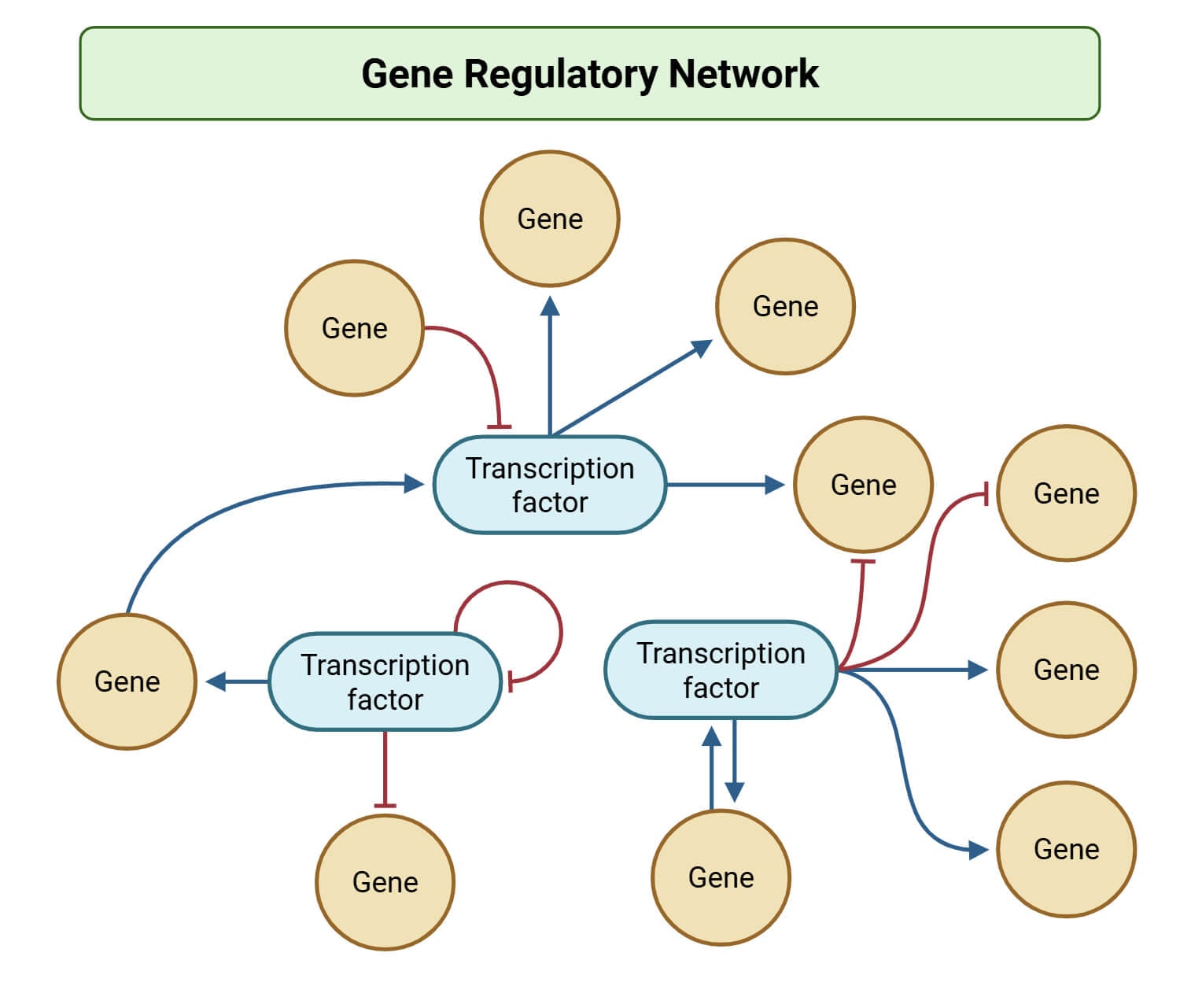
Gene regulatory elements function as the primary architectural framework for the spatiotemporal orchestration of the transcriptome.
These non-coding DNA sequences classified as promoters, enhancers, silencers, and insulators serve as high-affinity binding sites for trans-acting transcription factors and co-regulators. Unlike the static protein-coding regions of the genome, these elements act as dynamic biochemical switches that integrate extracellular signals and internal developmental cues to modulate the rate of transcriptional initiation.

While proximal promoters facilitate the assembly of the pre-initiation complex, distal enhancers can influence gene expression across vast genomic intervals. This long-range regulation is achieved through three-dimensional chromatin looping, where the physical folding of the DNA molecule brings distant regulatory modules into direct contact with their target promoters. This sophisticated regulatory landscape is fundamental to cellular differentiation, allowing identical genotypes to yield diverse phenotypic lineages.
The Promoter Region: Definition and Core Functions
The promoter region is a critical regulatory DNA sequence located upstream (5′) of a gene’s coding region that serves as the primary command centre for initiating transcription. Its fundamental role is to provide a physical docking platform for RNA polymerase and the specialized proteins, known as transcription factors, required to catalyze the synthesis of RNA.
The promoter serves as the fundamental regulatory gateway for gene expression by orchestrating four critical processes: site selection, which precisely locates the transcription start site; molecular assembly, which facilitates the formation of the pre-initiation complex; directional orientation, which ensures the correct 5′ to 3′ synthesis of RNA; and transcriptional gating, which modulates the frequency of mRNA production in response to cellular signals. Collectively, these functions ensure that the genetic blueprint is accessed with high fidelity and adapted to the specific physiological requirements of the cell.
Key components of Promoters: The TATA box and initiator sequences
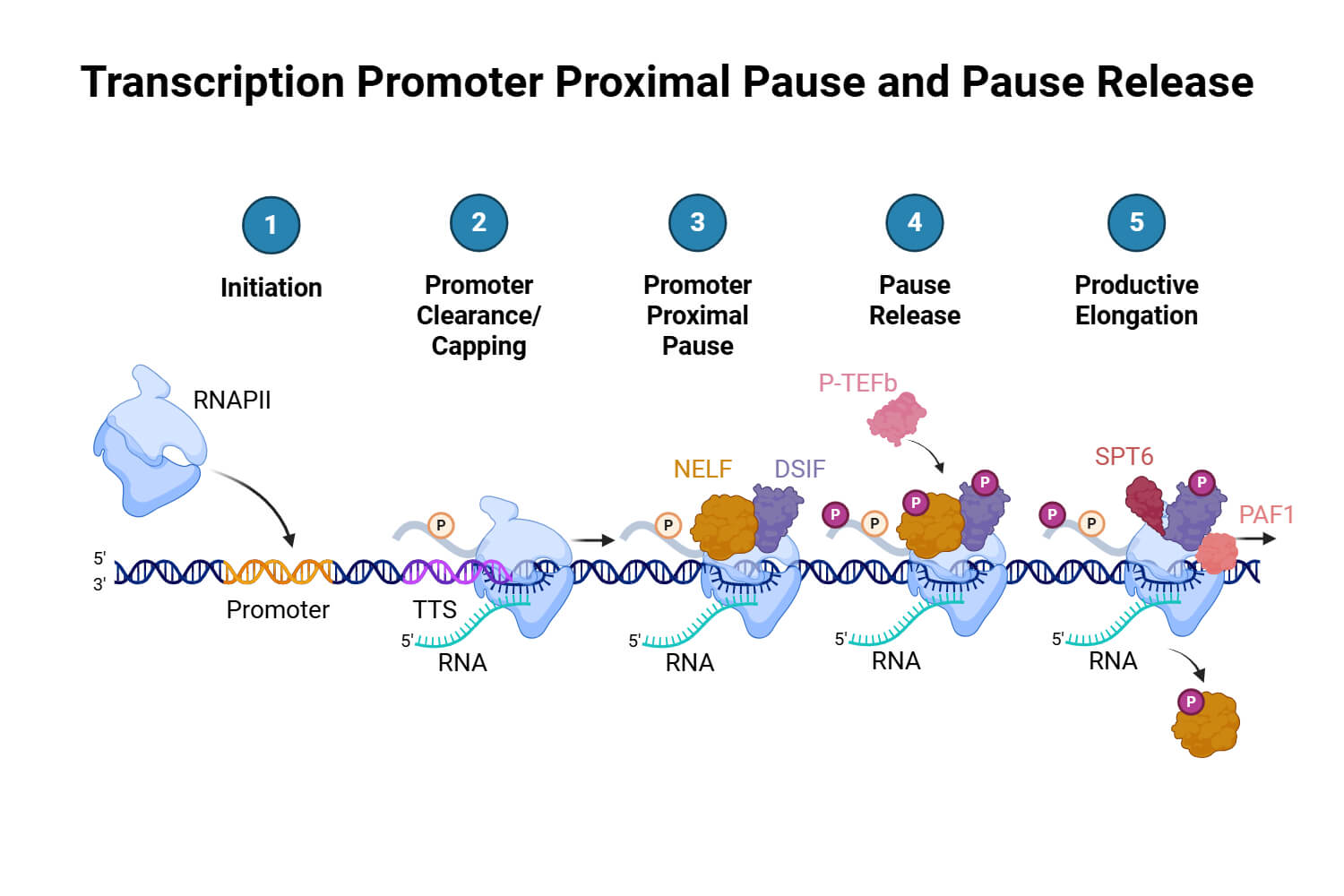
In eukaryotic transcription, the core promoter serves as the foundational regulatory scaffold for the assembly of the pre-initiation complex. This specialized DNA region directs RNA polymerase II to its correct genomic coordinates, ensuring the high-fidelity initiation of gene expression. Two of the most critical cis-regulatory elements within this architecture are the TATA box and the Initiator (Inr) sequence, which act as biochemical anchors for the transcriptional machinery.
The TATA box is a highly conserved, AT-rich motif typically located 25 to 31 base pairs upstream of the transcription start site. Its primary function is to serve as a high-affinity docking site for the TATA-binding protein (TBP). Upon binding, TBP induces a sharp, approximately 80-degree bend in the DNA, physically deforming the double helix to facilitate the recruitment of general transcription factors. Complementing this, the Initiator sequence directly encompasses the start site (+1), spanning from roughly -2 to +4. Recognized by subunits of the TFIID complex, the Inr provides the precise spatial coordinates required for the first nucleotide of RNA synthesis. While the TATA box nucleates structural assembly, the Inr is essential for accurate start-site selection, particularly in TATA-less promoters, ensuring the reliable execution of the genetic program.
Proximal vs. Core Promoters: Understanding the Difference
The transcriptional framework is defined by a distinct structural hierarchy, bifurcating the promoter into two primary functional domains: the core and the proximal regions.
The core promoter serves as the fundamental biochemical scaffold, typically spanning 40 base pairs upstream and downstream of the transcription start site. It houses essential motifs including the TATA box and the Initiator (Inr) which facilitate the direct recruitment of RNA polymerase II and the assembly of the pre-initiation complex. This region is primarily responsible for the mechanical execution of transcription, ensuring the holoenzyme Complex is correctly positioned to begin RNA synthesis at the precise +1 nucleotide.
Situated immediately upstream, the proximal promoter extends roughly 250 base pairs from the start site and functions as a sophisticated regulatory convergence Point. Unlike the relatively universal components of the core promoter, this region contains diverse cis-acting elements, such as GC-rich boxes and CAAT motifs, which bind sequence-specific transcription factors. These proteins modulate the efficiency of the core machinery, either accelerating or downregulating the rate of transcript initiation in response to cellular stimuli. Ultimately, while the core promoter provides the essential ignition for gene expression, the proximal promoter operates as a specialized control interface, mediating the intensity and timing of the transcriptional response.
What are Enhancers? Distal Regulation of Gene Expression
Enhancers represent a Complex Regulatory Modality of distal cis-regulatory elements that orchestrate the complex Dynamic Expression Profiles. Unlike proximal promoter elements, which are constrained by their orientation and immediate proximity to the transcription start site, enhancers exhibit unique position-independence. They can exert regulatory influence from hundreds of kilobases away, residing upstream, downstream, or even within the introns of the genes they regulate. These sequences serve as specialized docking platforms for sequence-specific transcription factors and co-activators, which integrate multifaceted cellular signals to amplify the transcriptional output.
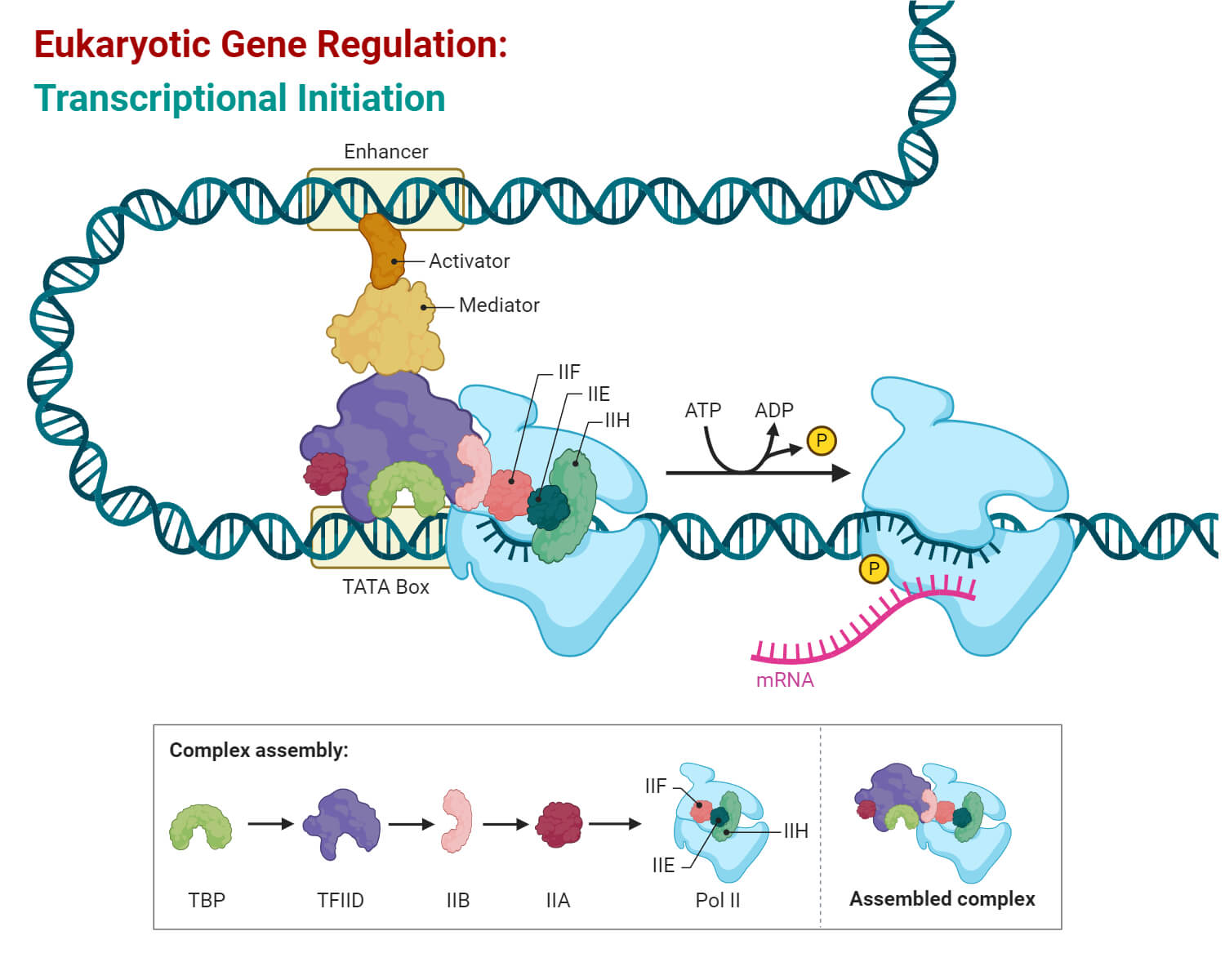
Enhancer activity relies on a physical process called chromatin looping. To overcome the vast distance between an enhancer and a gene, specialized architectural proteins such as cohesin, CTCF, and the Mediator complex bend the DNA strand into a loop. This structural shift bridges the gap, bringing the distant enhancer into direct physical contact with the gene’s core promoter.
This contact creates a crowded workspace or microenvironment where the necessary cellular machinery is concentrated. By stabilizing the basal transcription apparatus and pulling in RNA polymerase II, the enhancer acts like a power booster that increases the frequency of transcriptional bursting. Ultimately, these enhancers function as central regulatory nexuses, allowing the genome to follow specific instructions that ensure genes are turned on at exactly the right time and in the right tissue.
Promoters vs. Enhancers: Key Differences and Similarities
To elucidate the interplay between these two genomic features, it is helpful to view them as a hierarchy of control. Although they converge on the singular objective of modulating gene expression, they remain fundamentally distinguished by their biochemical signatures and their specialized mechanistic roles.
| Feature | Promoter | Enhancer |
| Primary Function | Initiation: Defines the exact site where transcription begins. | Amplification: Boosts the frequency and intensity of transcription. |
| Location | Fixed. typically, within 100–1000 bp of the Transcription Start Site (TSS). | Flexible; can be located up to 1 Mb away (upstream, downstream, or intronic). |
| Orientation | Strictly dependent: Must face the correct direction to drive RNA Polymerase. | Independent: Functional in both forward and reverse orientations. |
| RNA Product | Produces stable, polyadenylated mRNA. | Produces short-lived, non-coding eRNA (enhancer RNA). |
| Sequence Motif Composition | Contains GC-rich motifs and “fixed” elements like the TATA box. | Contains diverse, tissue-specific transcription factor binding sites. |
| Protein Binding | Binds General Transcription Factors (GTFs) and RNA Polymerase II. | Binds Sequence-specific Activators and tissue-specific factors. |
Mechanism of Action: DNA Looping and Mediator Complexes
The regulation of gene expression relies on a spatial mechanism known as DNA looping, which allows distal regulatory elements called enhancers to physically contact target promoters located thousands of base pairs away. This process is orchestrated by the cooperative dynamics of the Mediator complex and the Cohesin ring, which together reorganize chromatin to facilitate gene activation. Functioning as a modular coactivator scaffold, Mediator utilizes its Tail module to recruit sequence-specific transcription factors at the enhancer, while its Head and Middle modules anchor RNA Polymerase II and the Pre-Initiation Complex at the promoter.
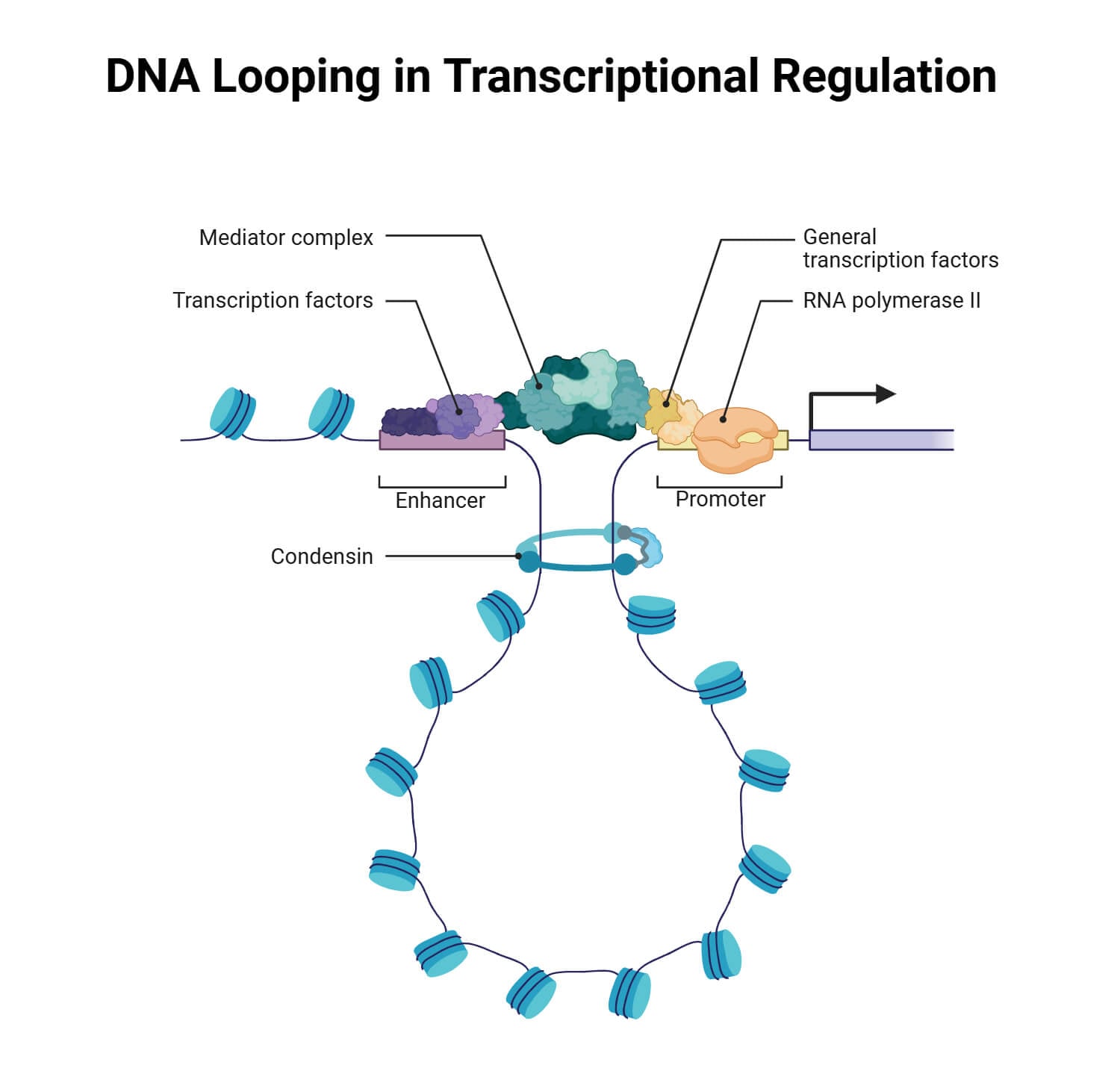
This biochemical bridge is structurally reinforced by Cohesin-mediated loop extrusion, a process where the ring-shaped complex utilizes ATP-dependent motor activity to thread chromatin through its lumen. The extrusion is halted when Cohesin is tethered by the Mediator complex or architectural proteins like CTCF, effectively stapling the genomic elements into a stable transcriptional hub. By creating this localized microenvironment, the cell ensures high concentrations of transcriptional machinery are available for precise, lineage-specific gene activation.
DNA looping is a 4-step process, involving mechanical forces and biochemical anchors.
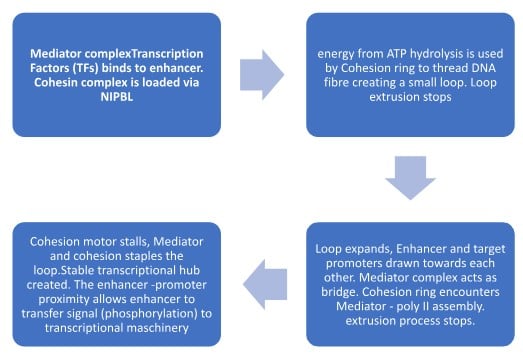
| Step | Phase | Key Component | Action |
| 1 | Loading | NIPBL & Cohesin | Cohesin ring is placed onto the DNA. |
| 2 | Extrusion | Cohesin + ATP | DNA is reeled through the ring to form a loop. |
| 3 | Bridging | Mediator Complex | Enhancer and Promoter are linked by the Mediator scaffold. |
| 4 | Capture | Mediator & Cohesin | Cohesin is stalled, stapling the loop for transcription. |
Transcription Factors: The Bridge Between Enhancers and Promoters
The transition from a silent genetic state to active transcription is governed by the hierarchical interaction between two classes of regulatory proteins: pioneer transcription factors and standard transcription factors.
Pioneer factors act as the primary initiators of gene activation due to their unique biochemical capacity to engage DNA motifs embedded within highly condensed heterochromatin. Unlike standard factors, which are sterically hindered by nucleosomes, pioneer factors recognize their cognate sequences on the surface of histone wrapped DNA. Once bound, they serve as molecular anchors, recruiting chromatin remodelers and histone modifiers that displace or rearrange nucleosomes to create an accessible open state.
Only after this structural barrier is removed can standard transcription factors, such as activators and the Mediator complex, bind to the newly exposed enhancers. While pioneer factors establish the competence of a gene to be expressed, essentially unlocking the DNA standard factors execute the final activation by recruiting RNA Polymerase II. Pioneer factors define a cell’s potential identity by selecting which genomic regions can be opened, while standard factors respond to immediate environmental cues to drive the actual synthesis of mRNA.
Super-Enhancers: Controlling Cell Identity and Disease
Super-enhancers (SEs) are high-order regulatory domains composed of dense clusters of individual enhancers that act in concert to drive robust, lineage-specific transcription. Unlike typical enhancers, SEs are defined by an exceptional accumulation of master transcription factors, the Mediator complex, and active chromatin signatures such as H3K27ac. These domains serve as the primary coordinators of cellular identity, ensuring that genes essential for a cell’s unique function and developmental fate are expressed at maximum capacity.
The exceptional potency of SEs is increasingly explained by the phase separation model. In this framework, the high concentration of proteins containing intrinsically disordered regions (IDRs) specifically within Mediator and transcription factors facilitates the formation of transcriptional condensates. These liquid-like droplets compartmentalize the genome, concentrating the necessary enzymes and RNA Polymerase II to facilitate high-frequency transcriptional bursting.
In clinical settings, SEs represent critical vulnerability cancer cells frequently hijack this machinery by creating de novo super-enhancers at oncogenes through genomic rearrangements or focal amplifications. This transcriptional addiction makes SE-associated cofactors, such as BRD4 and CDK7, prime targets for small-molecule inhibitors designed to collapse these oncogenic hubs and restore normal gene regulation.
Methods to Study Regulatory Elements: ChIP-Seq and Reporter Assays
To decode the intricate regulatory language of the genome, two primary complementary methodologies are used: correlative mapping (identifying physical presence) and functional validation (testing actual biological impact). ChIP-Seq and Reporter Assays serve as the cornerstone technologies for these distinct approaches.
ChIP-Seq
Chromatin Immunoprecipitation Sequencing (ChIP-Seq) is the standard correlative method used to chart the occupancy of proteins across the entire genome. It utilizes specific antibodies to isolate DNA fragments physically bound by target proteins, such as transcription factors or modified histones (e.g., H3K27ac for active enhancers). It provides a comprehensive, genome-wide atlas of potential regulatory elements in their native chromatin context. ChIP-Seq can capture tethered interactions where a protein is not bound to the DNA directly but is instead part of a large multi-protein complex that is cross-linked to the site.
Reporter Assays
Reporter Assays act as the functional counterpart, moving beyond mapping to measure a DNA sequence’s actual ability to drive gene expression. A sequence of interest is cloned into a plasmid (an episomal DNA vector) upstream of a reporter gene, such as Luciferase or GFP. This construct is then introduced into living cells.If the cell produces light or fluorescence, it confirms the sequence is a functional regulatory element. This method is essential for verifying hits from ChIP-Seq and for dissecting the specific nucleotides required for an enhancer’s activity through site-directed mutagenesis.
Conclusion
The eukaryotic genome operates through a rigorous structural hierarchy where core and proximal promoters provide the mechanical ignition for transcription, while distal enhancers modulate its intensity. This spatial coordination is facilitated by the Cohesin-Mediator axis, which utilizes ATP-dependent loop extrusion to transition the genome from a linear sequence into a functional 3D architecture.
By integrating the unique chromatin-penetrating capabilities of pioneer factors with the concentrated potency of super-enhancer condensates, the cell maintains a precise transcriptomic profile. The synergy between correlative ChIP-Seq data and functional reporter assays remains the gold standard for dissecting these interactions, revealing how biochemical switches translate a singular genotype into diverse cellular phenotypes.
References
- Doane, A. S., & Elemento, O. (2017). Regulatory elements in molecular networks. WIREs Systems Biology and Medicine, 9(3), Article e1374. https://doi.org/10.1002/wsbm.1374
- Cooper, S. J., Trinklein, N. D., Anton, E. D., Nguyen, L., & Myers, R. M. (2006). Comprehensive analysis of transcriptional promoter structure and function in 1% of the human genome. Genome Research, 16(1), 1–10. https://doi.org/10.1101/gr.4222606
- Dao, L. T. M., & Spicuglia, S. (2018). Transcriptional regulation by promoters with enhancer function. Transcription, 9(5), 307–314. https://doi.org/10.1080/21541264.2018.1486150
- Petrascheck, M., Escher, D., Mahmoudi, T., Verrijzer, C. P., Schaffner, W., & Barberis, A. (2005). DNA looping induced by a transcriptional enhancer in vivo. Nucleic Acids Research, 33(12), 3743–3750.
- Zhu, Q., Dabi, T., & Lamb, C. (1995). TATA box and initiator functions in the accurate transcription of a plant minimal promoter in vitro. The Plant Cell, 7(10), 1681–1689. https://doi.org/10.1105/tpc.7.10.1681
- D’Elia, B., & Fuxman Bass, J. (2025). Applications of high-throughput reporter assays to gene regulation studies. Current Opinion in Structural Biology, 94, 103105. https://doi.org/10.1016/j.sbi.2025.103105
- APA 7th Edition Citation: Whyte, W. A., Orlando, D. A., Hnisz, D., Abraham, B. J., Lin, C. Y., Kagey, M. H., Rahl, P. B., Lee, T. I., & Young, R. A. (2013). Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell, 153(2), 307–319. https://doi.org/10.1016/j.cell.2013.03.035
- Russell, P. J. (2010). Gene expression: Transcription. In iGenetics: A Molecular Approach (3rd ed., pp. 83–110). Pearson Benjamin Cummings.
