What are Caspases?
Caspases play a critical role in apoptosis and belong to a family of protease enzymes involved in apoptosis, inflammation, and other cellular processes. Structurally, caspases typically consist of three domains: the N-terminal prodomain, the large catalytic subunit (p20), and the small catalytic subunit (p10), characterized by a cysteine residue at their active site and a specificity for cleaving target proteins at specific aspartic acid residues.
- Apoptosis can be triggered by various internal and external signals, including DNA damage, cellular stress, or signals from neighboring cells. Once apoptosis is initiated, a cascade of molecular events is activated, leading to cell death.
- Caspases are key players in this process. Caspases are a family of protease enzymes that are present in cells as inactive proenzymes (procaspases). They become activated through cleavage by other caspases or proteases in response to apoptotic signals.
- There are two main types of caspases involved in apoptosis: initiator caspases and executioner caspases.
- Initiator caspases, such as caspase-8 and caspase-9, are activated in response to apoptotic signals and subsequently activate executioner caspases, such as caspase-3, caspase- 6, and caspase-7.
- Executioner caspases then cleave a wide range of cellular substrates, leading to the dismantling of the cell. The activation of caspases is tightly regulated to ensure that apoptosis occurs only when necessary and to prevent unwanted cell death.
- Dysregulation of caspase activity can contribute to various diseases, including cancer, neurodegenerative disorders, and autoimmune diseases. Therefore, understanding the role of caspases in apoptosis is crucial for developing therapeutic strategies targeting these pathways.
- The activation of initiator caspases typically leads to the cleavage and activation of effector caspases, which in turn execute the apoptotic program by cleaving specific target proteins within the cell. This sequential activation and cleavage of caspases ensure the orderly progression of apoptosis and help maintain tissue homeostasis.
c-FLIP (Cellular FLICE-inhibitory protein)
Cellular FLICE (FADD-like IL-1β-converting enzyme)-inhibitory protein, also known as “cFLIP” protein, is a significant contributor in controlling apoptosis, or programmed cell death, in cells.
- It is involved in the regulation of pathways leading to cell death, especially those that are mediated by death receptors such as TNF and Fas. cFLIP isoforms are formed by alternative splicing of the cFLIP gene, resulting in the long (cFLIPl) and short (cFLIPs) kinds.13 different splice variants exist for cFLIP, which is also referred to as Casper, iFLICE, FLAME-1, CASH, CLARP, MRIT, or usurpin.
- Different impacts of these isoforms on the control of apoptosis may occur. Depending on the species and isoform, cFLIP proteins have different molecular weights. The molecular weight of cFLIPl is roughly 55 kDa, but the molecular weight of cFLIPs is about 26 kDa. Twenty amino acids appear to be essential for ubiquitinating and targeting the two DEDs of cFLIPs for proteasomal destruction.
- Though it lacks a functional caspase domain, the C- terminus of cFLIPl is longer than that of cFLIPs and resembles the structures of caspases-8 and -10.
- A number of amino acid changes have resulted in this lack of caspase function, most notably the loss of the critical cysteine residue in the catalytic domain, which is required for caspase catalytic action. Furthermore, cFLIPl has a caspase-8 cleavage site located at Asp- 376 (LEVD); cleavage of cFLIPl at this location results in the proteolytic fragment variation p43 cFLIP. Inhibiting the activation of caspase-8, a crucial enzyme involved in the initiation of apoptosis, is one of cFLIP’s main roles.
- cFLIP blocks the activation of caspase-8 by binding to it, which stops the apoptotic signaling cascade from happening. Typically, the protein has several functional domains, such as caspase-like domains and death effector domains (DEDs).
- For the purpose of facilitating interactions with other proteins involved in apoptotic signaling cascades, these domains are essential. Usually, it has two DED domains that interact with other proteins, like caspase-8, that are part of the apoptotic signaling cascade. An essential caspase-like domain, which physically resembles the caspase active site, is also present in the cFLIP structure. This domain binds to caspase-8 and inhibits its catalytic activity, preventing its activation, which is essential for cFLIP’s inhibitory effect
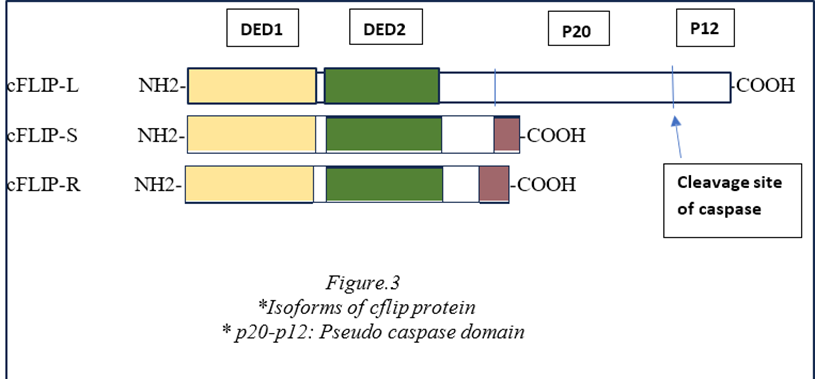
cFLIP (cellular FLICE-like inhibitory protein) has three main isoforms: cFLIP(L), cFLIP(S), and cFLIP(R). These isoforms are produced through alternative splicing and play roles in regulating apoptosis. cFLIP(L) can both inhibit and promote apoptosis, cFLIP(S) predominantly inhibits apoptosis, and cFLIP(R) is less well-characterized but is believed to have regulatory functions in cell death pathways. The p20 and p12 pseudo-caspase domains are parts of the caspase-8 enzyme, involved in apoptosis. These domains lack key catalytic residues, rendering them inactive. However, they still play essential roles in protein-protein interactions within apoptotic signaling pathways, contributing to the regulatory functions of caspase-8 in cell death processes.
Structural Similarities of Procaspase-8 & cFLIP
- The domains of procaspase-8 and cellular FLICE (FADD-like IL-1β-converting enzyme)- inhibitory protein (cFLIP) are structurally similar, especially the caspase-like and death effector domains (DEDs).
- DEDs are protein interaction domains that are involved in the formation of the Death-Inducing Signaling Complex (DISC) and are present in both ProCaspase-8 and cFLIP.
- During DISC formation, DEDs help procaspase-8/cFLIP engage with adaptor proteins, including Fas-associated death domain protein (FADD), through protein-protein interactions.
- Because of their structural similarity and shared sequence homology, the DEDs of ProCaspase- 8 and cFLIP can interact with homologous binding partners within the DISC. Caspase-like domains, which structurally mimic caspases’ active site, are present in both ProCaspase-8 and cFLIP.
- Although the active caspase-8 enzyme is produced by autocatalytic cleavage of the procaspase- like domain of ProCaspase-8, the caspase-like domain of cFLIP is devoid of essential catalytic residues required for protease activity.
- Because of its structural similarity to ProCaspase-8’s caspase-like domain, cFLIP can suppress caspase-8’s activation within the DISC in a competitive manner, even in the absence of enzymatic activity. Given that cFLIP inhibits the extrinsic apoptotic pathway, the structural similarities between ProCaspase-8 and cFLIP may have significant implications for the regulation of apoptosis.
- In the competition for binding to FADD and other elements of the Death-Inducing Signaling Complex (DISC), cFLIP faces up against ProCaspase-8. ProCaspase-8’s catalytic activity is absent from cFLIP, so its binding to the DISC stops procaspase-8 from activating and starting apoptosis.
- Competitive inhibition hinders the progression of apoptosis by efficiently blocking the apoptotic signal at an early stage. Because ProCaspase-8’s and cFLIP’s caspase-like domains are structurally similar, cFLIP can connect to ProCaspase-8 within the DISC. cFLIP stops autocatalytic cleavage, which is necessary for procaspase-8 to be activated into active caspase- 8, by binding to the active site of ProCaspase-8.
- Consequently, the reduction of apoptosis occurs as a result of the inhibition of the downstream apoptotic cascade, which is dependent on caspase-8 activation. When cFLIP is overexpressed or more abundant than ProCaspase-8, its inhibitory action may be predominant, resulting in longer cell survival.
- Even in the presence of death ligands, which ordinarily cause apoptosis through the extrinsic pathway, this life extension is possible. As a result, cells may resist death signals and carry on multiplying, which aids in the development of tumors and other pathological disorders.
References
- Pfeffer, C. M., & Singh, A. T. K. (2018, February 2). Apoptosis: A Target for Anticancer Therapy. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms19020448
- Hanahan, D., & Weinberg, R. A. (2011). Hallmarks of cancer: The next generation. Cell, 144(5), 646-674. https://doi.org/10.1016/j.cell.2011.02.013
- Riley, J., Malik, A., Holohan, C. et al. DED or alive: assembly and regulation of the death effector domain complexes. Cell Death Dis 6, e1866 (2015). https://doi.org/10.1038/cddis.2015.213
- Cui, Z., Dabas, H., Leonard, B.C. et al. Caspase-8 mutations associated with head and neck cancer differentially retain functional properties related to TRAIL-induced apoptosis and cytokine induction. Cell Death Dis 12, 775 (2021). https://doi.org/10.1038/s41419-021-04066-z
- Park, H.H. Molecular basis of dimerization of initiator caspase was revealed by crystal structure of caspase-8 pro-domain. Cell Death Differ 26, 1213–1220 (2019). https://doi.org/10.1038/s41418-018-0200-x
- Stöhr, D., Jeltsch, A., & Rehm, M. (2020). TRAIL receptor signaling: From the basics of canonical signal transduction toward its entanglement with ER stress and the unfolded protein response. In L. M. Glover & V. Nisenblat (Eds.), Advances in Clinical Chemistry (Vol. 95, pp. 63-104). Elsevier. https://doi.org/10.1016/bs.ircmb.2020.02.002
- Safa AR. c-FLIP, a master anti-apoptotic regulator. Exp Oncol. 2012 Oct;34(3):176-84. PMID: 23070002; PMCID: PMC4817998.
- Zhang, M., Abrams, C., Wang, L., Gizzi, A., He, L., Lin, R., Chen, Y., Loll, P. J., Pascal, J. M., & Zhang, J.-f. (2012). Structural basis for calmodulin as a dynamic calcium sensor. Structure, 20(5), 911-923. https://doi.org/10.1016/j.str.2012.03.019
- Gaidos G, Panaitiu AE, Guo B, Pellegrini M, Mierke DF. Identification and Characterization of the Interaction Site between cFLIPL and Calmodulin. PLoS One. 2015 Nov 3;10(11):e0141692. doi: 10.1371/journal.pone.0141692. PMID: 26529318; PMCID: PMC4631386.
- Yu, J. W., & Shi, Y. (2008). FLIP and the death effector domain family. *Oncogene, 27*(47), 6216–6227. https://doi.org/10.1038/onc.2008.302
