DNA is often described as a double helix, resembling a twisted ladder with each strand made of repeating units called nucleotides. A nucleotide consists of a phosphate group, a deoxyribose sugar, and a nitrogenous base (A, T, C, or G). Each strand has a direction, determined by the carbon atoms in the sugar molecule.

One end is called the 5′ (five-prime) end (finishing with a phosphate group), and the other is the 3′ (three-prime) end (finishing with a hydroxyl group). In the double helix, the two strands run in opposite directions. If one strand is oriented 5’→3′, its partner strand is oriented 3′ → 5’this is termed as the antiparallel structure of DNA.
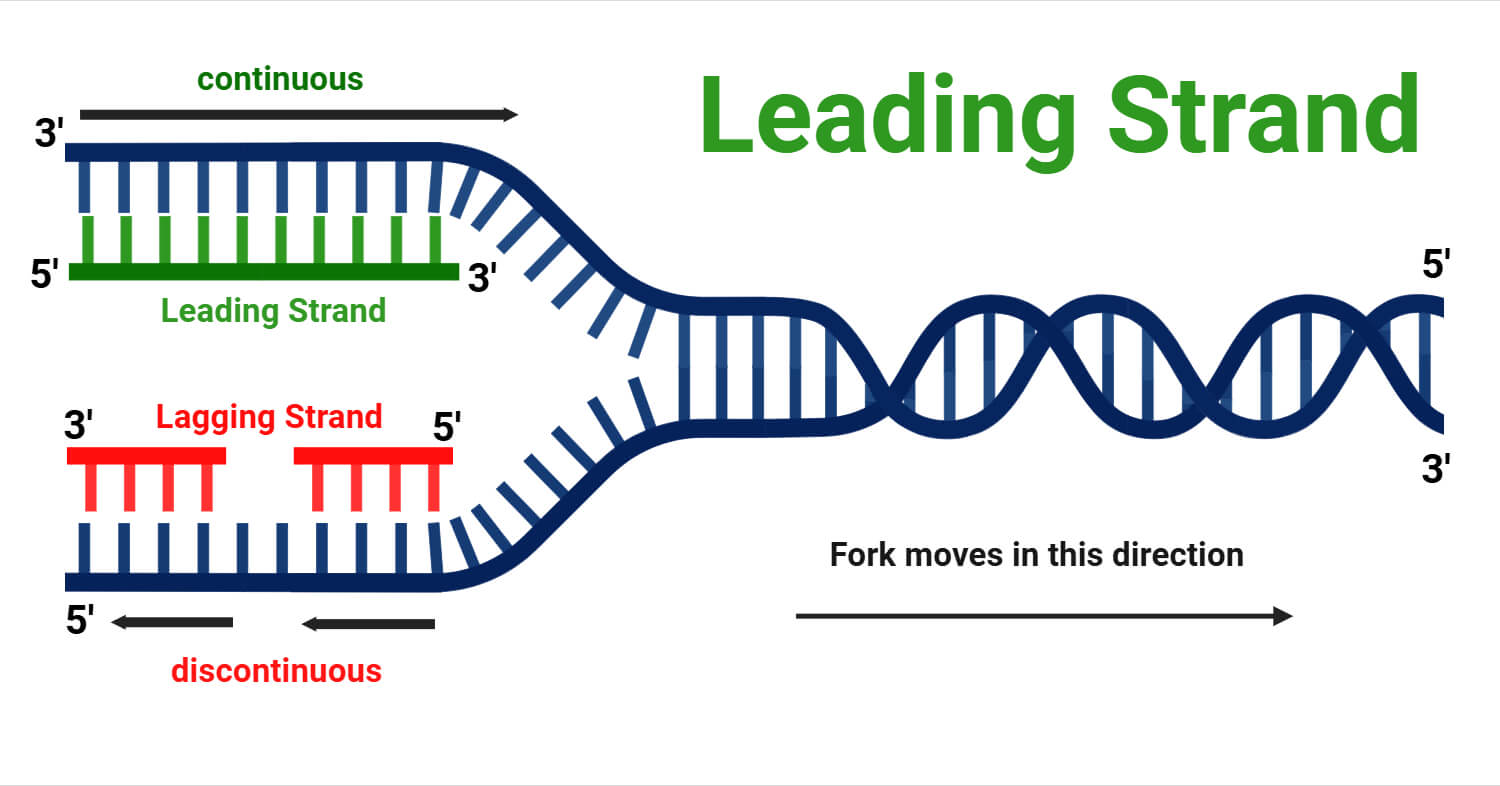
Defining from the structural perspective of DNA, a leading strand is the newly synthesized DNA strand that is built continuously in the 5’→ 3′ direction, moving in the same direction as the advancing replication fork. Using the 3’→ 5′ parental strand as its template because the new strand must be antiparallel to the template.
The Replication Fork: How the Leading Strand is Exposed
DNA replication operates like a precision engineered assembly line. At the centre of this process is the replication fork a Y-shaped junction where the double helix is unzipped, transforming a single stable molecule into two distinct templates.
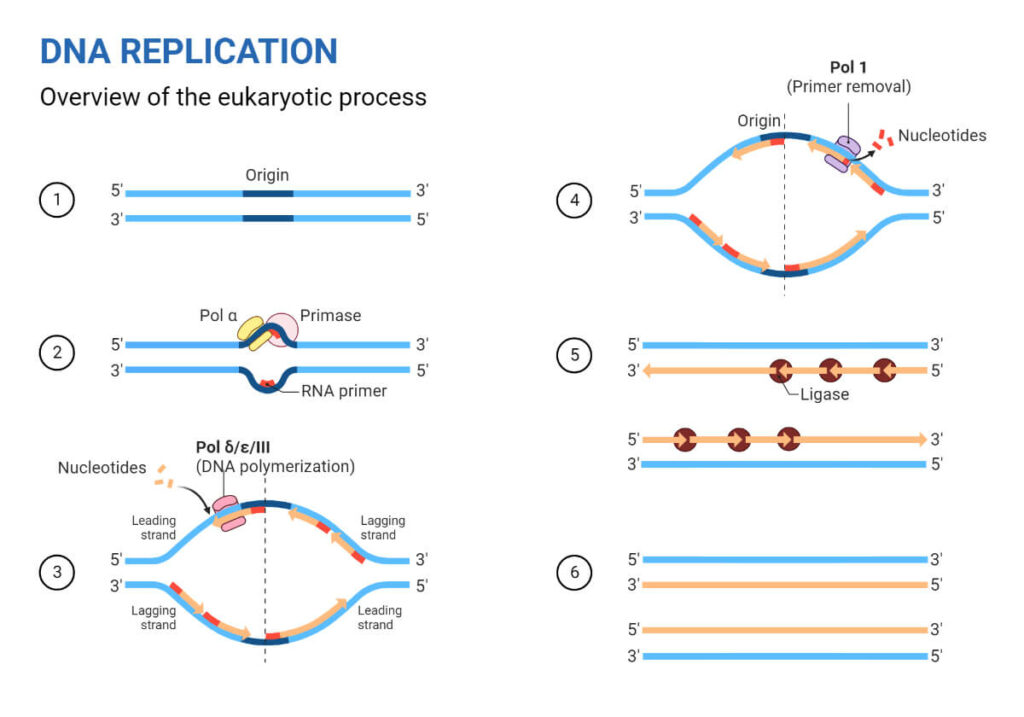
While both sides are copied simultaneously, the leading strand is the fast track of the operation. Due to its specific chemical orientation, it is exposed and synthesized in one smooth, unbroken motion, moving in perfect lockstep with the unwinding DNA.
The process is initiated by Helicase, a motor protein that uses energy to break the hydrogen bonds between nitrogenous bases. As Helicase moves forward, it creates the “V” shape of the fork, continuously peeling back the leading strand template. Once separated, the single strands are highly unstable and want to re-anneal. Single-Strand Binding (SSB) Proteins act as molecular stabilizers, coating the leading strand to keep it straight and accessible for the polymerase. The leading strand is oriented in a 3’→5′ direction toward the fork. Because DNA Polymerase III can only add nucleotides in the 5’→3′ direction, it can attach to the template and chase the helicase. This allows for continuous synthesis, requiring only one initial RNA primer to start a chain that can span thousands of base pairs without interruption.
Directionality Matters: Why Synthesis is Continuous (5′ to 3′)
DNA replication is a paragon of chemical coordination, where directionality is strictly dictated by the specific atomic structure of the deoxyribose sugar. Because DNA Polymerase can only function in a 5’→3′ direction, it requires a free 3′-hydroxyl (-OH) group to act as a chemical hook for incoming nucleotides. On the leading strand, this orientation is perfectly aligned with the mechanical movement of the replication fork. As the enzyme helicase unzips the double helix, it exposes the template strand in a 3′ →5′ orientation. The necessity of this directional movement is rooted in energy. This continuous synthesis is powered by the incoming dNTPs, which carry a high-energy triphosphate tail. When the 5′ phosphate of an incoming nucleotide attaches to the 3′-OH of the growing chain, it releases pyrophosphate, providing the fuel to forge a permanent phosphodiester bond. Once the chemical bond is forged, the replication machinery immediately locks the new nucleotide in place and prepares for the next addition. The discarded pyrophosphate is split into two individual phosphates by an enzyme pyrophosphatase, a reaction that releases a final burst of energy to ensure the synthesis is irreversible, essentially acting as a chemical lock. Simultaneously, DNA Polymerase undergoes translocation, sliding one step forward along the template to clear its active site for the next incoming base.
Key Enzymes: The Role of DNA Polymerase III and Primase
In the continuous synthesis of the leading strand, two enzymes act as the starter and the builder. While their roles are different, the work in perfect sequence to ensure the genetic code is copied without gaps.
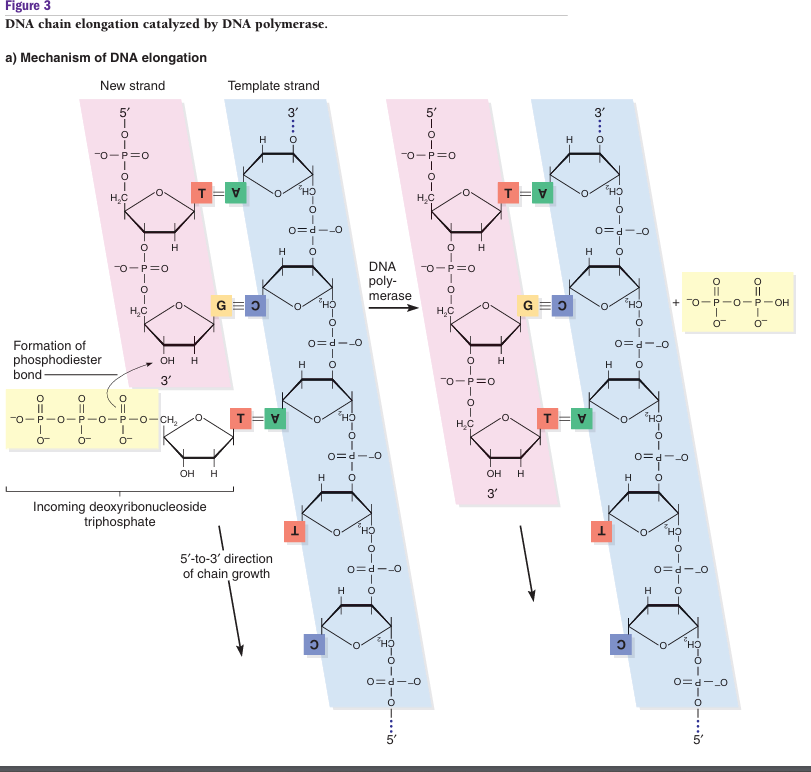
Russell, P. J. (2010). DNA replication. In iGenetics: A molecular approach (3rd ed., pp. 43–72). Pearson Benjamin Cummings.
- RNA Primase: RNA Primase also known as DNA-dependent RNA Polymerase is the essential starter enzyme of DNA replication, structurally it is a Domain-based monomer (in bacteria) or a Heterodimeric complex (in eukaryotes). Acting as a bridge between a single template strand and the main building machinery. Because DNA Polymerase is chemically incapable of starting a new chain from scratch, it relies on primase to provide a foundation i.e. it synthesizes a short stretch of RNA (about 10–12 nucleotides long) called a primer. The primary goal of this RNA primer is to provide a free 3′-OH (hydroxyl) group. This serves as the chemical hook that DNA Polymerase III needs to latch onto and begin adding DNA bases. Primase physically binds to Helicase to form a functional unit called the Primosome. This ensures that as soon as the DNA is unzipped, the template is immediately primed for copying. On the leading strand, Primase only acts once at the very beginning of the process.
- DNA Polymerase III: DNA Polymerase III is a massive, multisubunit holoenzyme that serves as the primary engine for bacterial DNA synthesis. Structurally, it is defined by its catalytic α-core, which builds the new strand, and the donut-shaped β-sliding clamp that hitches the enzyme to the DNA for extreme stability and high speed. Its primary role on the leading strand is the rapid, continuous addition of nucleotides in the 5’→3′ direction, moving in perfect synchronization with the replication fork. Beyond its role as a builder, the enzyme also acts as its own editor; the ɛ subunit provides a proofreading function that detects and removes mismatched bases, ensuring the genetic code is copied with near-perfect accuracy.
Leading Strand vs. Lagging Strand: Key Differences Explained
While both strands are synthesized at the same time and by the same molecular machinery, they are processed differently due to the vectorial nature of DNA construction. The primary difference between the leading and lagging strands lies in their directionality relative to the replication fork.
Comprehensive Comparison of Leading and Lagging Strands:
| Feature | Leading Strand | Lagging Strand |
| Orientation | The template is 3′ →5′ (relative to fork). | Template is 5′ →3′(relative to fork). |
| Synthesis Direction | 5′ →3′ (moving toward the fork). | 5′ → 3′ (moving away from the fork). |
| Continuity | Continuous: One long, unbroken chain. | Discontinuous: Short Okazaki fragments. |
| Primer Requirement | Only one RNA primer at the origin | Thousands of primers (one per fragment). |
| Main Enzyme (Prokaryotic) | DNA Polymerase III | DNA Polymerase III |
| Main Enzyme (Eukaryotic) | DNA Polymerase ɛ (Epsilon) | DNA Polymerase δ (Delta) |
| Ligase Activity | Minimal (only at the very end). | High Required to seal every fragment. |
| Looping | No looping required. | DNA is looped to fit into the polymerase. |
| Exonuclease Activity | Primarily proofreading. | High activity to remove numerous RNA primers. |
Processivity Factors: The Function of the Sliding Clamp
Processivity is a measure of an enzyme’s ability to catalyze continuous reactions without releasing its substrate. In DNA replication, this is achieved through the sliding clamp, a ring-shaped protein multimer (such as the β-clamp or PCNA) that encircles the DNA double helix. By forming a stable, toroidal structure that physically tethers the DNA polymerase to the template, the clamp prevents the enzyme from dissociating. This mechanism drastically increases replication efficiency, allowing the polymerase to add thousands of nucleotides at high speeds before falling off.
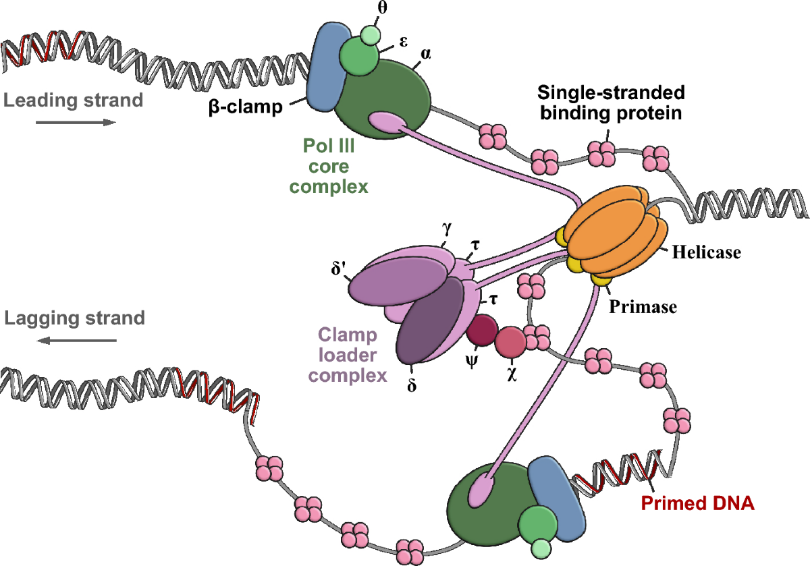
https://link.springer.com/article/10.1007/s00018-024-05252-w
In this context, the term substrate refers to the specific molecules that DNA polymerase acts upon to build the new DNA strand. There are two primary ways to look at the substrate:
- The Building Blocks (Chemical Substrates): These are deoxyribonucleoside triphosphates (dNTPs)—specifically dATP, dTTP, dCTP, and dGTP. Each dNTP provides both the monomer for the growing chain and the chemical energy (via phosphate bond hydrolysis) required to drive the reaction forward.
- The Structural Substrate: The enzyme requires a primer-template junction. This consists of a single-stranded DNA template that dictates the sequence and a short primer (RNA or DNA) that provides the necessary 3′-OH group for the polymerase to attach the next nucleotide.
Proofreading Mechanisms: Ensuring High Fidelity on the Leading Strand
High fidelity on the leading strand is maintained through a coordinated multi-step process that begins with nucleotide selectivity, where the polymerase active site acts as a geometric filter to ensure only correct Watson-Crick base pairs are incorporated. If a mismatch occurs, the enzyme utilizes its intrinsic 3′ →5′ exonuclease activity; the polymerase stalls, shifts the frayed DNA end from the polymerization site to the exonuclease site, and excises the incorrect base. This self-correcting mechanism is heavily supported by the sliding clamp, which prevents the polymerase from dissociating during the transition between synthesis and error correction. Together with post-replication mismatch repair, these mechanisms ensure an incredibly low error rate of approximately one in a billion nucleotides.
Termination of Synthesis: What Happens at the End?
The termination of DNA synthesis occurs when two converging replication forks meet and the replication machinery is dismantled. In prokaryotes, this is a highly regulated process involving specific Ter sequences and Tus proteins that trap forks in a designated terminus region, ultimately resulting in two interlinked (catenated) circular chromosomes that must be separated by topoisomerases. In eukaryotes, termination occurs more randomly across multiple linear replicons, but it faces the unique end-replication problem. Because DNA polymerase requires an RNA primer and only synthesizes in a 5′ → 3’ direction, the very end of the lagging strand cannot be fully copied, leading to progressive chromosomal shortening. To combat this, the enzyme telomerase, which is a ribonucleoprotein, adds repetitive G-rich sequences to the telomeres using its own internal RNA template. This extension provides enough space for a new primer to be laid down, allowing the lagging strand to be completed and preserving the integrity of the genetic code across cell divisions.
Clinical Significance: Targeting DNA Replication in Cancer Therapy
Targeting DNA replication is a primary strategy in cancer therapy due to the heightened replication stress and rapid division rates characteristic of malignant cells. Therapy focuses on disrupting the core machinery of the replication fork to induce genomic instability and trigger apoptosis. By inhibiting the enzymes responsible for unwinding DNA, synthesizing new strands, or relieving torsional strain, clinicians can selectively force cancer cells into a mitotic catastrophe. This approach often leverages the fact that cancer cells have compromised DNA repair pathways, making them unable to recover from stalled replication forks or DNA strand breaks that healthy cells might otherwise survive.
Clinical Concepts in Practice
- Nucleotide Mimicry: Introducing structural analogues that compete with natural dNTPs for the polymerase active site, effectively halting synthesis once incorporated into the growing strand.
- Topoisomerase Trapping: Inhibiting the enzymes that manage DNA supercoiling, which creates physical barriers that lead to permanent double-strand breaks as the replication fork attempts to pass.
- Processivity Inhibition: Disrupting the interaction between the DNA polymerase and the sliding clamp to prevent continuous synthesis, forcing the replication machinery to dissociate prematurely.
- Synthetic Lethality: Exploiting pre-existing genetic deficiencies in a tumour’s repair system by blocking a secondary, compensatory replication-restart pathway, leading to cell death.
- Replication Fork Collapse: Inducing massive DNA damage by causing the replication fork to stall and break, a state that rapidly proliferating cells cannot resolve.
Conclusion
DNA replication is a precision-engineered process where the leading strand is synthesized continuously in the 5′ →3′ direction, moving in lockstep with the unwinding helix. Facilitated by Helicase and stabilized by SSB proteins, this fast-tracks synthesis relies on DNA Polymerase III and the sliding clamp for high processivity and speed. Built-in proofreading mechanisms ensure near-perfect accuracy, while telomerase resolves end-replication issues in eukaryotes. Because malignant cells are addicted to this rapid division, targeting these replication forks through nucleotide mimicry or topoisomerase trapping is a vital strategy in modern cancer therapy to induce mitotic catastrophe.
References
- Cooper, G. M. (2000). DNA replication. In The cell: A molecular approach (2nd ed.). Sinauer Associates. https://www.ncbi.nlm.nih.gov/books/NBK9940/
- Author, A. A., & Author, B. B. 2018. DNA replication. [PDF file]. Source Name. https://www.url-to-pdf.com
- Song, H.-Y., Shen, R., Hamid, M., Guo, Y.-N., & Wang, D.-G. (2023). DNA replication: Mechanisms and therapeutic interventions for diseases. MedComm, 4(1), e210. https://doi.org/10.1002/mco2.210
- Lee, J., Chastain, P. D., II, Kusakabe, T., Griffith, J. D., & Richardson, C. C. (1998). Coordinated leading and lagging strand DNA synthesis on a minicircular template. Molecular Cell, 1(7), 1001–1010. https://doi.org/10.1016/s1097-2765(00)80100-8
- Chilkova, O., Stenlund, P., Isoz, I., Stith, C. M., Grabowski, P., Lundström, E.-B., Burgers, P. M., & Johansson, E. (2007). The eukaryotic leading and lagging strand DNA polymerases are loaded onto primer-ends via separate mechanisms but have comparable processivity in the presence of PCNA. Nucleic Acids Research, 35(19), 6588–6597. https://doi.org/10.1093/nar/gkm741
- https://www.youtube.com/watch?v=TNKWgcFPHqw&t=52
- Altieri, A. S., & Kelman, Z. (2018). DNA sliding clamps as therapeutic targets. Frontiers in Molecular Biosciences, 5, 87. https://doi.org/10.3389/fmolb.2018.00087
- Georgescu, R. E., Yurieva, O., Kim, S.-S., Kuriyan, J., Kong, X.-P., & O’Donnell, M. (2008). Structure of a small-molecule inhibitor of a DNA polymerase sliding clamp. Proceedings of the National Academy of Sciences, 105(32), 11116–11121. https://doi.org/10.1073/pnas.0804754105
- Russell, P. J. (2010). DNA replication. In iGenetics: A molecular approach (3rd ed., pp. 43–72). Pearson Benjamin Cummings.
