The Polymerase Chain Reaction was invented by Kary Mullis in 1983. It is a thermal process used to amplify a specific gene within a DNA sequence. The reaction works on the principle that specific genes can be amplified in vitro under optimal conditions. These conditions are provided by a thermocycler, which allows for consecutive changes in temperature. Replication occurs in three steps: Denaturation, Annealing, and Extension. This process uses Taq polymerase, dNTPs, primers, buffers, and more. Traditional PCR, though useful, has drawbacks. Primers may bind non-specifically to the template, resulting in useless amplicons.
To address this, a variant called Nested PCR was developed. Nested PCR is a modified form of polymerase chain reaction in which two sets of primers, external and internal, are used sequentially to amplify the same target nucleic acid. The first amplification is carried out using external primers, followed by a second amplification using internal primers, which increases the sensitivity and specificity of detection.

Objectives of Nested PCR
The objectives of Nested PCR are:
- To improve the sensitivity of the reaction
- To reduce the chances of non-specific amplification of the target DNA, ie, to increase the specificity of the reaction.
- To increase the number of cycles, thereby improving the specific amplification of the target DNA
- To prevent the production of non-specific products, thereby avoiding resource waste.
Requirements of Nested PCR
Nested PCR utilizes the same materials as a traditional PCR with an extra set of primers. The required materials include:
- Template DNA sample
- Primers: They are short stretches of oligonucleotides required for the replication process to begin. Two sets are required: first-round PCR primer (External Primer) and second-round PCR primer (Inner Primer).
- Taq Polymerase
- dNTP mixture (dATP, dCTP, dGTP, dTTP)
- 10 x PCR buffer
- MgCl2 Solution
- Thermocycler
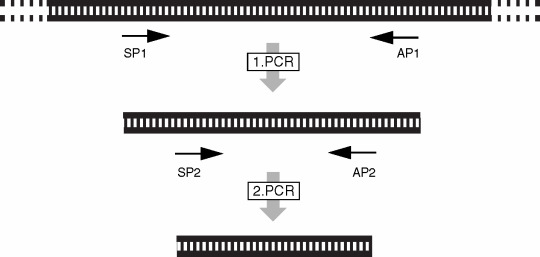
Principle of Nested PCR
Nested PCR increases the specificity and the sensitivity of the amplification reaction. This is done with the help of two sets of primers. Here, the first pair of primers, ie, the external primer, amplifies the target DNA first. The amplification takes place for 15-30 cycles, after which the second round of amplification is performed by the second set of primers, i.e., internal primers or nested primers. Before the second round of multiplication, part of the product from the first round is diluted 100-1000 times and then utilized as a template. The resulting product of the second round of amplification is shorter than that of the first round.
The use of two sets of primers is to increase the specificity and the sensitivity of the reaction by reducing the chances of production of non-specific amplicons, which arise due to non-specific binding between the primer and the target DNA. If the external primers result in non-specific binding, this product, when used as a template for the second round, it is highly unlikely that the tested primers will recognize the same non-specific region for amplification. This results in specific amplification of the target DNA.
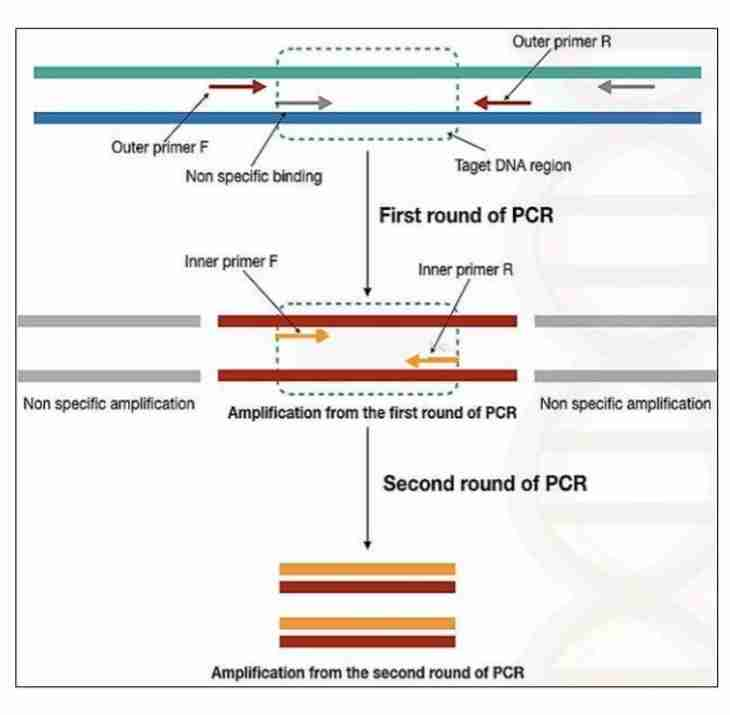
This method increases the specificity by two mechanisms:
- The product of the first round of amplification is used as a template for the second round, thereby reducing the chances of non-specific amplicon production.
- The higher number of cycles in the first round improves the sensitivity, since the number of target DNA is doubled here.
The primers used for this method are:
- External Primers: These are the first set of primers, the outer primers, which anneal to the outside or flanking the target sequence. They are present upstream of the inner set of primers. These primers are 18-22 nucleotides long.
- Inner Primers: Also known as “Nested primers”, they bind inside the outer primer regions. It is this binding that increases the reaction specificity. The primers are up to 20 nucleotides long.
Steps of Nested PCR
The steps involved in nested PCR are:
- Preparation of the reaction mixture
- Amplification
- Post-amplification analysis of the product.
1. Preparation of the reaction mixture
The reaction mixture is prepared to a final volume of 25μL with the following components:
- Template DNA: 1-2μL
- External primer (per pair): 0.5μL (final concentration 0.2μM)
- dNTP mixture: 0.5μL (final concentration of 200μM per dNTP)
- 10×PCR buffer: 2.5μL
- MgCl2: 1.5μL (final concentration is 1.5-2.0mM)
- Taq DNA polymerase: 0.25μL (1.25U)
- Sterile ultrapure water: Replenished to 25μL
The prepared reaction mixture was added to the thermocycler for amplification.
2. Amplification
The amplification process is divided into three steps, which include:
Denaturation: The first step in PCR, here the DNA template is separated into two strands to be utilised as templates for the amplicon production. Denaturation of the DNA strand is done at 94°C and continued for 2 minutes.
Annealing: After denaturation, the temperature is lowered to 45- 55°C for the hybridization of the external primer to the target DNA. The time taken for this is 30 seconds.
Extension: The final step in PCR, where the temperature is increased to 72°C, is the optimal temperature for the Taq polymerase to start functioning. The polymerase adds the added NTPs to form a double-stranded amplicon. These steps proceed for 1 minute, with the final elongation requiring 5 minutes.
After the first round of amplification, which has around 30-35 cycles, 1-2μL of the product is taken and diluted (eg: 1:10). The diluted product is then used as a template for the second round of amplification with the nested primers. The concentration of the materials is the same, with the inner primers having a concentration of 0.5μL/per pair (final concentration 0.2μM).
3. Analysis of post-amplification products
Upon amplification, the products are analyzed using agarose gel electrophoresis.
Types of Nested PCR
Nested PCR has 4 types. These include:
1. Semi-nested PCR
This method has the same principle as that of the nested principle, except that instead of using four primers (two sets), here only three primers are used. For the first round of the amplification, two primers are used, while for the second round, only one primer is used. For applications requiring nested PCR, this is a good alternative in case of not being able to design specific primers for the 5’ or 3’ end.
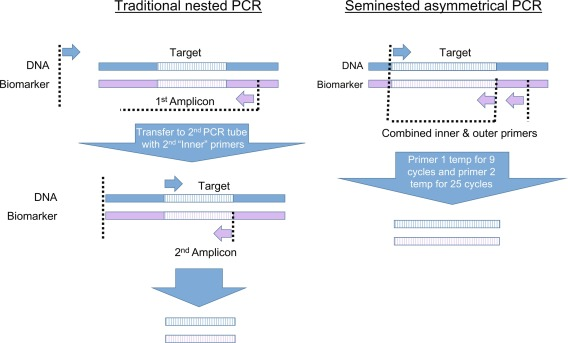
2. Reverse transcriptional nested PCR
Also, now RT-nested PCR combines the principles of both reverse transcription and nested PCR. This method is used for the amplification of cDNA, which has been obtained from the target RNA. This method is used to detect whether a certain RN has been expressed or to compare its relative expression level. This is a more specific and highly reliable method and can be used for samples of RNA with low copy number.
Eg: HCV genes in Hepatitis C virus-infected people.
3. One-tube nested PCR
This method is based on the traditional PCR method, where both rounds of the amplification are done in a single tube. The first set of primers is 25bp long, and the annealing temperature is relatively higher, at 68℃, while for the second round, the primers are 17bp long, at a lower temperature of 46℃. The first round is run for 20-30 cycles. Since both the PCR reactions are performed in a single tube, the chances of cross-contamination are reduced.
4. Consensus nested PCR
This method is used by designing degenerate primers based on the conserved sequences within the same genus. Degenerate primers are usually found in the first round of amplification, with fewer ones found in the second round, with the amplification length being 200~300. To differentiate between the different subtypes of the microorganisms, priors have been designed. This is done because there may be various types within the genus, and the type of organism within the tube is unknown and has to be determined. By amplifying using nested PCR to obtain the target sequence, it can help in finding the unknown organism. This makes it a simple and simple detection method.
To design the primer for consensus PCR, all the information related to DNA sequences should be obtained, and sequence comparison should be carried out via software to find the conserved sequences. Within these sequences, there may be nucleotide diversity that should be taken into consideration and designed as reduced bases. Keeping the diversity in mind, the length of the products of the second round of PCR amplification was controlled to about 200~300 bp. The appropriate annealing temperature should be used due to the presence of many degenerate bases.
Examples of Nested PCR Kits
HiPer® Nested PCR Teaching Kit (HiMedia)
- Used for amplifying the DNA sequence in two successive runs.
- Contains two sets of primers required to run the nested PCR protocol.
- Modified to be highly specific and sensitive by preventing non-specific binding
TRUPCR® One Step nested Kit (MTBC and NTM) (TRUPCR Europe)
- It is a real-time PCR kit using a single tube for the PCR reactions.
- It is an in-vitro diagnostics assay utilized for the detection of low-load Mycobacterium DNA in human clinical samples (Pulmonary and extrapulmonary), with the help of sequential PCR reactions.
- Each of the tests is a multiplexed reaction with three independent reactions running in parallel.
- Mycobacterium tuberculosis is detected by targeting the conserved sequence (two preferred gene sequences); additionally, one target is for the genus Mycobacterium to differentiate between MTB and NTM (MOTT). It also contains an internal control (IC) reaction.
- Provides accurate and simultaneous detection due to targeting three different genes.
Nested PCR Kit (SLS Research)
- Enhances the specificity of the PCR reaction
- Utilises two sets of primers; no additional components required.
- The components of the kit should be stored in -20℃.
Pro TechEx-Nested PCR Teaching Kit (Progene)
- Follows the principle of the nested PCR and contains the components required for performing the experiment in laboratory settings.
- Required detailed knowledge of the sequence of the target.
- With a single kit, 5 experiments can be performed. There are kits with a higher rate of experiment (25x).
- Contains Agarose, 20x TAE, Ethidium Bromide, Gel loading dye, Template DNA, Primer (Forward and Reverse), Taq Polymerase,10mM dNTP mix, 10x Taq polymerase Buffer, Distilled water, PCR tubes, and the protocol sheet. The component is to be stored at 20℃.
Applications of Nested PCR
High sensitivity modified nested PCR or detection of Leishmaniasis: The techniques available in the market for Leishmaniasis are inadequate and not sensitive. With a modified nested PCR, detection of the disease in blood and tissue samples can be done. By designing primers specific to the Leishmania genetic material, the parasite can be detected even in low-concentration samples. This allows for timely detection and intervention, reducing the progression of the disease.
Detection of Mycobacterium tuberculosis: Similar to Leishmaniasis, the techniques currently available for the detection of tuberculosis are not specific. Thus, a modified nested PCR method, referred to as a one-time nested PCR, has been developed, which utilizes real-time PCR and nested PCR principles to detect the pathogen from the samples. It is comparatively a simpler and more accurate method.
Detection of respiratory viruses: Since respiratory disease can be caused by a diverse group of viruses and bacteria, fast detection of them is necessary. A modified nested method utilizes the principle of the multiplex PCR method, which can be used, resulting in a higher specificity and sensitivity (100-1000 times). This allows for improving the diagnostic rate of the infection and early treatment.
Advantages of Nested PCR
The advantages of this method include:
- Improves the sensitivity by increasing the amplification factor and overcoming the limitation of the single amplification plateau effect.
- Non-specific binding of the primer and target sequence is greatly reduced by performing amplification in two rounds.
- The accuracy and the feasibility of the methods are increased since the amplification in the second round is done using the product of the first round of amplification.
- Can be used for the detection of samples with a lower load of the target microorganism.
- Provides good results for samples with high G+C content or longer templates.
Limitations of Nested PCR
Some of the limitations of the method are as follows:
- Since the template for the second round of amplification is the product of the first round, during the opening of the tube and transferring, there are chances of cross-contamination.
- Detailed knowledge of the target sequence is required to design specific primers for the template.
- The primer content should be maintained so that there is no residual primer during the amplification process.
- As it requires a separate reaction setup, it could be a time-consuming and tedious process, and could be expensive too.
Conclusion
Nested PCR is a good alternative to the traditional PCR methods as it overcomes the possibility of production of non-specific products due to the non-specific binding between the primer and the target DNA. It is a modification of the traditional PCR method, where sets of primers are used to increase the specificity and the sensitivity of the amplification. The external primers do the first round of amplification, the product of which is used for the second round with a pair of internal primers. Despite the chances of cross-contamination, this is a highly reliable and accurate method for the detection of target compounds in samples with lower load, or High G + C content, and longer templates.
References
- https://www.bocsci.com/resources/nested-pcr-principle-protocol-and-applications.html?srsltid=AfmBOorzjgN_N3vW8D1kinKQhficfxk8egWIa7e-TzZSkMnz2jpC_p_L
- Mulhardt, C. (2010). Molecular biology and genomics. Elsevier.
- Shen, C. H. (2019). Diagnostic molecular biology. Academic Press.
- Singh, Y., Anand, K. J., Singh, M., Ambawat, S., & Tiwari, G. (2023). Recent developments in PCR technology. Biological Sciences, 77, 79.
- Green, M. R., & Sambrook, J. (2019). Nested polymerase chain reaction (PCR). Cold Spring Harbor Protocols, 2019(2), pdb-prot095182.
- https://geneticeducation.co.in/what-is-nested-pcr/#Applications_of_Nested_PCR
- https://www.indiamart.com/proddetail/pro-techex-nested-pcr-teaching-kit-9553337397.html?srsltid=AfmBOorHnXvBi17x_tZvldxVGVu0UMAel6mmySPmrZkCxHcQHi6WS6Kl
- https://www.himedialabs.com/in/htbm043-hiper-nested-pcr-teaching-kit.html
- https://slsresearch.com/shop/nested-pcr-kit/
