The accuracy of the polymerase chain reaction (PCR), a method for copying DNA, is a key factor in the proper transfer of genetic information. One of the main components responsible for this accuracy is DNA polymerase fidelity.
DNA polymerase is the enzyme that builds new DNA strands during PCR. Fidelity refers to how well the DNA polymerase inserts the correct base (A, T, C, or G) into the growing DNA strand, matching the template strand. Maintaining the sequence, or order, of DNA bases relative to the template strand is important to ensure that genetic information is properly copied. The rate at which the enzyme inserts the wrong base is called the Polymerase’s error rate.
To overcome such errors, high-fidelity DNA polymerases were developed. These are specially engineered enzymes that produce accurate DNA copies with fewer errors.
High-fidelity PCR uses specialized DNA polymerases with low error rates, providing high-accuracy amplification in DNA replication.
Scientists at New England Biolabs were the first to identify and commercialize Vent® DNA Polymerase, a high-fidelity enzyme suitable for PCR. With advancements in technology, many high-fidelity DNA polymerases have been engineered to have better accuracy, strength, and speed.
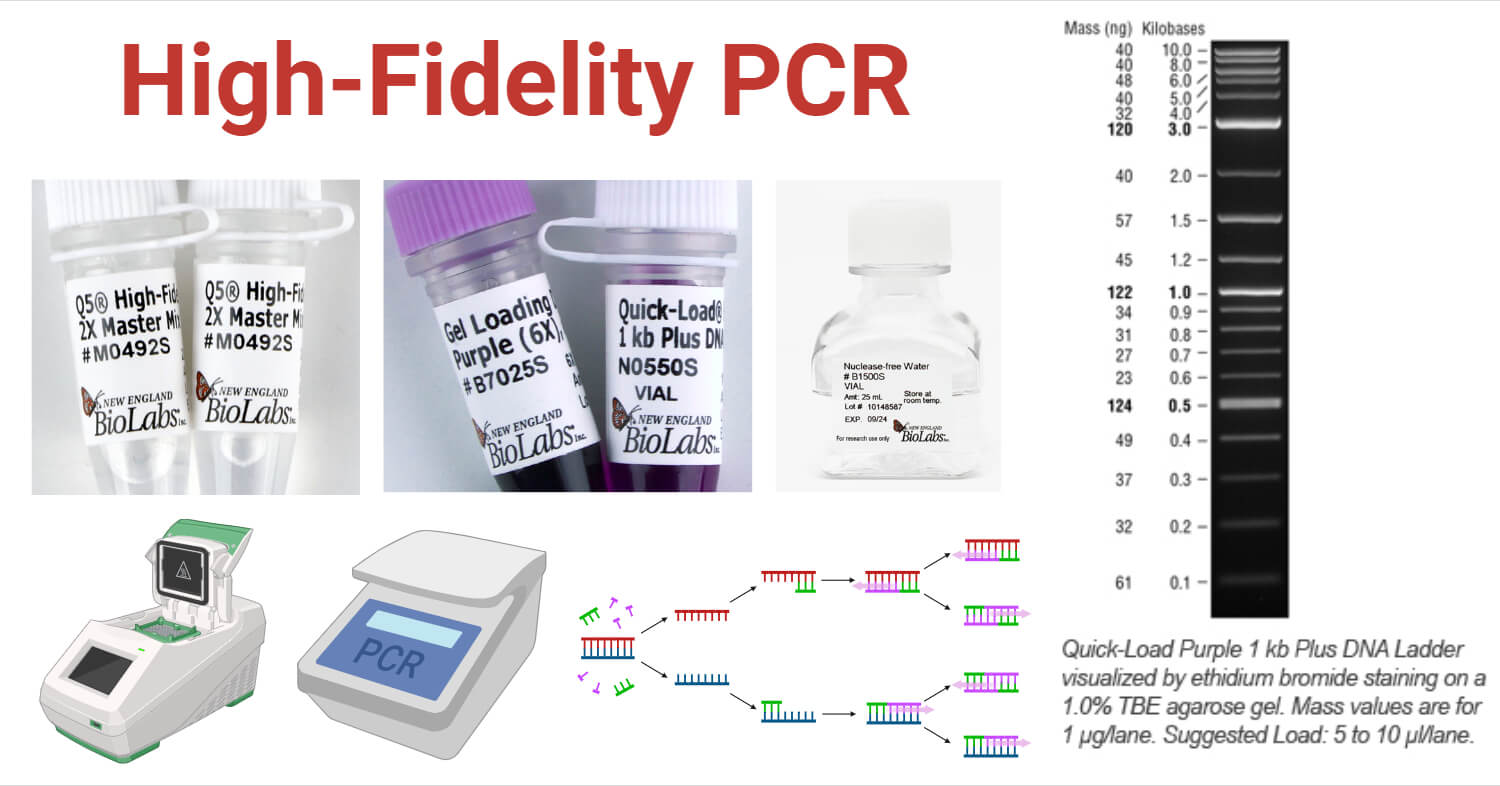
Objectives of High-Fidelity PCR
In a regular polymerase chain reaction (PCR), short strands of DNA called primers can attach to different parts of the template DNA. The 3’ end of a primer (the end with a free hydroxyl group) should match the target sequence exactly to allow for proper DNA amplification. However, if the primer’s 3’ end does not match perfectly, it can lead to non-specific DNA copies. To prevent this, researchers can add a chemical group called phosphorothioate to protect the primer’s 3’ end or use special DNA polymerases (such as Taq polymerase) that have a built-in proofreading ability. Proofreading means that the polymerase can remove incorrect bases using its 3’ to 5’ exonuclease activity (an enzyme function that removes bases from the end of DNA). Because this activity could also remove bases from primers, primers are often modified to protect them and reduce errors.
The objective of using high-fidelity polymerases is to reduce the chances of improper nucleotide incorporation during the amplification process. This reduces the Polymerase error rate and thereby reduces any chance of improper product and false-positive results.
Requirements for High-Fidelity PCR
The High-Fidelity PCR utilizes similar materials to those of traditional PCR, which include:
- Primers: Oligonucleotide primers (short pieces of single-stranded DNA) are generally 20–40 nucleotides long and ideally have a GC content (proportion of guanine and cytosine bases) of 40–60%.
- dNTP’s
- Reaction buffer which includes Tris-HCl, (NH4)2SO4, KCl, MgSO4, and Triton® X-100
- DNA polymerase: Taq polymerase that has been modified to provide high fidelity
- Thermocycler
Principle of High-fidelity PCR
DNA replication by PCR can be considered to be accurate if the DNA polymerase utilised can accurately read the template DNA to incorporate appropriate nucleotide bases. However, there are cases when correct base pairing is not performed due to the inaccuracy of the enzyme utilised. Such mistakes lead to permanent mutations in the product. Therefore, to prevent such mistakes, DNA polymerases can be modified to enhance their proofreading ability. This allows for the detection and correction of mistakes, thereby reducing the chances of mutations.
The geometry of the polymerase is such that the active site determines the selection of correct nucleotide sequences to align the catalytic groups to ensure efficient incorporation. In cases when incorrect nucleotide sequences are incorporated, this results in a time lag, which is needed to correct the mistake and allow for the process to start again with the correct nucleotide. Additionally, the polymerase also has a proofreading domain, i.e, it has a 3’-5’ exonuclease activity, which adds to the protection of incorporation of the wrong nucleotide. The enzymatic removal of incorrect nucleotides in the growing DNA strand from the 3’ end is referred to as proofreading.
Such an ability of the enzyme is what determines its fidelity. Fidelity, accuracy with which the polymerase can incorporate bases, is important to obtain correct replicates after amplification for various applications such as cloning of DNA for protein expression, SNP analysis, and next-generation sequencing applications. Enhancing the fidelity of DNA polymerase can help reduce any mistakes during the amplification, which can be done through improving the proofreading properties of the enzyme by genetic modification, chemical modification, mixing of several enzymes, etc.
High-fidelity DNA polymerases help in minimizing the mistakes that occur during DNA replication. They safeguard by having a significant binding preference for the correct nucleotide during the polymerization process. It utilizes the lag time and 3’-5’ exonuclease activity to detect and correct the mispaired bases and thereby reduces the chances of mutation. Upon detection, the chain is moved back to the polymerization domain, allowing polymerization to occur again.
Steps in High Fidelity PCR
High-fidelity PCR follows the same protocol as traditional PCR, but with a modified DNA polymerase. The protocol is as follows:
- Sampling: The sample to be replicated is obtained from the necessary sources.
- Amplification: This is done in the thermocycler, where the optimal conditions for DNA replication are provided.
- Analysis
1. Sampling
The target DNA is collected and prepared for the PCR process. The DNA polymerase is diluted in the buffer before use to reduce the chances of errors during pipetting. All the reaction components are added to a thin-walled PCR tube and mixed thoroughly to ensure a homogeneous, consistent mixture. It is better to use a pure template as it determines the amplicons produced.
2. Amplification
Upon preparation of the sample, it is then quickly transferred to the thermocycler, which is pre-heated to 98°C for the denaturation steps. This is the standard PCR reaction, which includes three steps:
- Denaturation: In this step, the template DNA is denatured by breaking the hydrogen bonds at the pre-heated temperature. This takes about 30 seconds and can even be longer (up to 3 minutes) depending on the type of template.
- Annealing: The step that follows denaturation, during which the binding of the primers to the template DNA occurs. The lowered temperatures aid in this process and set up the DNA polymerase for the amplification process. The temperature is lowered to 55 °C.
- Extension: The final step in the amplification process is the extension of the growing DNA strand. The DNA polymerase incorporates the correct nucleotide sequences to amplify the segment of the DNA of interest. The temperature is raised to 72°C for this process, and the process generally takes about 20-30seconds for 1kb of templates. The DNA polymerase adds the nucleotides to the annealed primer ends. In case of long products, i.e, more than 6kb, a longer extension time is preferred, up to 40-50 seconds/kb.
A single PCR cycle takes about 5 minutes, with the temperature raised back to 95°C – 98°C to repeat the cycle. The number of copies generated doubles with each cycle. To obtain a sufficient number of DNA copies, 25-30 cycles is the preferred amount.
3. Analysis
The DNA amplification can be monitored nd analysed with the use of fluorescent dyes, which can bind to the DNA or can be sequence specific in their binding. There is a quantification cycle, referred to as the fractional cycles, for the fluorescence to be able to be measured.
Types of High-Fidelity PCR
There are two types of high-fidelity PCR, they are:
- Classical High Fidelity PCR: This is the traditional PCR method with the use of a high-fidelity DNA polymerase whose properties have been enhanced by genetic modification, chemical modification, and mixing of several enzymes.
- Hot start DNA polymerase: This utilises a modified DNA polymerase referred to as Hot start. Such an enzyme avoids the use of non-specific amplification of DNA by the inactivation of the Taq polymerase at low temperatures with the use of specific antibodies.
Examples of High-Fidelity PCR Kits
Multiple companies have come up with PCR kits that utilise the high-fidelity PCR technology. Some of them are listed below:
1. Q5® High-Fidelity PCR Kit (New England Biolabs)
- This kit with the Q5 high-fidelity DNA polymerase shows a robust performance.
- The polymerase shows almost 280 times higher amplification than Taq polymerase with ultra-low error rates.
- The DNA polymerase utilised is engineered to be able to fuse to the processivity-enhancing Sso7d DNA binding domain, thereby enhancing the speed, fidelity, and reliability of the performance.
- High performance is seen for a broad range of amplicons from high AT to GC content.
2. iProof High-Fidelity DNA PCR kit (Bio-Rad)
- The DNA polymerase used in the kit is a unique Pyrococcus-like proofreading enzyme, which is fused to the Sso7d dsDNA-binding protein.
- The enzyme is a thermostable fusion polymerase that can accurately amplify long products from diverse DNA templates.
- The enzyme used is 52-fold more accurate than Taq polymerase.
- It has high processivity and reduces extension up to 15-30 sec/kb and thereby the overall reaction times.
- Longer templates of up to 37 kb can be amplified in less time while utilising less enzyme (0.25-1unit/reaction)
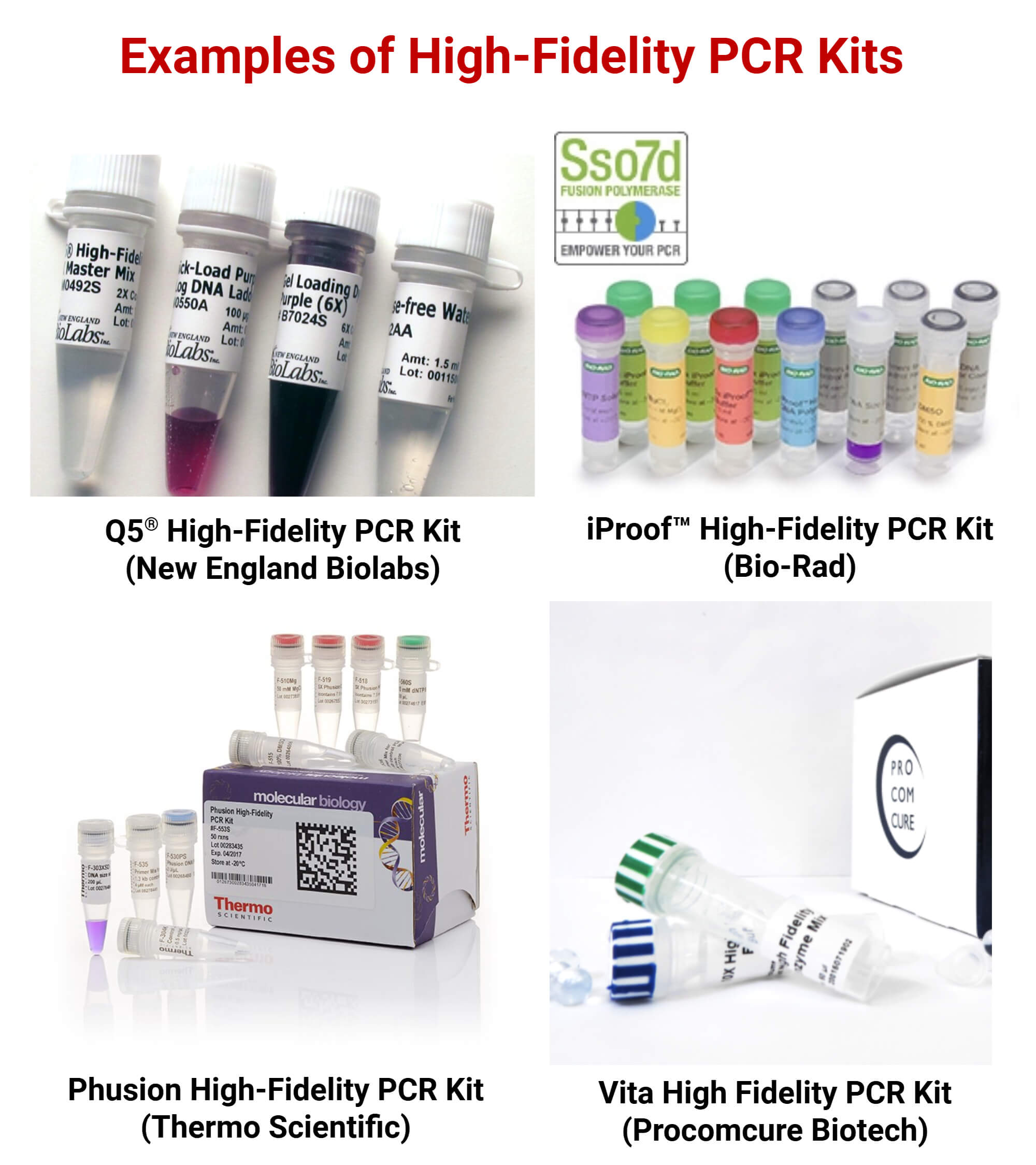
3. Phusion High Fidelity PCR Kit (ThermoFisher)
- The DNA polymerase in this kit has an error rate of 50-fold less than that of Taq polymerase and 6-fold less than that of Pfu.
- This kit provides robust performance with short protocol time, even in the presence of PCR inhibitors.
- The copies produced are much higher with less requirement for the enzyme, unlike other DNA polymerases on the market.
- The kit contains all the necessary reagents, including control lambda DNA template and primers for 1.3kb to 10 kb amplicons.
- It has a short extension time (15-30 s/kb), thus providing faster performance.
Applications of High-Fidelity PCR
For applications that require error-free products, fidelity is important. Some of these include:
- Cloning: For cloning/subcloning purposes, it should be ensured that the in vitro amplified material is an exact copy of the template. This is important for protein expression and gene studies.
- Single Nucleotide Polymorphism: For SNP analysis, done by cloning and sequencing.
- RNA analysis for Reverse Transcriptase Polymerase Chain Reaction.
- Any application that requires in vitro amplified material.
In cases where only the presence or absence of the product is required, Fidelity is not applicable.
Advantages of High-fidelity PCR
Advantages of High Fidelity PCR include:
- Produces error-free copies of the template DNA.
- Reduces the chances of permanent mutation in the product.
- Highly sensitive and therefore allows for smooth extension.
- High precision reduces the time taken for amplification
- More efficient compared to Taq polymerase
Limitations of High-Fidelity PCR
Some of the limitations of this method include:
- It is an expensive process since the DNA polymerase should be modified through chemical means, genetic modification, etc.
- Sometimes, the annealed primers may get proofread by the exonuclease activity, which can lead to mismatched extension of the edited primer.
- In cases of longer templates, the amplification process becomes more complex, and thus the reaction time increases.
- In certain kits, the modified DNA polymerases require a higher temperature for them to work, which can lead to DNA damage.
Conclusion
Fidelity is a feature of DNA polymerase, which scythe-like ability to incorporate the correct nucleotide bases with minimal error. This feature comes from the 3’-5’ exonuclease activity, i.e, the proofreading property of the enzyme. By exploiting this feature, the enzymes can be modified via genetic engineering, chemical means, etc., to enhance their fidelity. By doing so, PCR-produced copies have fewer errors, thereby reducing the chances of mutation.
High-Fidelity PCR has found application in cloning, RNA analysis, SNP analysis, Next Generation Sequencing, and so on. Despite the limitations of this method, its pros outweigh the cons, making it useful for PCR.
References
- https://www.thermofisher.com/in/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/dna-polymerase-characteristics.html
- https://www.thermofisher.com/in/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/dna-polymerase-characteristics.html
- https://www.thermofisher.com/in/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/dna-polymerase-characteristics.html
- https://www.neb.com/en/products/e0555-q5-highfidelity-pcr-kit
- Pezza, J. A., Kucera, R., & Sun, L. J. N. E. B. (2014). Polymerase fidelity: what is it, and what does it mean for your PCR. New England Biolabs.
- Yang, H. L., Jiang, H. J., Fang, W. Y., Xu, Y. Y., Li, K., Zhang, J., … & He, F. C. (2005). High fidelity PCR with an off/on switch mediated by proofreading polymerases combining with phosphorothioate-modified primer. Biochemical and biophysical research communications, 328(1), 265-272.
- https://www.thermofisher.com/in/en/home/life-science/cloning/cloning-learning-center/invitrogen-school-of-molecular-biology/pcr-education/pcr-reagents-enzymes/dna-polymerase-characteristics.html
- https://www.ncbi.nlm.nih.gov/books/NBK589663/
- https://www.britannica.com/science/polymerase-chain-reaction
