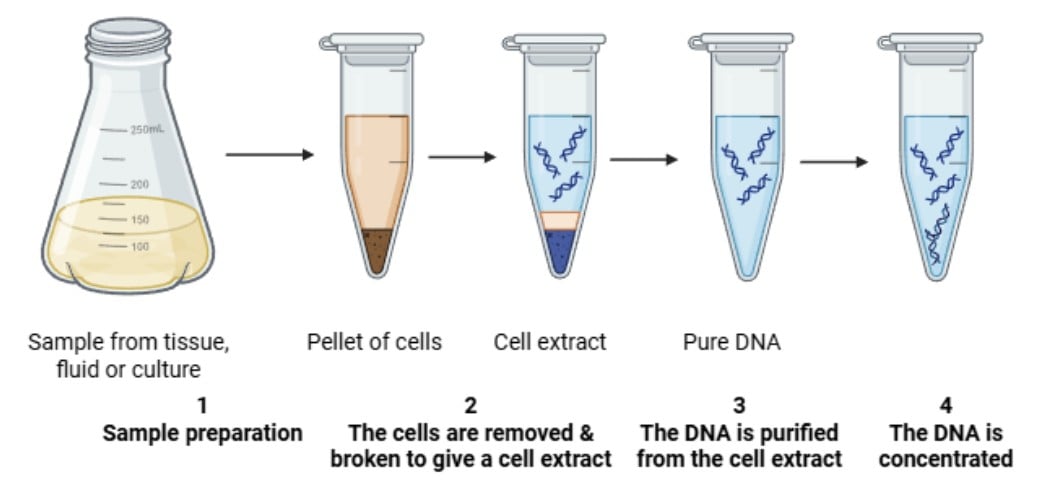
The phenol-chloroform method of DNA extraction is the technique that separates nucleic acids from proteins and other cellular components by exploiting their different solubilities in water and organic solvents.
After breaking open the cells, the mixture is combined with a phenol : chloroform : isoamyl alcohol solution. In this process, proteins and debris move into the organic layer, while DNA and RNA stay in the water-based (aqueous) layer. The DNA is then carefully collected from this layer and precipitated for purification.
This approach provides high-quality, intact DNA that can be used for a wide range of molecular biology applications, including restriction enzyme analysis, cloning, and sequencing (Brown, 2020).
Key Reagents of the Phenol‑Chloroform Method of DNA Extraction
| Reagent | Typical Concentration / Amount | Purpose |
| EDTA (0.6 M) | 1 mL per 14 mL sample (for biological fluids) | Chelates divalent cations (Mg²⁺, Ca²⁺) to inhibit DNases and stabilize DNA. |
| Extraction Buffer (50 mM Tris-Cl pH 8.0, 100 mM NaCl, 10 mM EDTA, 0.5 % SDS, 0.2 mg mL⁻¹ Proteinase K, 1 % 2-mercaptoethanol) | Sufficient to submerge tissue (~500 µL per 50–100 mg tissue) | Disrupts cell membranes and denatures proteins; Tris stabilizes pH, EDTA chelates metal ions, SDS lyses cells, Proteinase K digests proteins, 2-mercaptoethanol reduces disulfide bonds. |
| Phosphate Buffered Saline (PBS) | 500 µL | Resuspends pellet and maintains isotonic conditions before lysis. |
| 10 % SDS | Included in lysis buffer | Solubilizes lipid membranes and denatures proteins. |
| Proteinase K (20 mg mL⁻¹ stock) | Included in lysis buffer (final 0.2 mg mL⁻¹) | Hydrolyzes proteins and facilitates DNA release. |
| RNase A (10 mg mL⁻¹) | 5 µL per sample; incubate 45 min at 37 °C | Degrades RNA, ensuring DNA purity. |
| Phenol : Chloroform : Isoamyl Alcohol (25 : 24 : 1) | Equal volume to lysate (typically 0.5–1 mL) | Denatures and removes proteins; chloroform improves phase separation, isoamyl alcohol minimizes foaming. |
| Chloroform (100 %) | Equal volume to aqueous phase | Removes residual phenol from aqueous DNA phase. |
| 3 M Sodium Acetate (pH 5.2) | 1/10 volume of aqueous phase | Provides Na⁺ ions to neutralize DNA’s phosphate backbone, promoting aggregation during ethanol precipitation. |
| 100 % Ethanol (chilled) | 2 × volume of aqueous phase | Precipitates DNA at low temperature for recovery. |
| 70 % Ethanol (chilled) | Two washes of DNA pellet | Removes residual salts and organic contaminants. |
| TE Buffer (10 mM Tris-Cl pH 8.0, 1 mM EDTA) | 50 µL for resuspension | Solubilizes and preserves DNA for storage; Tris stabilizes pH and EDTA inhibits nucleases. |
Principle of the Phenol-Chloroform Method of DNA Extraction
Phenol, a polar organic solvent, disrupts hydrogen bonds and hydrophobic interactions in proteins, causing them to partition into the organic layer. Chloroform helps make the separation between layers sharper and clearer, while isoamyl alcohol prevents excessive foaming during mixing. DNA and RNA stay dissolved in the water-based (aqueous) layer because their negatively charged phosphate backbone is stabilized by the surrounding water.
Finally, adding ethanol along with sodium ions causes the DNA to clump together into a visible pellet, which can then be collected, leaving most contaminants behind (Brown, 2020).
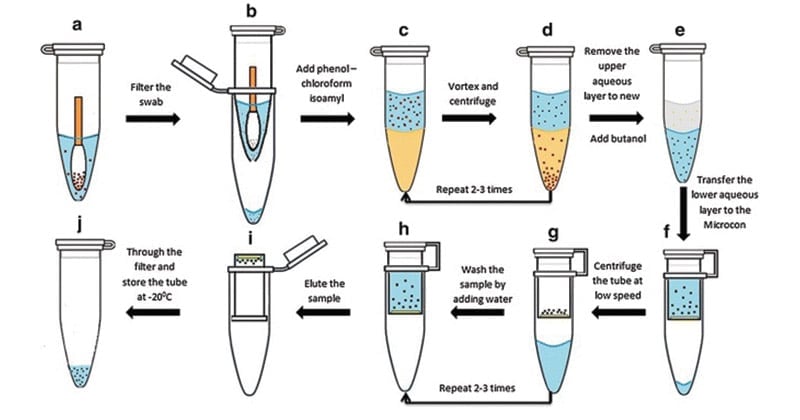
Protocol of the Phenol‑Chloroform Method of DNA Extraction
A. Sample Preparation
- For biological fluids and culture media, transfer 7 mL of the sample to a tube pre‑filled with 0.51 mL 0.6 M EDTA. Centrifuge at 15,000 g for 15 min; discard supernatant. Repeat if cell pellet is not visible.
- For tissue samples, add small pieces to the extraction buffer (50 mM Tris-Cl pH 8.0, 100 mM NaCl, 10 mM EDTA, 0.5 % SDS, 0.2 mg mL⁻¹ Proteinase K, 1 % 2-mercaptoethanol). Homogenize it on ice in a mortar and pestle until no large pieces remain, avoiding foaming (Kathmandu University, 2025).
B. Cell Lysis
- Resuspend the pellet in 500 µL PBS if the sample is to be temporarily stored. Add 700 µL of the prepared lysis buffer (10 % SDS + Proteinase K). Vortex 1 min.
- Add 5 µL RNase A (10 mg mL⁻¹) and incubate at 37 °C for 45 min, gently inverting every 10 min.
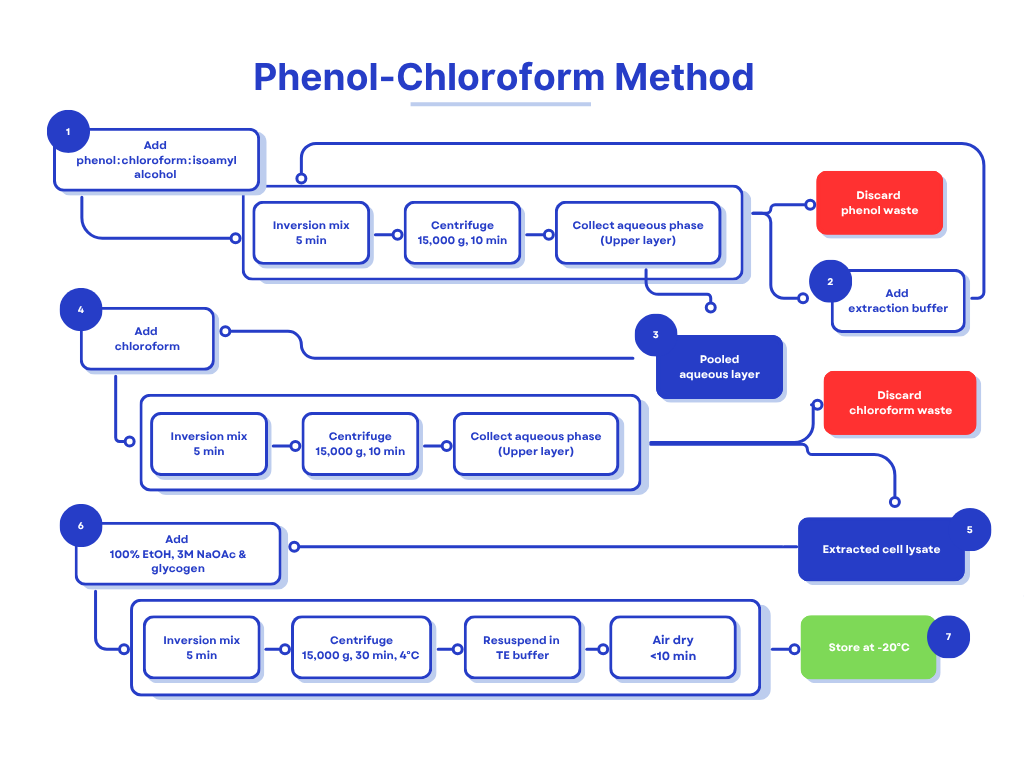
C. Phenol‑Chloroform Extraction
- Add an equal volume of phenol : chloroform : isoamyl alcohol (25 : 24 : 1). Mix gently by inversion for 30 s.
- Centrifuge at 16,000–18,000 g for 5 min (room temperature). Observe three layers:
- Bottom (organic) – phenol/chloroform with denatured proteins.
- Middle (interphase) – white protein debris.
- Top (aqueous) – DNA-containing layer.
- Carefully pipette the aqueous layer into a fresh tube; repeat the extraction 2–3 times until the interphase is clear.
D. Phenol Removal & DNA Precipitation
- Add an equal volume of chloroform to the final aqueous layer to strip residual phenol; mix and centrifuge as above.
- Transfer the aqueous phase, add 1/10 volume of 3 M sodium acetate, and 2 × volume of chilled 100 % ethanol. Mix by inversion and incubate on ice for 20 min.
E. DNA Recovery
- Centrifuge at 16,000 g, 4 °C for 15 min. Discard supernatant, wash pellet twice with chilled 70 % ethanol.
- Air‑dry briefly (≤10 min) and resuspend in 50 µL TE buffer. Store at –20 °C.
F. Optional Clean‑up
- If high protein carryover is suspected, perform an additional phenol‑chloroform step before ethanol precipitation.
Modifications of the Phenol‑Chloroform Method of DNA Extraction
- Carrier-aided precipitation: Adding glycogen or linear polyacrylamide improves DNA recovery from low-concentration samples (Liu et al., 2022).
- Isopropanol substitution: Using 0.7× volume of 2-propanol accelerates DNA precipitation in dilute lysates (Liu et al., 2022).
- Magnetic bead capture: Silica-coated magnetic beads allow DNA to bind and be washed quickly after lysis, reducing centrifugation steps (Green& Sambrook, 2017 et al., 2022).
- CTAB pre-treatment for plant tissues: For carbohydrate-rich samples, CTAB forms an insoluble DNA–CTAB complex, which is later dissolved in 1 M NaCl before phenol extraction (Van der Merwe, 2019).
Troubleshooting of the Phenol‑Chloroform Method of DNA Extraction
| Problem | Likely Cause | Solution |
| Low DNA yield | Incomplete cell lysis or insufficient precipitation | Increase SDS (to 1 % w/v) or add a second freeze‑thaw cycle (protocol.io).• Extend Proteinase K incubation (20 mg mL⁻¹, 1 h at 55 °C).Use ethanol: sample ≥ 2.5 : 1 or add 0.7 vol isopropanol (Bite‑size Bio). |
| Protein contamination | Inadequate number of phenol extractions | Perform 3–4 phenol‑chloroform extractions; pre‑treat lysate with Proteinase K (20 mg mL⁻¹) and, if needed, a second Pronase step (CSH Protocols). |
| Phenol carry‑over (orange tint) | Incomplete chloroform wash | Add a fresh chloroform‑isoamyl step; use Phase‑Lock Gel tubes to prevent interphase bleed‑through (Bite‑size Bio). |
| Sheared DNA | Vigorous vortexing or multiple extractions | Mix gently by inversion or rotate at ≤ 20 rpm (CSH Protocols). Limit bead‑beating to ≤ 2 min for soil samples. |
| RNA contamination | Insufficient RNase A treatment | Increase RNase A to 10 µg mL⁻¹, incubate 30 min at 37 °C, and verify activity with a control RNA. Keep RNase‑free tips and surfaces (Bite‑size Bio). |
| Poor phase separation (cloudy interface) | Low temperature of chloroform, presence of salts, or expired phenol | Use chilled chloroform (4 °C) and phenol equilibrated to pH 7.8–8.0; add extra 0.1 vol 3 M sodium acetate before extraction (protocol.io). |
| Small or invisible pellet | Over‑drying, low DNA concentration, or absence of carrier | Add 1 µg glycogen or 10 µg linear polyacrylamide before precipitation; avoid > 10 min air‑drying; use cold ethanol (‑20 °C). |
| Humic‑acid or polysaccharide contamination (A₂₆₀/A₂₃₀ < 1.8) | Soil/plant matrix rich in organic inhibitors | Include a CTAB pre‑extraction step for plant tissue; after phenol extraction, perform a second chloroform wash and optional PVPP column clean‑up (protocol.io). |
| DNA degradation (smearing on gel) | Prolonged exposure to phenol at room temperature | Perform extraction at 4 °C (automated robot protocol) and keep samples on ice during phase separation. |
| Ethanol residue after wash | Inadequate removal of ethanol during aspiration | Use Phase‑Lock Gel tubes; after the final 70 % ethanol wash, spin 5 min at 14,000 g, then invert tube on a lint‑free tissue to blot residual ethanol before drying. |
Quality Assessment of the Phenol‑Chloroform Isolated DNA
- Spectrophotometry: Assess DNA purity by measuring absorbance ratios. An A₂₆₀/A₂₈₀ ratio around 1.8 indicates minimal protein contamination, while an A₂₆₀/A₂₃₀ ratio ≥2.0 suggests low polysaccharide or organic solvent contamination (Brown, 2020; Green & Sambrook, 2017).
- Agarose gel electrophoresis: Run DNA on a 0.8% agarose gel. High-molecular-weight DNA appears as a bright band near the top of the gel with minimal smearing, indicating intact DNA (Brown, 2020; Liu et al., 2022).
- Fluorometric quantification: Use the Qubit dsDNA HS assay for accurate measurement of DNA concentration, especially useful for low-yield or precious samples (Liu et al., 2022).
Safety Tips and Precautions of the Phenol‑Chloroform Method of DNA Extraction
- Personal Protective Equipment (PPE): Always wear a lab coat, nitrile gloves (double-gloving recommended), and chemical splash goggles or a face shield. Phenol and chloroform can penetrate latex and cause burns or irritation (Green & Sambrook, 2017; Reina, 2025).
- Ventilation: Perform all phenol and chloroform steps in a certified fume hood. Phenol is corrosive, and chloroform is hepatotoxic and potentially carcinogenic (Green & Sambrook, 2017; Liu et al., 2022).
- Automation advantage: Using robotic or enclosed extraction systems reduces direct contact with phenol vapors and minimizes manual handling during phase separation (Liu et al., 2022).
- Temperature safety: Keep phenol chilled at 4 °C in secondary containment to preserve DNA integrity and limit volatilization. Avoid freezing, which can cause crystallization and container rupture (Green & Sambrook, 2017).
- Waste disposal: Collect phenol–chloroform waste in properly labeled halogenated solvent containers. Neutralize residual phenol with NaOH before disposal according to institutional biosafety guidelines (Green & Sambrook, 2017; Van der Merwe, 2019).
- Spill management: Absorb spills using inert materials such as vermiculite or pads, then rinse affected areas with plenty of water for at least 15 minutes. Avoid skin contact and report all exposures immediately (Green & Sambrook, 2017).
Storage and Long‑Term Stability of Phenol‑Chloroform Isolated DNA
- Short-term storage: Keep aqueous DNA at 4 °C for up to one week to minimize nuclease activity and maintain integrity for immediate downstream applications (Green & Sambrook, 2017).
- Long-term storage: Aliquot DNA in TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0) and store at –20 °C or –80 °C. Avoid repeated freeze-thaw cycles, which can cause shearing and depurination (Liu et al., 2022).
- Phenol-containing extracts: If DNA remains in phenol, store it at 4 °C for no more than 48 hours before ethanol precipitation, as phenol can oxidize over time and inhibit polymerases (Van der Merwe, 2019; Green & Sambrook, 2017).
- Ethanol preservation: For field collection or long-term archival purposes, DNA pellets can be stored in 70–100 % ethanol at –20 °C, which helps prevent hydrolytic damage and microbial growth (Liu et al., 2022).
Applications of the Phenol‑Chloroform Method of DNA Extraction
- Genomic DNA isolation: Produces high-quality genomic DNA suitable for restriction mapping, library construction, and whole-genome sequencing (Green & Sambrook, 2017; Liu et al., 2022).
- RNA-free DNA for epigenetic studies: Allows complete removal of RNA using RNase, which is essential for methylation analyses and bisulfite-conversion assays that require RNA-free DNA (Green & Sambrook, 2017).
- High-molecular-weight DNA recovery: Efficiently recovers intact, megabase-sized DNA from degraded or challenging samples such as forensic tissue, plant residues, or ancient specimens (Liu et al., 2022).
- Viral and environmental nucleic acids: Provides highly purified viral or microbial DNA from soil and water samples, ensuring compatibility with infectivity assays and metagenomic sequencing (Van der Merwe, 2019).
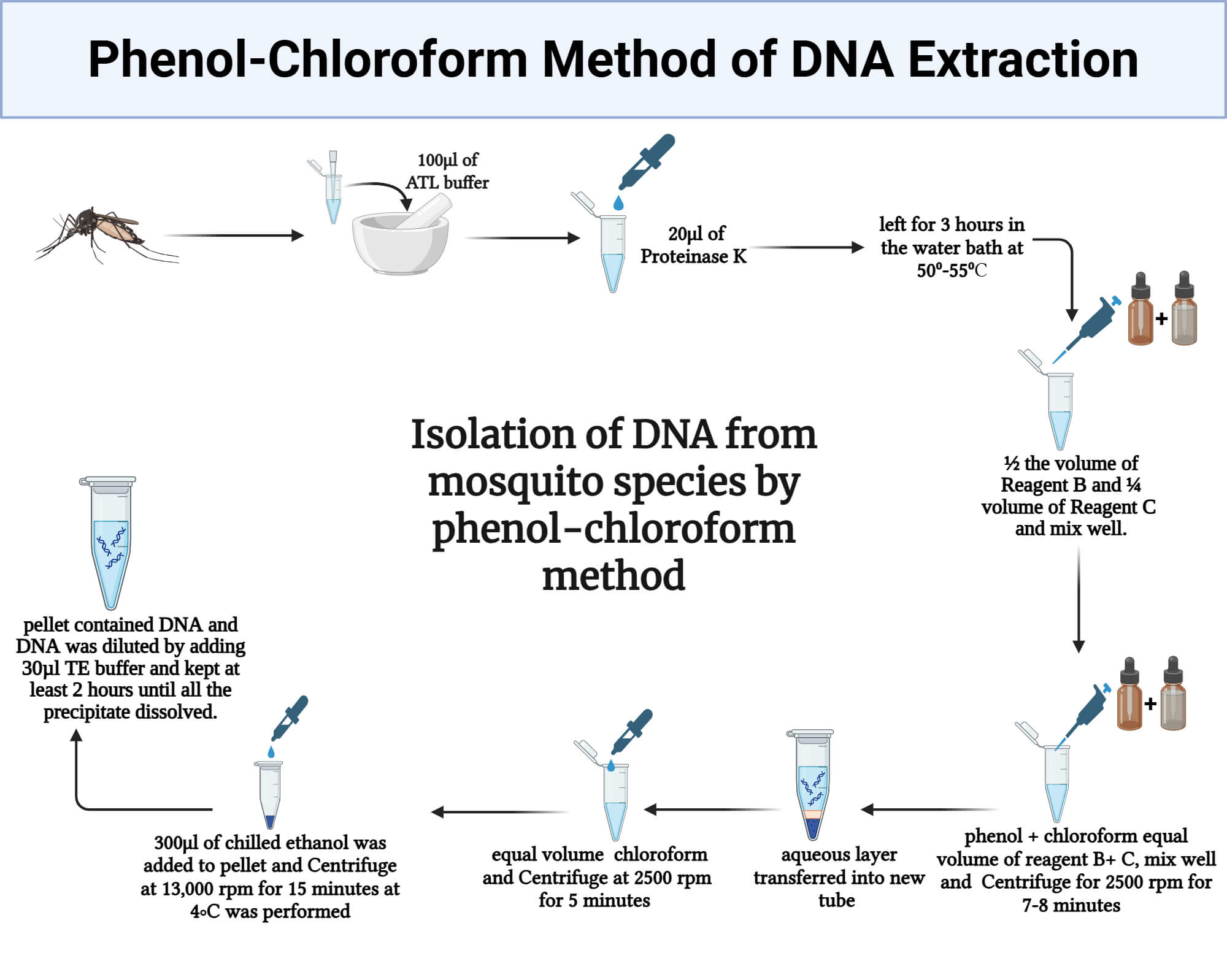
Advantages of the Phenol‑Chloroform Method of DNA Extraction
- Cost-effective: Relies on inexpensive, readily available reagents, providing a budget-friendly alternative to commercial silica- or magnetic-bead-based kits (Green & Sambrook, 2017).
- High yield and integrity: Generates large amounts of DNA with minimal shearing when handled gently, making it suitable for long-read sequencing and high-molecular-weight applications (Liu et al., 2022).
- Scalable: Works efficiently for large-volume extractions, even hundreds of milliliters, without being limited by column binding capacity (Van der Merwe, 2019).
Limitations of the Phenol‑Chloroform Method of DNA Extraction
- Chemical hazards: Phenol and chloroform are toxic and corrosive, so all work must be performed in a fume hood with strict adherence to safety and disposal protocols (Green & Sambrook, 2017).
- Labor-intensive: The method involves multiple mixing, centrifugation, and phase-separation steps, which increase manual handling and overall processing time (Liu et al., 2022).
- Phenol carry-over: Any residual phenol can interfere with PCR, restriction digestion, or other enzymatic reactions if not completely removed (Green & Sambrook, 2017).
- Limited throughput: This approach is not ideal for high-volume laboratories; automated or column-based systems are preferred for greater efficiency and safety (Liu et al., 2022).
Conclusion
The phenol-chloroform extraction method remains a reliable, flexible, and cost-effective approach for isolating high-quality DNA from a wide range of biological sources. When reagent ratios are optimized, phase separation is performed carefully, and washing steps are thorough, the method consistently yields DNA suitable for demanding molecular biology applications.
Although phenol and chloroform require strict safety measures, modern adaptations- such as magnetic-bead capture and CTAB pre-treatment- have improved their efficiency and broadened their use for complex or inhibitor-rich samples.
Despite the rise of commercial kits, this classic technique continues to hold value for laboratories seeking high-purity, high-integrity DNA at a low cost (Green & Sambrook, 2017; Liu et al., 2022; Van der Merwe, 2019).
References
- Liu, A. W., Villar-Briones, A., Luscombe, N. M., & Plessy, C. (2022). Automated phenol-chloroform extraction of high molecular weight genomic DNA for use in long-read single-molecule sequencing. bioRxiv. https://doi.org/10.1101/2022.01.26.477939
- Reina, O. (2025, June 5). Top 10 tips for phenol-chloroform extractions and ethanol precipitations. Bitesize Bio. https://bitesizebio.com
- Van der Merwe, S. (2019). DNA extraction from water/soil: 50-50-50 buffer-chloroform/phenol method. protocols.io. https://doi.org/10.17504/protocols.io.8yphxvn
- Miller, D. N., Bryant, J. E., Madsen, E. L., & Ghiorse, W. C. (1999). Evaluation and optimization of DNA extraction and purification procedures for soil and sediment samples. Applied and Environmental Microbiology, 65(11), 4715–4724.
- Green, M. R., & Sambrook, J. (2017). Isolation of high-molecular-weight DNA using organic solvents. Cold Spring Harbor Protocols, 2017(4), pdb.top093450. https://doi.org/10.1101/pdb.top093450
- Brown, T. A. (2020). Gene cloning and DNA analysis: An introduction (8th ed.). Wiley-Blackwell.
- Kathmandu University. (2025). Phenol–chloroform extraction manual for human urine samples. Unpublished laboratory protocol.
