The Polymerase Chain Reaction is a thermal cycling process invented by Kary Mullis in 1983, used to make multiple copies of nucleic acid regions of interest. In the traditional PCR method, the basic materials include: primers, buffers, DNA polymerase, dNTP’s, etc. PCR is used for various applications, some of which include the detection of single-copy DNA molecules, single-nucleotide polymorphisms, forensic samples, etc. It is also used for cloning and next-generation sequencing applications. With the advent of the PCR reaction, multiple derivatives of the method have come about, each improving aspects of traditional PCR. Hot-start PCR is one such PCR method.
Hot-start PCR is a modified form of polymerase chain reaction in which DNA polymerase is kept inactive at room temperature and only activated during the initial denaturation step, thereby preventing premature primer extension.

Hot-start PCR was developed to reduce non-specific amplification, a common issue in conventional PCR caused by unintended primer binding and extension. By using a chemically or antibody-modified DNA polymerase that remains inactive during reaction setup, the method minimizes primer-dimer formation and off-target amplification. The enzyme becomes active only at high temperature during the first denaturation step, improving both specificity and overall reaction efficiency.
Objectives of Hot Start PCR
The objectives of starting PCR are the same as those of the traditional PCR, but with certain changes, such as:
- Prevent the extension of the primers binding to the template sequences with lower homology (Mispriming).
- To prevent the formation of primer dimers during the amplification.
- To increase the precision, sensitivity, and yield from the reaction.
- Prevent the amplification process from occurring at room temperature,i.e, before entering the thermocycler.
Requirements of Hot Start PCR
Both Hot-start PCR and Traditional have the same requirements, with the former using a modified polymerase. The requirements are as follows:
- Primers: Both 10 µM forward and 10 µM reverse primers are required; the concentration required depends on how much of the reaction mixture is used. The primers to be used are generally 20-40 nucleotides in length with a GC content of 40-60%.
- Reaction buffers: A standard 10x Taq reaction buffer is used.
- DNA Polymerase: The modified hot start Taq DNA polymerase is used.
- Template DNA
- Nuclease-free water
All the components can be prepared before the reaction separately and then mixed before adding them to the thermocycler for the reaction to take place. All of this is mixed, and the liquid is removed by giving a spin if necessary. The PCR tubes are then transferred to the PCR machine to enter the thermocycling process. Due to the hot-start nature of the enzyme, no enzyme activation step is required to remove any inhibitors for the reaction to take place.
Principle of Hot-start PCR
The polymerase chain reaction is a method used to amplify specific DNA targets with the help of the thermocycler. In the case of traditional PCR techniques, during the assay setup, even before the thermal cycling occurs, there are chances of non-specific amplification occurring at room temperature. This leads to inaccurate amplification, providing false products and results. Such inaccurate amplification can be a hindrance when using low concentrations of complex templates, such as mammalian genomic DNA templates.
To prevent such amplifications from occurring, the DNA polymerase can be modified in such a way that the reaction is halted at room temperature and occurs only upon high-temperature conditions. Antibodies, Aptamers, Chemical modifications, Amino acid mutations, temperature-dependent inhibitors, etc, can modify the polymerase.
Steps in Hot-start PCR
The protocol to be followed in Hot-start PCR is:
- Sample preparation
- Reaction mixture preparation
- Amplification
- Product retrieval
Sample preparation
The use of high-quality DNA that has been purified is preferred as it enhances the results of the PCR. The recommended amounts of DNA template for a 50 µl reaction are:
| DNA | Amount |
| Genomic | 1 ng–1 µg |
| Plasmid or viral | 1 pg–10 ng |
Reaction materials preparation
The materials required include primers, dNTPs, reaction buffers, Mg++, and additives, Hot Start Taq DNA Polymerase, etc. To monitor the efficiency of the PCR and detect contamination with the DNA that contains the target sequence, positive and negative controls are required, respectively.
Amplification
The common step in every type of PCR is. Upon addition of the sample and the reaction mixture into the thermocycler, optimal conditions are provided, and the amplification process begins. This step includes
- Denaturation: This is the first step, where the thermocycler heats up to 95 °C for 30 seconds for the template DNA to be denatured into two parental strands.. This is the standard temperature and time for most templates, but for certain GC-rich templates, a longer denaturation of 2–4 minutes at 95°C is required for the amlicons to be produced. Before denaturation, a separate activation step to release the hot start inhibitor from the enzyme is not required. Additionally, this temperature is required for the activation of the Hot-start Taq DNA polymerase.
- Annealing: The primers added attach to the template strands in this step. The temperature is cooled down to 45-65°C. The activated DNA polymerase begins adding the deoxynucleotides. This step is typically 15-60 seconds long.
- Extension: The temperature increases to 65-75°C, and the addition of nucleotides extends for minutes/kb. Extension of sequence-specific primers with the incorporation of nucleotides complementary to the DNA template occurs in this step.
The entire amplification steps occur approximately 35-40 times, and the resulting amplicons are taken for post-amplification analysis.
Types of Hot-start PCR
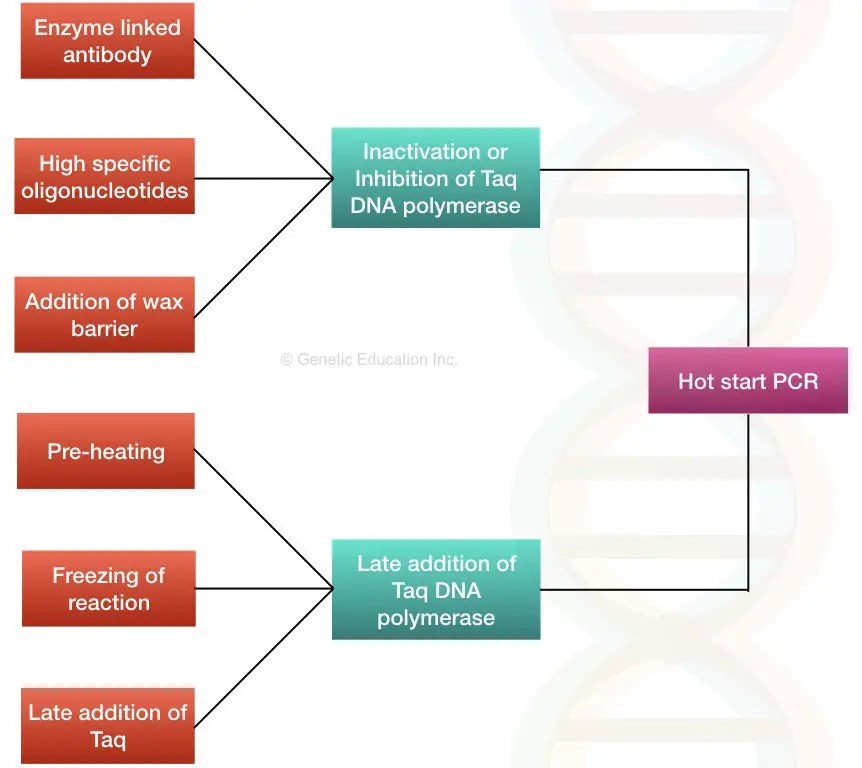
The first time hot-start PCR was practiced, prevention of non-specific amplification was done by withholding a key component during the reaction step, which includes magnesium cofactor, DNA polymerase, etc. Once temperatures are elevated, these components are added for the amplification to take place. However, upon the introduction of manual Hot-start techniques, variations of Hot-start PCR have emerged. Some of these include:
Manual Hot Start/ Pre-heating
This is the first type of Hot-start PCR that was discovered, which gave rise to the newer and upgraded PCR types. Here, the thermocycler is pre-heated, i.e., it is heated to 95°C before the preparation of the reaction. To prevent the non-specific amplification during the preparation of the mixture, the components are prepared on ice at 4°C, which is then immediately transferred to the pre-heated machine. This way, the Taq DNA polymerase cannot start the amplification process early on. Such a method has also proven to release mispaired primers. Preparing the mixture on ice ensures that the activity of the DNA polymerase is lowered, thereby preventing the synthesis of the amplicons. However, this method is not a surefire method since there are still chances of the production of undesirable products.
Barriers/encapsulated reagents
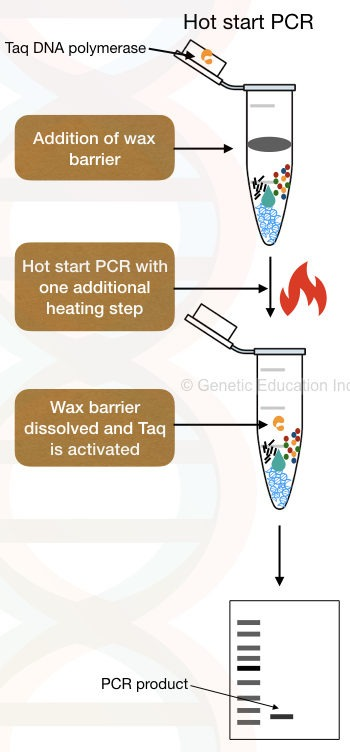
In this type of PCR technique, physical separation of the key components is done. Here, a wax barrier in the form of wax beads is added, which serves as a barrier between the Taq polymerase and other PCR components. The wax used is temperature-sensitive and therefore melts when the right temperature is provided. The wax beads can be used as a barrier in two ways:
- The DNA polymerase containing the wax beads is placed inside the ‘tube-mouth’ while the other components are placed at the bottom of the tube.
- In this method, instead of using wax beads with the NDA polymerase already incorporated in them, the beads are placed in the tube, and the enzyme and MgCl2 are added to the surface of these beads. Once the temperature rises, the beads melt, allowing the components to mix with the reaction, and synthesis begins.
This method is widely optimized and quite accurate.
- DNA polymerase-mediated
The DNA polymerase is modified to prevent the non-specific amplification. The modifications can be done by:
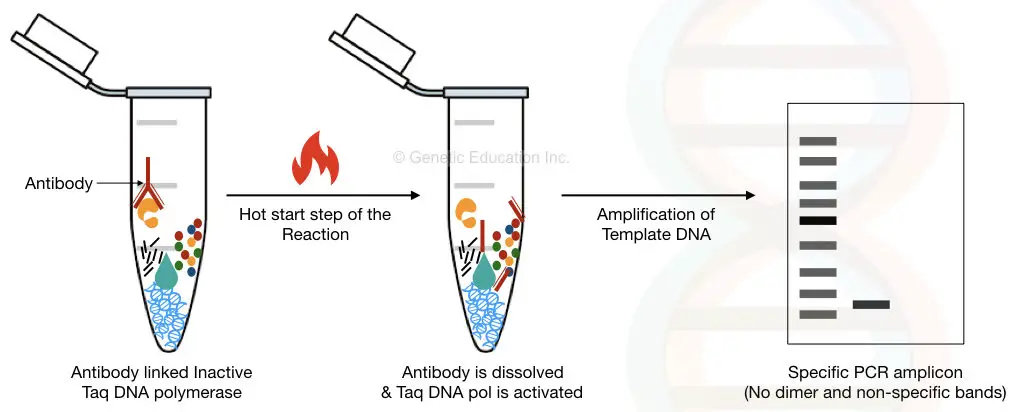
Antibodies: They are used to block the enzyme activity at a lower temperature. They do so by binding to the enzyme active site and dissociating at elevated temperatures since the antibody used is temperature sensitive. The features of the enzyme are unchanged, since the antibody doesn’t alter its structure. The limitation of such a method is the cost, since enzymes require specific antibodies. The antibody to be used should be of animal origin.
- Aptamers: Aptamers are oligonucleotides that can bind to the active site of the enzyme and prevent the amplification at room temperature. Dissociation of the aptamer occurs at elevated temperatures. Unlike antibodies, aptamers are free of animal origin. However, aptamers may not work well with primers of low melting point.
- Chemical Modification: Polymerases are covalently linked with chemical groups to prevent non-specific amplification at room temperature. This method requires a longer activation time.
- Affibody molecules: Affibody molecules are alpha helical peptides that can bind at the enzyme active site to prevent amplification. It has a short activation time.
- Fusion protein: This induces a hyperstable DNA-binding domain and topoisomerase
Freezing the PCR reaction
The PCR reaction mixture is frozen within the tube. To the frozen mixture, the Taq DNA polymerase and PCR additives (like MgCl2) are added. The tubes are then placed in the thermocycler, and the process takes place as usual. This method prevents non-specific amplification, early polymerization, and primer dimers.
The addition of Taq DNA polymerase separately
The reaction mixture is prepared without the addition of the enzyme. It is added only when the mixture is added to the thermocycler, and it reaches the optimal temperature for the amplification process. The limitation of this method is the higher chance of cross-contamination when opening the thermocycler lid in the middle of the process, which may also lead to reaction failure.
Examples of Hot-Start PCR Kits
DreamTaqTM Hot Start PCR master mix (ThermoFisher)
- A 2x reaction mixture with the DNA polymerase, dNTPs, Buffer, and Magnesium.
- Allows for easy setup with fewer pipetting steps, with only the primer, template, and water to be added.
- The kit contains a density reagent and two tracking dyes for direct loading of the PCR products on a gel.
- The DreamTaq DNA polymerase is a modified enzyme for higher specificity, yields, and sensitivity compared to conventional hot start polymerases available in the market.
- The DreamTaq DNA polymerase utilizes antibody-based inhibition to prevent non-specific amplification at room temperature.
- Higher yield of amplicons from low concentrations of template and reaction specificity are the benefits of this kit. The enzyme also allows for genomic fragment amplification of templates up to 6kb in length.
Q5® Hot Start High-Fidelity 2X Master Mix (New England BioLabs)
- Provides high fidelity amplification of ~280 higher than the conventional Taq Polymerase and ultra-low error rates.
- The master mix contains the Q5 Hot Start High-Fidelity DNA Polymerase, dNTPs, and Mg2+.
- High performance is seen for a broad range of amplicons from high AT to GC content.
- The hot start ability of the enzyme comes from the modification with eh help of an aptamer.
- Provides speed, fidelity, and reliability, additionally preventing amplification at room temperature.

2X Hot-Start PCR Master Mix (biotech rabbit)
- High PCR sensitivity without prolonged reactivation and high-throughput hot start PCR up to 3kb.
- Allows for amplification of low copy number targets.
- Has a fast reaction setup by reduced calculation time, reduced pipetting requirement, and no need for buffer optimization.
- The enzyme is bound with the antibody, and thereby amplification occurs at elevated temperature. Minimizes primer-dimers and mispriming.
GoTaq® Hot Start Polymerase (Promega)
- The GoTaq® Hot Start Polymerase is bound to an antibody which blocks the enzyme activity at low temperatures.
- The kit is supplied with 5X Green GoTaq® Flexi Buffer and 5X Colorless GoTaq® Flexi Buffer. These buffers contain a compound that allows for an increased sample density, helping the samples to easily sink into the wells of an agarose gel.
- The green buffer contains a yellow and blue buffer that separates for easy monitoring during electrophoresis.
- The green reaction buffer is used for direct gel analysis, and the colorless buffer for amplification in cases where there may be hindrance during post-amplification analysis, such as fluorescence or absorbance testing.
Applications of Hot-start PCR
- It can be used for the diagnosis of monogenic defects at the DNA level of a single cell.
- Used for templates with low copy number (less than 10⁴ copies).
- In case of complex templates, such as mammalian genomic DNA, hot start PCR is useful.
- Used for reactions with several pairs of oligonucleotide primers
Advantages of Hot-start PCR
- Prevents the chances of non-specific amplification at low temperatures.
- Reduce the occurrence of mmispriming and primer-dimer formation, a nd non-specific bindings.
- Since the modification enzyme prevents non-specific amplification, the reaction mixture can be prepared at room temperature.
- Improve the yield of the product and the accuracy of the reaction.
Limitations of Hot Start PCR
- Due to modifications that are required for the DNA polymerase, this method is expensive and tedious.
- The high temperature required for the activation of the DNA polymerase may damage and denature the template DNA
- Useful results cannot be obtained for templates longer than 2kb.
Conclusion
Conventional PCR techniques often yield improper products due to non-specific synthesis of the target DNA. Such non-specific amplification is the result of the enzyme activation at room temperature during the reaction setup. To combat this issue, Hot-Start PCR was invented. Here, the DNA polymerase is modified in such a way that its activity is not affected while allowing for its activation only at the elevated temperature provided by the thermocycler. This increases the specificity, sensitivity, and reliability of the reaction, providing proper amplicons for post-amplification analysis. Despite the cost that comes with the modification of the enzyme, this is a useful method for PCR since it prevents the non-specific amplification, mispriming, and the formation of primer-dimers.
References
- https://geneticeducation.co.in/what-is-a-hot-start-pcr/#3_Freezing_the_PCR_reaction
- https://www.promega.in/products/pcr/taq-polymerase/gotaq-hot-start-polymerase/
- https://www.thermofisher.com/order/catalog/product/in/en/K9011
- https://www.neb.com/en/products/m0494-q5-hot-start-high-fidelity-2x-master-mix
- https://www.biotechrabbit.com/hot-start-pcr-master-mix-2x.html
- https://www.thermofisher.com/in/en/home/brands/thermo-scientific/molecular-biology/molecular-biology-learning-center/molecular-biology-resource-library/spotlight-articles/hot-start-technology-benefits-PCR.html
- Paul, N., Shum, J., & Le, T. (2010). Hot start PCR. In RT-PCR Protocols: Second Edition (pp. 301-318). Totowa, NJ: Humana Press.
- Paul, N., Shum, J., & Le, T. (2010). Hot start PCR. In RT-PCR Protocols: Second Edition (pp. 301-318). Totowa, NJ: Humana Press.
- https://www.sigmaaldrich.com/IN/en/technical-documents/technical-article/genomics/pcr/what-is-hot-start-pcr
- Kim, Sung Ah, et al. “Application of Hot Start PCR Method in PCR-based Preimplantation Genetic Diagnosis.” Journal of genetic medicine 9.1 (2012): 11-16.
- https://www.bocsci.com/resources/hot-start-pcr-definition-protocol-and-application.html
