The genome serves as the comprehensive biological library of an organism, containing all the hereditary information required to build, operate, and repair a living being.
Its fundamental function is to provide the precise codes for protein production and to ensure that life’s developmental blueprint is passed accurately from one cell to its successors. Genome stability is the critical state in which a cell successfully protects this library, using specialized repair mechanisms to maintain the integrity of DNA over time.
If this stability is lost, the genetic instructions become corrupted, leading to a cascade of cellular failures. A lack of stability results in genomic turbulence where mutations occur rapidly, often leading to the onset of malignant tumours, premature cellular aging, and the failure of vital organ systems. Consequently, genome stability is of paramount importance because it acts as the primary defence against life-threatening diseases and ensures the faithful continuity of biological information.
Sources of Genome Instability: Endogenous vs. Exogenous Factors
Genome is a biological library of an organism, functioning as a master manual that stores all hereditary information and genetic instructions. To protect this vital record, cells rely on genome stability. This stability is constantly challenged by two primary categories of threats:
- Endogenous sources, which are internal pressures like DNA replication errors, metabolic byproducts such as reactive oxygen species (ROS), and spontaneous chemical changes within the cell
- Exogenous sources, which are external environmental stressors like ultraviolet (UV) radiation, ionizing radiation, and exposure to chemical mutagens from tobacco or pollutants.
The following table provides specific examples of these endogenous and exogenous sources, highlighting how everyday biological processes and environmental exposures can physically alter our molecular blueprint.
| Source Type | Causes | Examples | Type of DNA damage |
| Endogenous Sources | Metabolism | Reactive Oxygen Species (ROS) from mitochondria | Oxidized bases, single-strand breaks |
| Replication | DNA polymerase errors during S-phase | Base mismatches, insertions, or deletions | |
| Biochemistry | Spontaneous hydrolysis (water reactions) | Depurination (loss of A/G bases) | |
| Metabolic Products | Aldehydes (e.g., from alcohol or digestion) | DNA adducts and cross-links | |
| Exogenous Sources | Radiation | UV sunlight, X-rays, cosmic rays | Thymine dimers, double-strand breaks |
| Chemicals | Tobacco smoke, air pollutants, charred food | Bulky DNA adducts (e.g., Benzopyrene) | |
| Medicine | Chemotherapy drugs | Interstrand cross-links | |
| Biological | Viruses (HPV, Hepatitis B) | Viral DNA integration into the host genome |
If these internal and external threats overwhelm the cell’s repair systems, the resulting genomic instability leads to the disruption of the genetic code, which can trigger catastrophic events such as the development of cancer, accelerated aging, and the loss of vital cellular functions.
The DNA Damage Response (DDR): Signalling Pathways (ATM and ATR)
The DNA Damage Response (DDR) is a robust, hierarchical signal transduction network essential for maintaining genomic integrity. Rather than a simple repair mechanism, the DDR functions as a comprehensive surveillance system that coordinates DNA lesion detection with downstream cellular outcomes.
Depending on the extent of the damage, the DDR orchestrates one of three primary fates:
- Cell Cycle Arrest: Activation of checkpoints to allow time for enzymatic repair.
- Senescence: Permanent exit from the cell cycle for damaged but viable cells.
- Apoptosis: Programmed cell death to prevent the propagation of oncogenic mutations.
At the apex of this signalling cascade are two members of the PI3K-related kinase (PIKK) family: ATM (Ataxia-Telangiectasia Mutated) and ATR (Ataxia-Telangiectasia and Rad3-related). These kinases act as the primary transducers that convert physical DNA damage into a biochemical signal.
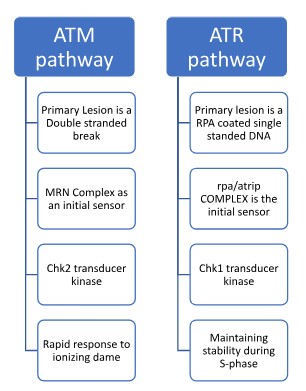
- ATM: Ataxia-Telangiectasia Mutated:
ATM is the primary mobilizer in the event of Double-Strand Breaks (DSBs). These breaks are highly cytotoxic and can be induced by ionizing radiation or oxidative stress. ATM is recruited to the lesion by the MRN sensor complex, which recognizes the physical break in the phosphodiester backbone. Once active, ATM phosphorylates the histone variant H2AX (creating gamma H2AX) to mark the site and activates the effector kinase Chk2.
- ATR: Ataxia-Telangiectasia and Rad3-related
ATR serves a broader, more essential role in monitoring Single-Stranded DNA (ssDNA) gaps, which are hallmark intermediates of UV-induced damage and replication stress (e.g., stalled replication forks). ATR is recruited to ssDNA that has been coated with Replication Protein A (RPA). In conjunction with its partner protein ATRIP, ATR activates Chk1, which is critical for stabilizing replication forks and preventing mitotic catastrophe.
The synergy between ATM and ATR ensures that the genome is monitored throughout all phases of the cell cycle.
Cell Cycle Checkpoints: The Guardians of Genetic Integrity
The cell cycle represents a highly orchestrated sequence of events through which a cell replicates its genetic material and divides to produce two identical daughter cells, serving as the fundamental mechanism for growth, development, and tissue repair. This process is primarily divided into interphase comprising the G1 (growth), S (DNA synthesis), and G2 (preparation) phases and the mitotic (M) phase, where nuclear and cytoplasmic division occurs. To preserve the fidelity of this complex transition, the cell employs a sophisticated network of internal checkpoints that act as guardians of genetic integrity.
At the G1 transition, the tumour suppressor protein p53 evaluates DNA integrity; upon detecting lesions, it stimulates p21 to inhibit Cyclin-Dependent Kinases (CDKs), thereby arresting the cycle. Simultaneously, the Retinoblastoma protein (Rb) functions as a gatekeeper by sequestering the E2F transcription factor, ensuring the cell does not initiate DNA synthesis in the S phase until environmental conditions are optimal.
Following replication, the G2 checkpoint serves as a final quality-control station. Here, the Cyclin B-CDK1 complex is often termed as the maturation-promoting factor monitor for incomplete replication or double-strand breaks, preventing the onset of mitosis in compromised cells. The final safeguard occurs during the M phase, where the Anaphase-Promoting Complex (APC/C) verifies the stable attachment of spindle fibres to kinetochores. Only when chromosomal alignment is perfected does the APC/C facilitate the activation of separase, an enzyme that triggers the symmetrical division of sister chromatids. By enforcing these stringent molecular protocols, the cell minimizes the risk of aneuploidy and oncogenic transformations.
Mechanisms of DNA Repair: BER, NER, and Mismatch Repair
To maintain genomic integrity, cells utilize DNA repair mechanisms to detect and fix lesions that threaten the genetic code. DNA is under persistent pressure from internal factors, such as reactive oxygen species generated during metabolism, and external stressors like UV light and toxins. These repair pathways act as a critical defence if they fail, the resulting accumulation of mutations can trigger unregulated cell growth, leading to cancer or premature cell death.
- Base Excision Repair: BER is the primary defense against localized, non-distorting damage, such as bases altered by oxidation or alkylation. Because these small lesions do not significantly warp the DNA’s physical structure, specialized enzymes called DNA glycosylases must actively scan the genome to flip out and identify damaged bases. DNA glycosylases patrol the helix, detecting and snipping out specific damaged bases. This leaves an empty spot called an AP site. An endonuclease then nicks the DNA backbone, allowing DNA polymerase to insert the correct replacement and ligase to seal it shut.
- Nucleotide Excision Repair: NER targets helix-distorting damage that creates significant physical obstacles in the DNA strand. Examples include covalent adducts and UV induced thymine dimers cross-links that warp the geometry of the double helix and can stall vital processes like transcription. NER performs a patch-based repair. It excises a wide segment of nucleotides (approx. 24–32 bases) surrounding the distortion. The gap is then filled by DNA polymerase using the non-defective strand as a guide.
- Mismatch Repair: MMR acts as a post-replication checker. It identifies mispaired bases (like a G paired with a T) or small insertions/deletions that occurred during DNA synthesis. It distinguishes the newly synthesized daughter strand from the original parental template to ensure it fixes the error rather than the original code. It removes a large section of the new strand containing the mismatch and resynthesizes it with high precision.
Double-Strand Break Repair: Homologous Recombination vs. NHEJ
Double-strand breaks (DSBs) are characterized by the simultaneous cleavage of both strands of the DNA phosphodiester backbone. This damage is highly cytotoxic, as it can lead to chromosomal translocation or fragmentation. To preserve genomic integrity, eukaryotic cells utilize two sophisticated biochemical pathways: Non-Homologous End Joining (NHEJ) and Homologous Recombination (HR).
- Non-Homologous End Joining (NHEJ): NHEJ is a template-independent repair mechanism that operates throughout the cell cycle, though it is most prevalent in the G1 phase. The process is initiated by the binding of the Ku70/Ku80 heterodimer to the exposed DNA termini. This complex recruits the DNA-dependent protein kinase catalytic subunit (DNA-PKcs) to form a synaptic bridge. Because the broken ends are often chemically incompatible, nucleases (such as Artemis) process the ends. This nucleolytic processing frequently results in the loss or gain of nucleotides, rendering NHEJ an error-prone or mutagenic pathway. Final ligation is executed by the DNA Ligase IV/XRCC4 complex.
- Homologous Recombination (HR): HR is a template-directed, high-fidelity repair modality restricted to the late S and G2 phases, when a sister chromatid is available to serve as a genetic blueprint. The process begins with resection, where the 5′ ends of the break are enzymatically degraded to create 3′ single-stranded DNA (ssDNA) overhangs. These overhangs are coated by RPA proteins, which are subsequently replaced by RAD51 (a process mediated by BRCA2). This nucleoprotein filament facilitates strand invasion into the homologous sister chromatid, forming a D-loop. DNA polymerase then extends the invading strand using the intact template as a guide, ensuring an error-free restoration of the original sequence.
Replication Stress: Stalled Forks and Genome Fragility
Replication stress occurs when the DNA replication machinery (the replisome) encounters obstacles that impede its progress, threatening the fidelity of genome duplication. When a replication fork encounters a physical or chemical barrier such as DNA lesions, transcription machinery collisions, or secondary structures the polymerase stalls while the helicase may continue unwinding. This uncoupling leads to an accumulation of single-stranded DNA (ssDNA).
To manage this, the cell triggers the ATR-mediated checkpoint, which stabilizes the stalled fork and prevents its collapse. If this protection fails, the replisome dissociates, leading to fork collapse and the formation of a double-strand break (DSB).
Genome fragility is particularly evident at Common Fragile Sites (CFS). These specific genomic regions are intrinsically difficult to replicate due to their length or complex sequences. Under conditions of replication stress, these sites often remain unreplicated as the cell enters mitosis. The resulting unfilled gaps lead to chromosomal breakage during anaphase, causing large-scale deletions and translocations. This cycle of stress and fragility is a primary driver of chromosomal instability, a hallmark of cancer development and cellular aging.
The Role of Telomeres and Telomerase in Chromosome Stability
To ensure genomic integrity, linear chromosomes are capped at their ends by specialized nucleoprotein complexes known as telomeres. These structures consist of repetitive DNA sequences and associated proteins that prevent the cell’s surveillance machinery from incorrectly identifying natural chromosome termini as hazardous double-strand breaks (DSBs).
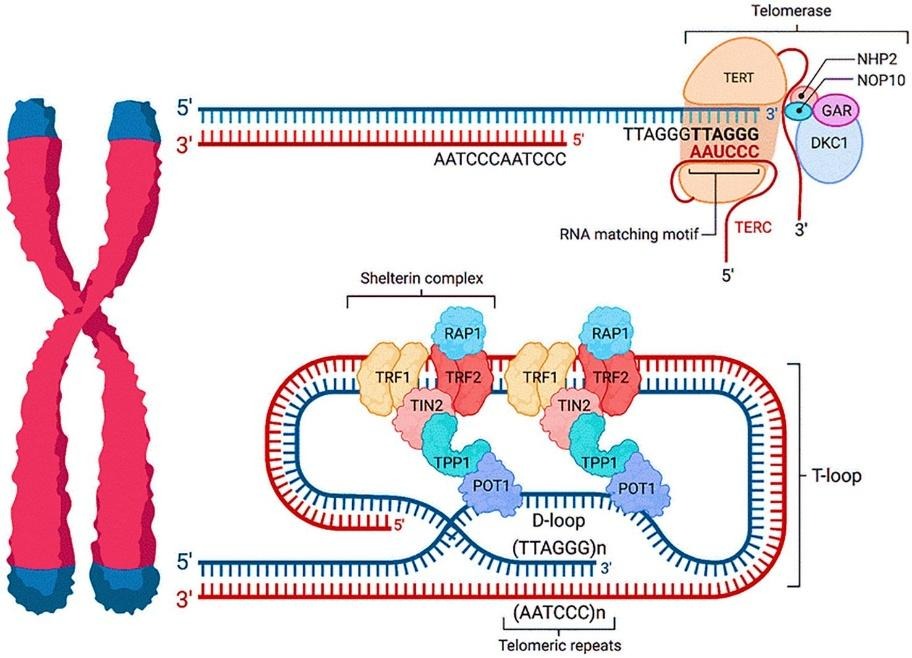
- Telomeres: Telomeres are repetitive nucleoprotein structures (specifically TTAGGG repeats in humans) located at chromosomal termini. Their primary function is to act as a physical buffer and protective shield. Telomeres are coated by the Shelterin complex (comprising six proteins: TRF1, TRF2, RAP1, TIN2, TPP1, and POT1). This complex remodels the DNA into a T-loop, where the single stranded 3′ overhang tucks into the double-stranded region. This looping mechanism hides the chromosome ends from the cell’s DNA damage response (DDR). Without this cap, the cell would mistakenly identify the natural end as a double-strand break, leading to lethal end-to-end fusions or degradation.
- Telomerase: Due to the end-replication problem, DNA polymerase cannot fully copy the extreme tips of linear chromosomes, causing telomeres to shorten with every cell division. Telomerase is a specialized ribonucleoprotein that counteracts this attrition. It uses its own internal RNA template (TERC) to synthesize and add telomeric repeats back onto the 3′ end.
Genome Instability as a Hallmark of Cancer
Genome instability is an enabling characteristic of cancer defined by a mutator phenotype that accelerates the acquisition of hallmarks like metastasis and drug resistance. It arises from the failure of DNA repair pathways, such as mismatch repair or homologous recombination and the loss of cell cycle checkpoints like p53. These defects lead to chromosomal instability (CIN), characterized by aneuploidy and structural rearrangements, and microsatellite instability (MSI). Furthermore, telomere attrition triggers breakage fusion bridge cycles, driving massive genomic reshuffling. While this genetic flux fuels clonal evolution, it also exposes vulnerabilities; therapeutic strategies like synthetic lethality exploit these deficiencies to induce selective mitotic catastrophe in malignant cells.
Therapeutic Exploitation: Synthetic Lethality and PARP Inhibitors
The therapeutic application of synthetic lethality exploits the inherent vulnerabilities created by genome instability. Cells possess redundant DNA repair pathways to maintain genomic integrity. Poly (ADP-ribose) polymerase (PARP) facilitates the repair of single-strand breaks, while the BRCA1/2 proteins govern high-fidelity homologous recombination (HR) for double-strand breaks. In malignancies harbouring BRCA mutations, the HR pathway is functionally absent.
By introducing PARP inhibitors, clinicians pharmacologically ablate the secondary repair mechanism. This leads to the conversion of routine single-strand nicks into lethal, unrepairable double-strand breaks during DNA replication. While healthy cells remain viable due to their intact HR machinery, the mutated cancer cells succumb to mitotic catastrophe and apoptosis. This strategy represents a paradigm shift in precision oncology, transforming a driver of tumour evolution into a target for selective cytotoxicity.
Conclusion
Genome stability represents the essential biological frontier between cellular homeostasis and disease. The intricate interplay between DNA Damage Response (DDR) signalling, high-fidelity repair mechanisms like Homologous Recombination, and the protective buffering of telomeres ensures the faithful transmission of life’s blueprint. While healthy cells rely on this stability to prevent premature aging and organ failure, cancer cells exploit its absence to fuel their evolutionary trajectory.
This mutator phenotype allows tumours to bypass physiological constraints, yet it simultaneously creates unique molecular dependencies. By mapping these defects, modern medicine has moved toward precision oncology, using strategies like synthetic lethality to transform genomic chaos into a therapeutic target. Ultimately, understanding the balance between stability and instability is not only fundamental to biology but is the key to unlocking future curative strategies that turn a tumour’s primary engine of growth into its ultimate cause of destruction.
References
- Vijg, J., & Montagna, C. (2017). Genome instability and aging: Cause or effect? Translational Medicine of Aging, 1, 5–11. https://doi.org/10.1016/j.tma.2017.09.003
- Veschetti, L., Treccani, M., De Tomi, E., & Malerba, G. (2023). Genomic instability evolutionary footprints on human health: Driving forces or side effects? International Journal of Molecular Sciences, 24(14), 11437. https://doi.org/10.3390/ijms241411437
- Guo, X., Zhu, X., & Zhou, X. (2023). Genomic instability drives tumorigenesis and metastasis and its implications for cancer therapy. Biomedicine & Pharmacotherapy, 157, 114036. https://doi.org/10.1016/j.biopha.2022.114036
- Maréchal, A., & Zou, L. (2013). DNA damage sensing by the ATM and ATR kinases. Cold Spring Harbor Perspectives in Biology, 5(9), a012716. https://doi.org/10.1101/cshperspect.a012716
- Barnum, K. J., & O’Connell, M. J. (2014). Cell cycle regulation by checkpoints. Methods in Molecular Biology, 1170, 29–40. https://doi.org/10.1007/978-1-4939-0888-2_2
- Mao, Z., Bozzella, M., Seluanov, A., & Gorbunova, V. (2008). Comparison of nonhomologous end joining and homologous recombination in human cells. DNA Repair, 7(10), 1765–1771. https://doi.org/10.1016/j.dnarep.2008.06.018
- Kaushal, S., & Freudenreich, C. H. (2019). The role of fork stalling and DNA structures in causing chromosome fragility. Genes, Chromosomes & Cancer, 58(5), 270–283. https://doi.org/10.1002/gcc.22721
- Deng, Y., & Chang, S. (2007). Role of telomeres and telomerase in genomic instability, senescence and cancer. Laboratory Investigation, 87(11), 1071–1076. https://doi.org/10.1038/labinvest.3700673
- Lee, J., & Pellegrini, M. V. (2024). Biochemistry, telomere and telomerase. StatPearls Publishing. https://www.ncbi.nlm.nih.gov/books/NBK576429/
- Lord, C. J., & Ashworth, A. (2017). PARP inhibitors: The first synthetic lethal targeted therapy. Science, 355(6330), 1152–1158. https://doi.org/10.1126/science.aam7344
