOP or CATH to ensure a wide variety of shapes are represented.
- An Intricate energy function is designed for measuring compatibility. This function doesn’t just look at letters it calculates the physical and chemical fitness of the target sequence within a fold by assessing Pairwise Potential, Solvation/Burial Potential, Secondary Structure Match.
- The target sequence is physically threaded through each template in the library. Unlike simple sequence alignment, this involves mapping each amino acid to a specific X, Y, Z coordinate on the template’s backbone and calculating the alignment’s total energy score.
- The software evaluates the scores from all possible alignments. The fold that results in the lowest global energy (the most stable physical fit) is identified as the likely structure. Once selected, this best-fit scaffold is used to generate the final 3D coordinates for the target protein.
The AI Revolution: AlphaFold and the Future of Structure Prediction
AlphaFold, represents a paradigm shift in structural biology by solving the 50-year-old protein folding problem from a linear string of amino acids. By applying deep learning to biological data, AlphaFold predicts a protein’s 3D structure from its primary sequence with accuracy comparable to traditional experimental methods like X-ray crystallography, which previously took years to perform. While AlphaFold 2 mastered single proteins, AlphaFold 3 has sparked a broader “AI revolution” by moving toward holistic biological systems. It replaces traditional rigid-body modelling with a generative diffusion model, enabling it to predict interactions between proteins, DNA, RNA, and small-molecule ligands. This leap transforms drug discovery into a predictive science, allowing researchers to visualize how a drug binds to its target in seconds.
Comparison of Predictive Methods:
| Feature | Homology Modelling | Threading | AlphaFold (AI) |
| Logic | Evolutionary relative | Physical fit (fold library) | Deep Learning/Diffusion |
| Identity | >30% (Safe) | <25% (Twilight Zone) | Independent of Identity |
| Speed | Minutes | Hours | Seconds |
| Scope | Related Proteins | Known Folds | Nearly all Biomolecules |
Molecular Docking: Simulating Drug-Target Interactions
Molecular docking is a computational technique that simulates the molecular recognition process to predict the preferred orientation and binding affinity between a ligand (usually a small-molecule drug) and a receptor (typically a protein). It serves as a digital lock-and-key experiment, identifying the most stable 3D configuration, or pose by minimizing the system’s Gibbs free energy.
The process relies on two pillars: a search algorithm, which explores thousands of potential spatial orientations and conformations, and a scoring function, which evaluates these poses based on physical forces like hydrogen bonding and van der Waals interactions. Depending on the complexity, docking can be rigid (fixed structures) or induced-fit (flexible structures).
By enabling virtual screening of vast chemical libraries, molecular docking helps to identify promising hit compounds in seconds rather than years. This dramatically reduces the cost and time of drug development, making it an indispensable tool in modern medicine.
Structure Visualization Tools: PyMOL, Chimera, and VMD
Protein structure visualization tools are essential for structural biologists since it transforms the abstract 3D coordinate data into intuitive, interactive models. Programs like PyMOL, UCSF Chimera, and VMD decode the complex architecture of biomolecules, investigate atomic interactions in drug binding, and communicate scientific discoveries through high-resolution imagery.
- PyMOL: It is widely regarded as the best tool for creating high-quality, static images for research papers and presentations. It includes a high-end Ray Tracer that adds realistic shadows and lighting to models, making them look professional.
- UCSF Chimera / ChimeraX: Chimera is favoured for complex analytical tasks and visualizing very large data sets. It analyzes molecular density maps (from Cryo-EM), visualizing large complexes (like viruses), and performing basic modelling tasks like mutagenesis. It can handle massive structures (millions of atoms) and overlay sequence alignments directly onto the 3D structure.
- VMD (Visual Molecular Dynamics): VMD was built to handle movement. It is the industry standard for viewing the results of Molecular Dynamics (MD) simulations. It is best for watching a protein fold or a drug enter a pocket and analysing large-scale motions over time. It is optimized to play thousands of frames of movement smoothly.
Evaluating Model Quality: Ramachandran Plots and RMSE
Evaluating the quality of a 3D protein model is a mandatory final step in the modelling. It ensures that the predicted structure is physically realistic and biologically plausible. Two of the most critical metrics for this are Ramachandran Plots (for local geometry) and RMSD (for global accuracy).
A Ramachandran plot is a 2D map used to evaluate protein model quality by plotting backbone dihedral angles: Phi Φ and Psi Ψ. Because of steric hindrance, atoms cannot overlap, restricting residues to favoured regions representing stable structures like alpha-helices and beta-sheets. A high-quality model features over 90% of residues in these regions; outliers in disallowed areas typically signal errors in the structural coordinates.
RMSE (Root Mean Square Error) is a standard metric that quantifies a model’s prediction accuracy. It calculates the square root of the average squared differences between predicted and actual values. Lower values indicate a better fit. Because it squares errors, it heavily penalizes large outliers. Since it uses the same units as the target data, it provides an intuitive measure of the typical error magnitude.
Applications of Structural Bioinformatics in Drug Discovery
Structural bioinformatics shifts drug discovery from an experimental search to a rational design process. It has numerous uses that can be incorporated into drug discovery.
- Drug identification and validation is the utmost important component when it comes to designing drugs. Identifying deep cavities or pockets on a protein surface where a drug can physically bind helps in binding site analysis.
- Finding hidden sites away from the main active area that can regulate protein function when a molecule binds.
- Protein structure and prediction, by using homology modelling and AI driven modelling
- Virtual Screening computationally screens millions of compounds to find potential hits without expensive lab work.
- Pharmacophore Mapping identifies the essential 3D chemical features (like hydrogen bonds) required for a drug to be active
- Scanning known drug structures to find new therapeutic uses for existing, safe medications
Conclusion
Structural bioinformatics is the computational study of 3D biological macromolecules, driven by the principle that structure dictates function. By integrating data from the Protein Data Bank (PDB) with experimental methods like X-ray Crystallography, NMR, and Cryo-EM, researchers can map atomic coordinates. Advanced modelling techniques including Homology Modelling, Threading, and the AI-powered AlphaFold predict how sequences fold into functional machines. These insights are essential for rational drug discovery, enabling Molecular Docking and Virtual Screening to identify therapeutic hits. Ultimately, this field bridges the gap between raw genetic sequences and the complex molecular mechanisms of life.
References
- Helliwell, J. R. (2021). Structural biology: A golden era. Journal of Applied Crystallography, 54(Pt 1), 1–2. https://doi.org/10.1107/S160057672100045X
- Blundell, T. L., Sibanda, B. L., Montalvão, R. W., Brewerton, S., Chelliah, V., Worth, C. L., … & Mizuguchi, K. (2006). Structural biology and bioinformatics in drug design: opportunities and challenges for target identification and lead discovery. Philosophical Transactions of the Royal Society B: Biological Sciences, 361(1467), 413–423. https://doi.org/10.1098/rstb.2005.1800
- Foundations for the study of structure and function of proteins. (2020). PubMed Central (PMC). https://pmc.ncbi.nlm.nih.gov/articles/PMC7123217/
- Berman, H. M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T. N., Weissig, H., Shindyalov, I. N., & Bourne, P. E. (2000). The Protein Data Bank. Nucleic Acids Research, 28(1), 235–242. https://doi.org/10.1093/nar/28.1.235 RCSB PDB: Homepage
- Adams, P. D., Afonine, P. V., Bunkóczi, G., Chen, V. B., Echols, N., Headd, J. J., Hung, L. W., Jain, S., McCoy, A. J., Moriarty, N. W., Oeffner, R. D., Read, R. J., Richardson, D. C., Richardson, J. S., Terwilliger, T. C., & Zwart, P. H. (2013). How cryo-electron microscopy and X-ray crystallography complement each other. Methods (San Diego, Calif.), 59(3), 329–336. https://doi.org/10.1016/j.ymeth.2013.01.002
- Marion, D. (2013). An introduction to biological NMR spectroscopy. Molecular & Cellular Proteomics, 12(11), 3006–3025. https://doi.org/10.1074/mcp.O113.030239
- Milne, J. L., Borgnia, M. J., Bartesaghi, A., Tran, E. E., Earl, L. A., Schauder, D. M., Lengyel, J., Pierson, J., Patwardhan, A., & Subramaniam, S. (2013). Cryo-electron microscopy: A primer for the non-microscopist. The FEBS Journal, 280(1), 27–45. https://doi.org/10.1111/febs.12078
- https://www.youtube.com/watch?v=Qq8DO-4BnIY
- Hameduh, T., Haddad, Y., Adam, V., & Heger, Z. (2020). Homology modeling in the time of collective and artificial intelligence. Computational and Structural Biotechnology Journal, 18, 3494–3506. https://doi.org/10.1016/j.csbj.2020.11.007
- Peng, J., & Xu, J. (2009). Boosting protein threading accuracy. In S. Batzoglou (Ed.), Research in Computational Molecular Biology. RECOMB 2009. Lecture Notes in Computer Science (Vol. 5541, pp. 31–45). Springer. https://doi.org/10.1007/978-3-642-02008-7_3
- What is AlphaFold? | AlphaFold
- umper, J., Evans, R., Pritzel, A., Green, T., Figurnov, M., Ronneberger, O., Tunyasuvunakool, K., Bates, R., Žídek, A., Potapenko, A., Bridgland, A., Meyer, C., Kohl, S. A. A., Ballard, A. J., Cowie, A., Romera-Paredes, B., Nikolov, S., Jain, R., Adler, J., … Hassabis, D. (2021). Highly accurate protein structure prediction with AlphaFold.
- Agu, P. C., Afiukwa, C. A., Orji, O. U., Ezeh, E. M., Ofoke, I. H., Ogbu, C. O., Ugwuja, E. I., & Aja, P. M. (2023). Molecular docking as a tool for the discovery of molecular targets of nutraceuticals in diseases management. Scientific Reports, 13(1), Article 13398. https://doi.org/10.1038/s41598-023-40160-2
- Pinzi, L., & Rastelli, G. (2019). Molecular docking: Shifting paradigms in drug discovery. International Journal of Molecular Sciences, 20(18), 4331. https://doi.org/10.3390/ijms20184331
- Haddad, Y., Adam, V., & Heger, Z. (2021). Bioinformatics tools in protein analysis: Structure prediction, interaction modelling, and function relationship. Progress in Molecular Biology and Translational Science, 183, 1–45. https://doi.org/10.1016/bs.pmbts.2021.06.014
- Protein Structure Visualization Tools – DrOmics Labs
- https://www.youtube.com/watch?v=LHLA0wNH1dM
- https://www.youtube.com/watch?v=QGBxXHqdC9E
- Skariyachan, S., Prasanna, A., & Manjunath, S. S. (2019). The role of structural bioinformatics in drug discovery and development. In Bioinformatics (pp. 173–201). Academic Press. https://doi.org/10.1016/B978-0-12-814682-8.00008-0
