Restriction enzymes are “molecular scissors” that precisely cut DNA at specific nucleotide sequences. Their primary importance lies in enabling Recombinant DNA Technology, where they cut plasmids and genes to create new genetic combinations.
They are critical for DNA analysis, allowing for mapping and the detection of genetic variations (RFLP). Specialized enzymes like DpnI are essential for cleaning up reactions in site-directed mutagenesis by eliminating unwanted DNA. Dpn1 belongs to the Type II restriction enzyme class and it is the most widely used endonuclease type since it cleaves the DNA very precisely. Unlike Type I and Type III enzymes, the restriction (cutting) activity is separate from the modification (methylation) activity and the enzyme does not require ATP for cleavage. Overall, they are the foundational tools that made genetic engineering and modern biotechnology possible.
The Unique Mechanism of DpnI
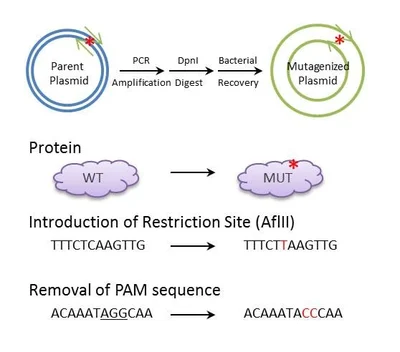
https://blog.addgene.org/site-directed-mutagenesis-by-pcr
The DpnI restriction enzyme stands apart from most conventional molecular scissors due to its strict and unusual cutting requirements. The enzyme is unique since its activity is strictly restricted to the methylation status of its DNA recognition site.
DpnI recognizes the short, four-base-pair sequence 5′-GATC-3′. However, unlike most restriction enzymes, it will only cleave the DNA if the Adenine (A) base within that sequence is modified by a methyl group (specifically, N6-methyladenine). This specific requirement allows DpnI to distinguish between different DNA molecules based on their methylation history.
As mentioned before, Dpn1 RE belongs to the Type II restriction endonucleases meaning it hydrolyzes the phosphodiester bond within the DNA backbone. The cleavage of the double helix at the recognition site is particularized specifically on the 5′ end of the Am (methylated adenine) base. This cleavage action results in DNA fragments with sticky ends (a two-base 5’overhang).
The origin of Dpn1 can be traced to Streptococcus pneumoniae. The reason a bacterium possesses this highly specific, methylation-dependent enzyme is rooted in its natural defence mechanism which is the restriction-modification system.
When an invading foreign DNA from bacteriophages (methylated) enters the host i.e. S. pneumoniae (unmethylated) The restriction enzyme cuts and destroys this foreign DNA. The DpnI-producing strain protects itself by not methylating its DNA, allowing DpnI to destroy any incoming DNA that is methylated.
The S. pneumoniae strain that produces DpnI typically lacks methylase (DpnM). Therefore, its own DNA is unmethylated and is preserved. If the strain takes up DNA from a rival strain that does have methylase (DpnM), that foreign DNA is methylated. DpnI then destroys the methylated foreign DNA.
The Cornerstone Application of DpnI: Site-Directed Mutagenesis
Site-Directed Mutagenesis (SDM) is a fundamental and precise technique in molecular genetics that grants researchers the ability to make predetermined, specific alterations to the sequence of a nucleic acid (DNA or RNA). Its defining characteristic is the capacity to introduce a targeted mutation, most commonly the substitution of a single base pair within a gene. Following the in vitro synthesis phase of PCR-based Site-Directed Mutagenesis (SDM), the resulting reaction mixture presents a significant challenge that is an undesirable blend of two distinct DNA populations that contains the original parental template DNA and the newly synthesized, mutated DNA. The original, parental template DNA is typically intact, closed, and highly efficient at replicating in the host bacteria. The parental template is usually present in much greater quantity than the newly synthesized mutant product, this leads to the overwhelming growth of background colonies which are bacterial clones that contain the original, unwanted, non-mutated plasmid.
This high background colony severely dilutes the desired mutant colonies, making the identification and isolation of a successful mutant clone extremely time consuming and often impractical. Hence for this reason a mechanism is required to selectively eliminate this abundant methylated plasmid DNA template.
The unique, methylation-dependent activity of DpnI makes it an indispensable clean-up enzyme. DpnI is added directly to the post-PCR reaction mixture to execute a highly selective digestion. This action is based entirely on its stringent requirement to only cut methylated DNA. It specifically recognizes and hydrolyses the unwanted parental template DNA because it is methylated at its GATC sites (Standard laboratory strain of E. coli that carries the active dam gene.).
Conversely, the desired newly synthesized, mutated DNA is unmethylated (since the synthesis occurred in vitro). DpnI ignores this unmethylated product, leaving it perfectly intact and ready for the next step of the protocol.
By degrading this background, DpnI ensures that when the unmethylated DNA is transformed into E. coli, the vast majority of resulting colonies contain the correct, mutated plasmid.
This transformation eliminates the need for labor-intensive, time-consuming, and expensive screening (like sequencing many colonies) to find a single mutant. DpnI effectively transforms a potentially low-yield reaction into a high-throughput, reliable technique.
Other Applications and Considerations of DpnI
- Methylation Detection: DpnI serves as a highly specific analytical tool to directly determine if the GATC sequence in a given DNA sample is methylated.
- Plasmid Preparation: Historically, DpnI was sometimes used to help purify plasmid DNA from contaminating bacterial genomic DNA.
- Validation: It’s often used to validate the activity of a methylase enzyme in vitro or to verify that a plasmid has been grown in a dam+ versus a dam- strain
- Contaminant Removal: It can be used to eliminate traces of methylated bacterial genomic DNA from plasmid preparations, or to specifically verify that a prepared plasmid is indeed methylated.
Future Outlook of DpnI
While the primary current application of DpnI is its indispensable role in Site-Directed Mutagenesis (SDM), its unique and highly specific methylation-dependent cleavage property suggests potential future applications, particularly in advanced genomic research and diagnostics.
The ability of DpnI to distinguish between methylated and unmethylated GATC sites makes it a valuable enzyme for epigenetic studies, especially when integrated into sequencing workflows.
In methods involving simultaneous assembly of multiple DNA fragments (like Gibson Assembly or Golden Gate Assembly), DpnI could be engineered into specifically eliminate template DNA used in initial PCR steps, reducing background from complex starting materials. The specificity of DpnI could be harnessed in diagnostics where the methylation status of a pathogen or a specific gene is a key indicator of a pathogen’s genome has a unique or distinct methylation pattern at GATC sites compared to the host (due to different Dam methylases), DpnI could be part of a diagnostic panel to selectively amplify or destroy pathogen DNA, leading to faster and clearer detection.
References
- Primrose, S. B., Twyman, R., & Old, R. (2002). Principles of gene manipulation (6th ed.). Wiley.
- Czapinska, H., Zaremba, M., Czapinski, J., Bochtler, M., & Bujnicki, J. M. (2012). Crystal structure and mechanism of action of the N6-methyladenine-dependent type IIM restriction endonuclease R.DpnI. Nucleic Acids Research, 40(15), 7524–7534. https://doi.org/10.1093/nar/gks428
- Barnes HE, Liu G, Weston CQ, King P, Pham LK, Waltz S, et al. (2014) Selective Microbial Genomic DNA Isolation Using Restriction Endonucleases. PLoS ONE 9(10): e109061. https://doi.org/10.1371/journal.pone.0109061
- Enam, S. U., Cherry, J. L., Leonard, S. R., Zheludev, I. N., Lipman, D. J., & Fire, A. Z. (2023). Restriction Endonuclease-Based Modification-Dependent Enrichment (REMoDE) of DNA for Metagenomic Sequencing. Applied and Environmental Microbiology, 89(1), e01670–22. https://doi.org/10.1128/aem.01670-22
- Reece, R. J. (2013). Site-directed mutagenesis. In R. J. Reece (Ed.), Analysis of genes and genomes (pp. 395–403). Academic Press. https://doi.org/10.1016/B978-0-12-418687-3.00019-7
- Plasmids 101: Methylation and Restriction Enzymes
- Mierzejewska, K., Siwek, W., Czapinska, H., Kaus-Drobek, M., Radlinska, M., Skowronek, K., Bujnicki, J. M., Dadlez, M., & Bochtler, M. (2014). Structural basis of the methylation specificity of R.DpnI. Nucleic Acids Research, 42(13), 8745–8754. https://doi.org/10.1093/nar/gku546
