In conventional PCR, amplification occurs at both ends of the target DNA. Some applications require specific amplification of a DNA fragment containing only one known end. These include DNA fingerprinting, determination of cysteine methylation patterns, primary DNA nucleotide sequences, DNA lesion formation and repair, and in vivo protein-DNA footprints. In genomic sequencing and in vivo footprinting, it is useful to amplify a family of fragments with one end in common. Ligation-mediated Polymerase Chain Reaction (LMPCR) was developed for this purpose.
Ligation-mediated PCR (LMPCR) is a PCR-based technique in which short DNA linkers (adaptors) are ligated to fragmented target DNA, enabling selective amplification using one primer specific to the linker and another specific to a gene of interest.

A key advantage is that the method preserves the original fragment lengths with base-pair resolution, which makes it particularly useful in genomic sequencing and in vivo footprinting studies.
Objectives of Ligation-Mediated PCR
The objectives of ligation-mediated PCR include
- To perform genomic typing and fingerprinting without prior knowledge of the sequence
- To detect an unlimited number of restriction fragments virtually and analyze them in genomes.
- To selectively amplify a subset of DNA fragments generated by restriction enzymes.
- To perform PCR for processes, such as DNA sequence ladders, to retain single-base resolution.
Requirements for Ligation-Mediated PCR
The materials required for Ligation-mediated PCR follow the basic pattern, which includes:
- Primer: Two primers are to be prepared; one for the linker that is used and the other for the gene of interest.
- Deoxynucleotides (dNTPs)
- Linkers: These are small fragments of DNA that can be ligated onto the gene of interest for the amplification to be made more specific.
- Buffers
- Ethanol
- DNA ligase
- Taq DNA Polymerase
- Magnesium Chloride
- Purified template DNA
- Thermocycler
Principle of Ligation-Mediated PCR
The Traditional Polymerase Chain Reaction is used to exponentially amplify the target DNA. However, this method cannot specifically amplify the required gene without prior sequence knowledge of the template DNA. In genomic sequencing and in vivo footprinting, there is a need to amplify DNA containing only one end of a known sequence. Ligation-mediated Polymerase Chain Reaction (LMPCR) is a single-sided PCR method that addresses these challenges.
In this method, a short fragment of known DNA sequence, called a “linker” or “adaptor,” is ligated onto the template DNA. This allows specific amplification of any DNA fragment. Only one primer must hybridize to the template; the second primer binds to the linker. The added linker gives specificity by providing a defined and discrete sequence. This helps retain single-base resolution in the final product.
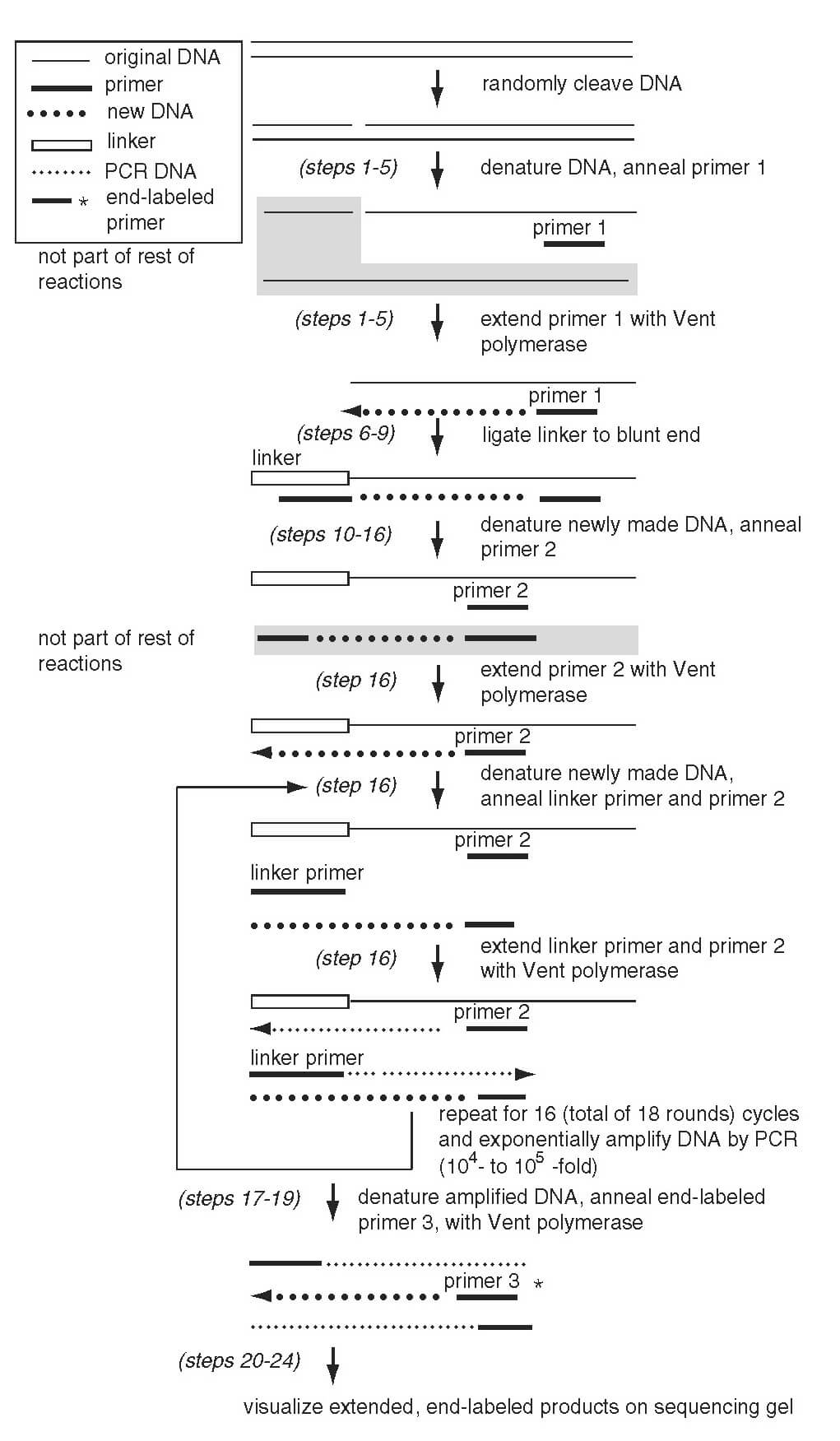
To perform Ligation-mediated PCR, the genomic DNA to be used as the template is first cleaved so that the 5’-phosphate is retained at the cleavage site. Then the cleaved DNA is denatured using the appropriate restriction enzymes, and a gene-specific primer is annealed to the region of interest. Such an annealing step is required so that synthesis can be done at the cleavage site to create a blunt end.
The DNA ligase is used to ligate the 3’ end of the unidirectional (staggered) linker to the 5’ end of the genomic DNA (the blunt end). Since the 5’ end of the linker lacks the phosphate group, the linker cannot be ligated onto the extension product of the gene-specific primer. In the next cycle, the DNA is denatured, and a second gene-specific primer is annealed, and extension of the strand takes place through the ligated linker region. The generated product contains two ends, one with a specific linker end and the other with the genomic sequence. The extended product is specific in the way that only the molecules with both the specified ends can be amplified in further cycles, whereas the molecules with only one end can only be linearly amplified.
For the post-PCR analysis, a third gene-specific primer is used, which is used to label the DNA indirectly, and it overlaps with the second primer. Such a labeled DNA can be visualized on a sequencing gel.
Steps in Ligation-Mediated PCR
The steps in Ligation-mediated PCR can be divided into 5:
- First strand synthesis
- Ligation
- Amplification
- End labelling
- Analysis
1. First strand synthesis
- 5 µl (2 µg) of the cleaved genomic DNA was added to a microcentrifuge tube and chilled for several minutes in an ice water bath. The sample used is made sure to be clean and free of contaminants.
- The reaction mixture is prepared by mixing gene-specific primer (Primer 1) and chilled for several minutes in an ice water bath. The 25 µl of this primer is added to he previously chilled DNA sample and gently mixed with a pipette. This mixture is chilled once again.
- The reaction mixture is added to the thermocycler for the synthesis process. The DNA is first denatured at 95°C for 5 minutes, the primer is then annealed at 60°C for 30 minutes, and the extension temperature is at 76°C for 10 minutes. The extension results in the creation of a blunt end for the linker ligation to take place.
- A ligase mixture and ligase dilution were chilled in an ice water bath, whereas the unidirectional linker was allowed to thaw.
- Upon completion of the third step, the sample is transferred to an ice water bath.
2. Ligation
- To the ice-cold ligase mix, the unidirectional linker and the DNA ligase were added, mixed well, and kept in an ice waterbath
- The sample was prepared by making it precipitate by placing it in an ice-water bath for a few minutes, whereas the microcentrifuge tube was chilled at 4°C so that the transfer of the sample to the tube would be smooth and the sample would not get warm in the process.
- Salt mix was prepared, and it was added to the sample along with 100% ethanol. It was mixed well through inversion and allowed to chill for more than 2 hours at -20°C. The ethanol keeps the sample stable for weeks when stored at -20°C
3. Amplification
- Centrifuge the sample to take out the pellet. The collected pellet was washed with water to dilute. The dissolved pellet is then kept in an ice-water bath.
- The reaction mixture with the linker primer and the gene-specific primer, and keep it in an ice water bath.
- Add 30 µl of the chilled mix to the sample and place it in the ice-water bath after mixing it with a pipettor.
- DNA polymerase was added to the other sample and mixed with a pipette, and the mix was chilled.
- The final mixture was covered with 90 µl mineral oil and microcentrifuged at 4°C.
- The mix was then used for amplification. 18 cycles of PCR are to be performed. The first denaturation is done for 3-4 minutes at 95°C, while the rest is for 1 minute.
- Annealing of primers is done at a temperature between 0 °C to 2°C for 1 minute. The extension is allowed to take place for 3 minutes at 76°C, and for the subsequent cycles, an extra 5 seconds is added, with the final extension taking place for 10 minutes.
- The sample is transferred to an ice water bath and microcentrifuged at 4°C to remove any condensation.
4. End labeling
- The end labeling mixture with the third primer was prepared and chilled, which was then added to the sample.
- Two rounds of PCR were carried out for the labeling processing; denaturation for3-4 minutes at 95°C, with the second round for 2minutes, Annealing of primer 3 at 0° to 2°C for 2 minutes, and finally extension at 76°C for 10 minutes.
- For the analysis of the product, a loading buffer was prepared and added to the sample, which was microcentrifuged at 4°C to obtain the pellet.
5. Analysis
- The samples were denatured at 85° to 90°C fr minutes and the contents loaded onto a sequencing gel to obtain the DNA ladders.
- After the run, autoradiograph the gel that has been fixed and dried for 6-24 hours.
Types of Ligation-Mediated PCR
Different types of molecular methods based on the Ligation-mediated PCR principle include:
Amplified fragment length polymorphism (AFLP)
Used for the detection of genomic restriction sequences with the use of PCR. It is used for studying the DNA of any complexity of origin to study the genetic relationship at the species level. The DNA is digested using restriction enzymes, and complete digestion should be ensured to prevent misinterpretation when reading the AFLP. Incompletely digested fragments generate an altered banding pattern, which can be misinterpreted as only polymorphisms. LMPCR is performed, and the results are analyzed using agarose gel electrophoresis, and the patterns are viewed under UV light after being stained with ethidium bromide.
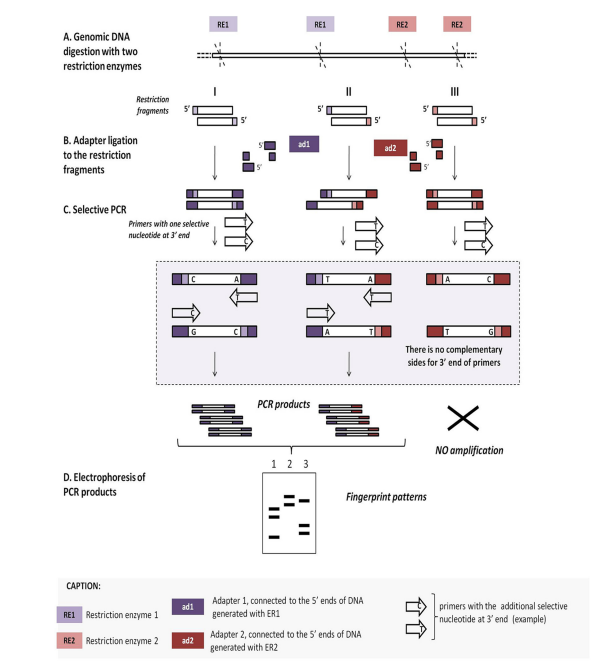
Amplification of DNA fragments surrounding rare restriction sites (ADRSS)
This technique is based on the digestion of the bacterial DNA with two different restriction enzymes that differ in the cleavage frequency. Such a method is used for fingerprinting purposes. Here, ligation is performed with two different oligonucleotide adaptors, and a suppression of PCR is done. PCR suppression, also known as Suppression Subtractive Hybridization (SSH), allows for obtaining a limited amount of restriction fragments. This allows for the mapping of only a limited number of fragments that have been linked with both the adaptor.
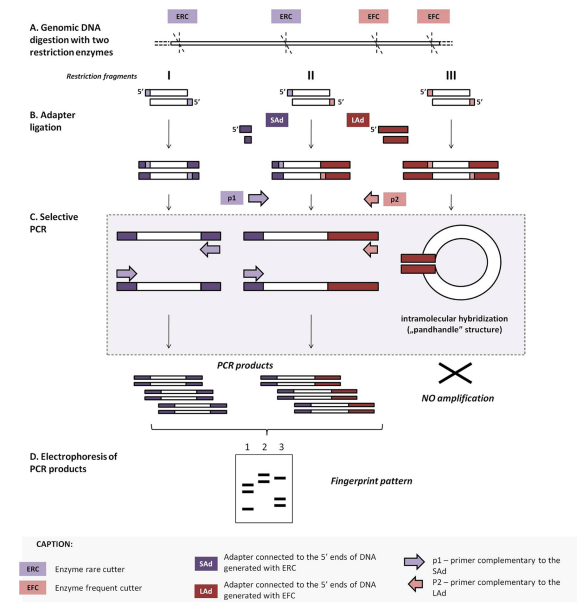
Infrequent-restriction-site amplification (IRSPCR)
Here, double-digestion of the template DNA using two restriction enzymes, one that cuts it infrequently, giving 5’ sticky ends (RE1), whereas the second one cuts the genome frequently, giving 3’ sticky ends (RE2). The end cut with RE1 is ligated with an adaptor with the help of a helper oligonucleotide with a phosphate group at the 5’ end, and the other end with a 3’ end is ligated with an adaptor by using a helper oligonucleotide lacking a phosphate group. The latter helper oligonucleotide is removed during the denaturation step since the 3’ end is modified by a synthetic adaptor lacking a phosphate group. This leads to the loss of the binding sites for the primer. Therefore, amplification of the fragments digested by both RE1 and RE2, and RE1 will be performed.
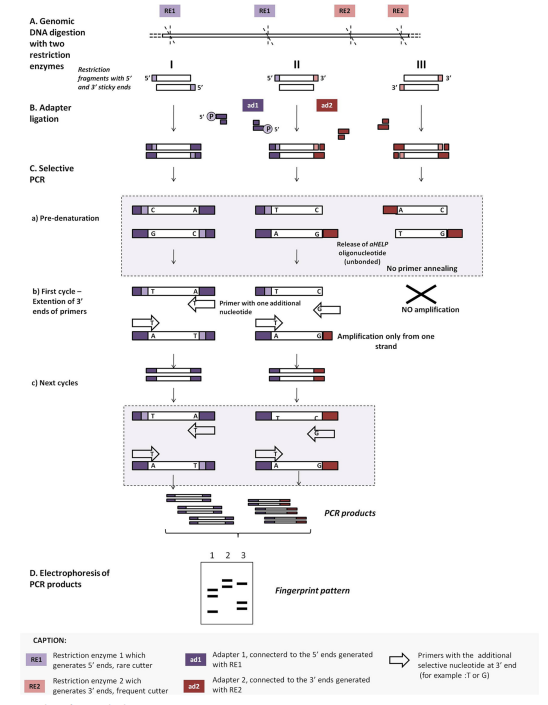
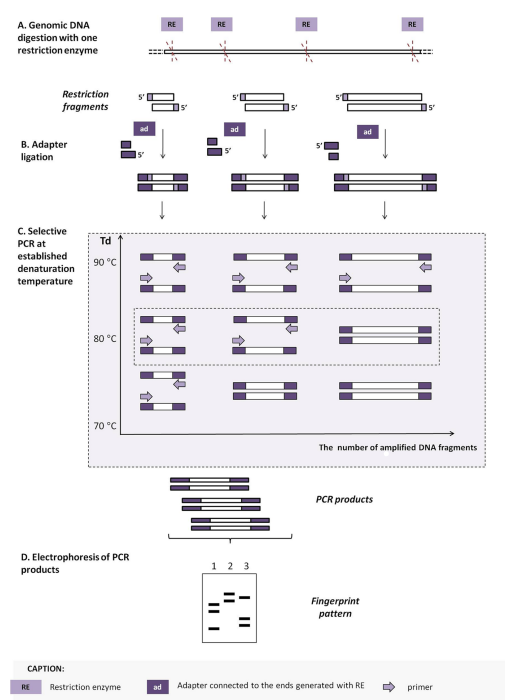
PCR melting profiles (PCR MP)
A method used for genetic typing of microorganisms. Here, the complete digestion of the genomic DNA is done using a single type of restriction enzyme. The resulting 5’ends are then modified by ligating the same synthetic oligonucleotide. Selectivity and the differentiating capacity of this method come from lowering the denaturation temperature. This results in only the single-stranded DNA fragments being amplified.
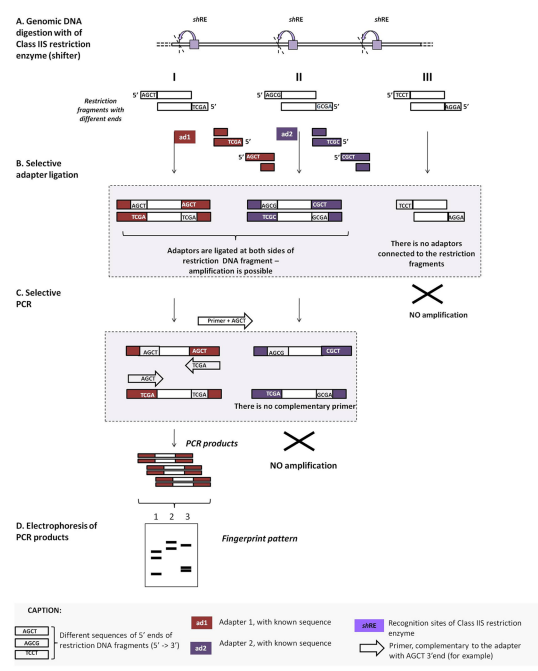
Ligation-mediated PCR/shifter (LM PCR/SHIFTER)
The restriction enzyme used for this method is a Class IIS restriction enzyme, which can recognize non-palindromic DNA sequences between 4 and 7base pairs in length. It cuts up to 20 bases outside their target sites, which helps in creating different types of sticky ends that are up to 5 bases long. Using oligonucleotides with varying degrees of degeneration or mixing several primers in a single reaction can determine the differentiating potential of this method. Genotyping can be done using LMPCR/shifter.
Ligation-Mediated PCR Kits
Ligation-mediated PCR kits are not commercially available because it is a method that requires modification and performance in lab settings.
Applications of Ligation-Mediated PCR
Ligation-mediated PCR is used mainly in genomic sequencing studies. Some of the applications include:
- Used for the determination of primary DNA nucleotide sequences.
- Cytosine methylation patterns can be studied using LMPCR.
- To study and determine the DNA lesion formation and its repair.
- For in vivo protein and DNA footprinting.
- In genome sequencing, DNA ladder analysis, genome walking, and DNA footprinting are widely used.
Advantages of Ligation-Mediated PCR
- Retention of single-base resolution allows the generation of discrete DNA ladders.
- Amplification of specific strands with the use of different types of Ligation-mediated PCR methods.
- Prior sequencing knowledge of the genomic DNA is not required.
- The LMPCR methods are reproducible, simple, and cost-effective.
Limitations of Ligation-Mediated PCR
- The repeatability of the methods can vary depending on the use of a template and personnel experience.
- Chances of contamination during the ligation and end-labeling are high.
- Errors in the method can occur from incomplete restriction digestion.
Conclusion
Ligation-mediated Polymerase Chain Reaction was developed to perform DNA fingerprinting and genomic sequencing. These methods require fragment amplification with only one known end sequence. Here, the use of linkers has allowed for the specific amplification by ligating known linkers to the end of the fragments with the end sequence of interest. Such a method allows for the retention of single-base resolution, which aids in applications requiring that, for example, DNA lesion determination. Different methods of Ligation-mediated PCR are utilized for various applications, and these methods are cost-effective, providing results. Ligation-mediated PCR can serve as a good alternative for differentiating high clonal strains.
References
- Krawczyk, B., Kur, J., Stojowska-Swędrzyńska, K., & Śpibida, M. (2016). Principles and applications of Ligation Mediated PCR methods for DNA-based typing of microbial organisms. Acta Biochimica Polonica, 63(1), 39-52.
- Mueller, P. R., Wold, B., & Garrity, P. A. (2001). Ligation‐mediated PCR for genomic sequencing and footprinting. Current protocols in molecular biology, 56(1), 15-3.
- https://www.ste-mart.com/ligation-mediated-pcr-lm-pcr-151234.htm
- Dai, S. M., Chen, H. H., Chang, C., Riggs, A. D., & Flanagan, S. D. (2000). Ligation-mediated PCR for quantitative in vivo footprinting. Nature biotechnology, 18(10), 1108-1111.
