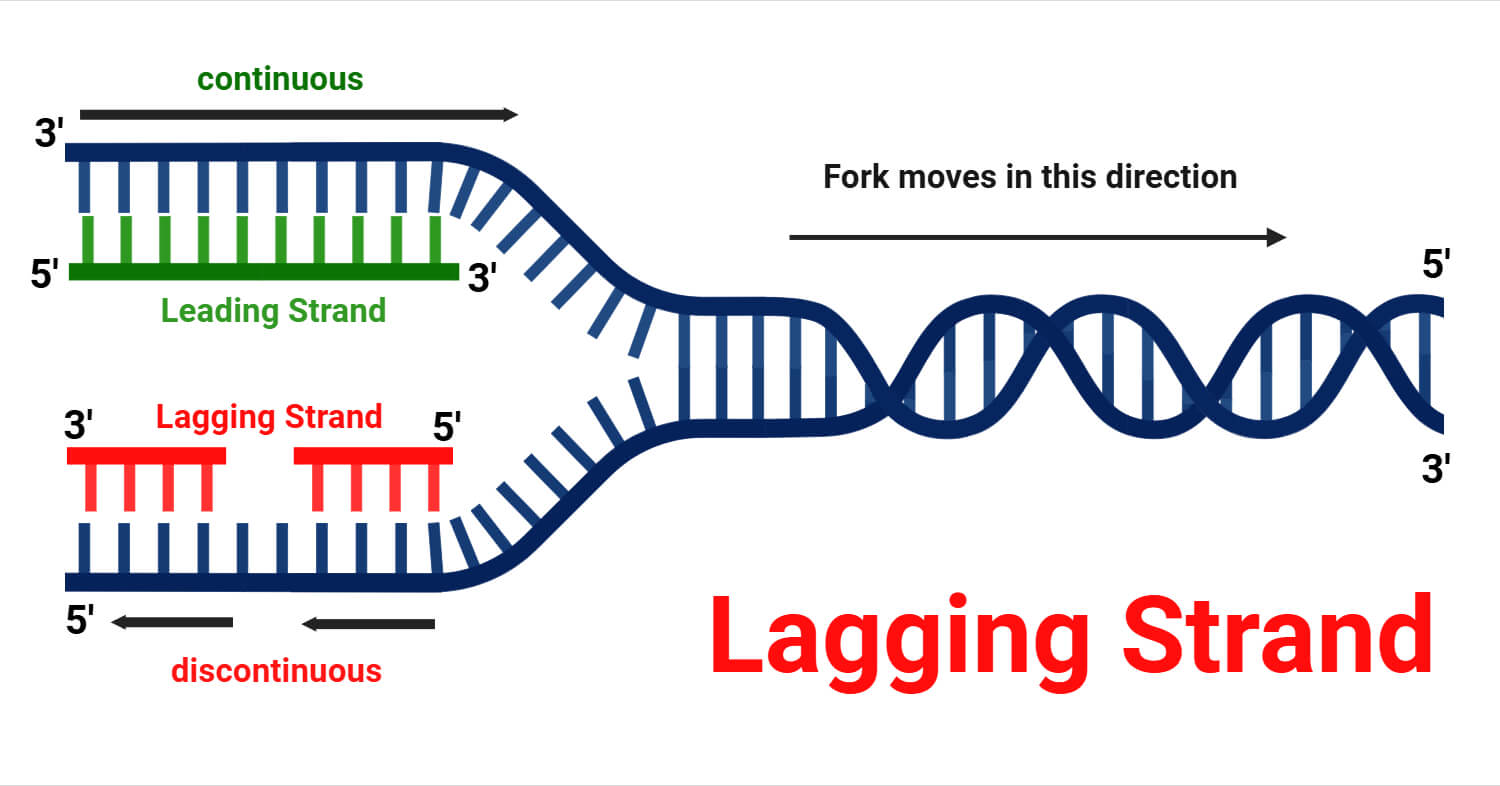
The lagging strand is the nascent daughter DNA that is assembled discontinuously in the opposite direction of the unwinding replication fork. Due to the antiparallel nature of DNA, its template is oriented 5′ → 3′, forcing DNA Polymerase, which can only build in a 5′ → 3′ direction, to replicate the strand in a backstitching motion.

As Helicase unzips the double helix, the lagging strand is formed in short, separate stretches known as Okazaki fragments. Unlike the leading strand, which needs only one starter, each of these fragments requires its own RNA primer to begin. After synthesis, the temporary primers are excised, the resulting gaps are filled with DNA, and DNA Ligase seals the fragments into a single, cohesive backbone.
The Directionality Dilemma: Why Synthesis Must Be Discontinuous
The Directionality Dilemma arises from two non-negotiable rules of molecular biology: the antiparallel nature of DNA and the strict 5→3′ biosynthetic requirement of DNA Polymerase. Because of this, a new strand can only grow in the 5’→3′ direction. As the DNA double helix unzips at the replication fork, one strand (the leading strand) is oriented perfectly for the polymerase. The enzyme can follow directly behind the unzipping machinery, building a single, long, continuous chain of DNA in the same direction the fork is opening. The other strand is incongruent because its orientation is upside down relative to the fork’s movement. To maintain the 5′ → 3′ chemical rule, the polymerase must move away from the opening fork.
To solve this, the cell resorts to a backstitching technique:
- Discontinuous Synthesis: Instead of one long piece, the strand is built in short, separate bursts.
- Okazaki Fragments: These are the small segments of DNA produced during these bursts.
- DNA Ligase: Once the fragments are completed, this enzyme acts as molecular glue to stitch the pieces together into a finished strand.
The directionality dilemma is essentially a trade-off for accuracy. Because DNA polymerase only works in the 5′ →3′ direction, it can perform proofreading without stalling the entire process. In this orientation, the energy required to form the chemical bond is provided by the incoming nucleotide’s own triphosphate group. If an incorrect base is added, the enzyme can snip it out and simply try again with a new, energized nucleotide.
In contrast, if a hypothetical enzyme synthesized DNA in the 3’→ 5′ direction, the energy for the bond would be stored at the growing tip of the DNA strand itself. Removing a mismatched base in that scenario would strip the energy away from the attachment site, leaving a dead end that could no longer accept new nucleotides.
Okazaki Fragments: The Building Blocks of the Lagging Strand
During replication, on the lagging strand, short newly synthesized DNA sequences are formed. These are formed discontinuously while the leading strand is copied in one long, unbroken chain. Since the two strands of the DNA double helix are antiparallel, only one strand (the leading strand) can be built continuously as the replication fork opens. The other strand, known as the lagging strand, must be synthesized in the opposite direction. To solve this, the cell uses a backstitching mechanism to create short, discrete segments of DNA known as Okazaki fragments.
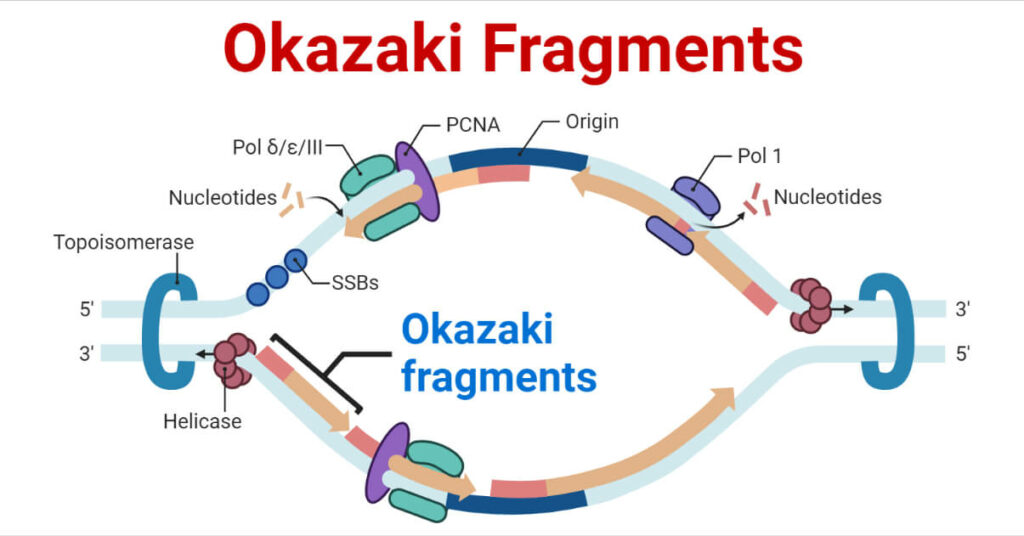
Formation of Okazaki fragments:
- Priming: Since DNA polymerase cannot start from scratch, an enzyme called primase lays down a short RNA primer.
- Synthesis: DNA polymerase III extends from this primer, moving away from the replication fork, creating a fragment typically 1,000 to 2,000 nucleotides long in bacteria (shorter in eukaryotes).
- Ligation: Once the fragment is complete, DNA polymerase I removes the RNA primer and replaces it with DNA. Finally, DNA ligase acts as the glue, sealing the nicks between fragments to create a continuous strand.
Without Okazaki fragments, the lagging strand could not be replicated accurately. These fragments ensure that even though the replication machinery moves in one physical direction, both strands of the genetic code are copied simultaneously and with high fidelity.
The Role of RNA Primase: Starting the Process Repeatedly
DNA replication relies on a sophisticated backstitching mechanism to overcome the directional constraints of DNA polymerase, which can only synthesize in the 5’→3′ direction. On the lagging strand, this results in the creation of Okazaki fragments.
The process is initiated by RNA primase, a specialized enzyme composed of a zinc-binding domain for template recognition and a catalytic core for nucleotide assembly. Because DNA polymerase cannot start a chain from scratch, primase repeatedly synthesizes short RNA primers to provide the necessary 3′-OH attachment point.
As the replication fork opens, DNA polymerase extends these primers into discrete DNA segments. These fragments are later processed by DNA polymerase I which removes the RNA primers and fills the gaps with DNA, while DNA ligase acts as molecular glue to seal the phosphodiester backbone. This coordinated effort ensures that both antiparallel strands are copied simultaneously, maintaining genomic integrity with remarkable precision.
Elongation and Replacement: DNA Polymerase III vs. DNA Polymerase I
DNA replication requires the coordinated activity of specialized enzymes to synthesize the lagging strand through a process of discrete fragment production and maturation. RNA primase, characterized by a zinc-binding domain and a catalytic core, initiates synthesis by generating short RNA primers that provide the required 3′-OH terminus. DNA polymerase III, a multi-subunit holoenzyme stabilized by a β-sliding clamp, then extends these primers into Okazaki fragments. Because Pol III lacks the ability to remove RNA, DNA polymerase I utilizes its unique 5’→3′ exonuclease domain to excise the primers while simultaneously filling the gaps with deoxyribonucleotides.
The structure of DNA polymerase I, consisting of a single polypeptide with distinct domains for polymerization and exonuclease activity, allows for this simultaneous nick translation. Once the RNA is fully replaced by DNA, DNA ligase catalyzes the formation of the final phosphodiester bond to join the fragments. This enzymatic sequence ensures that the discontinuous synthesis on the lagging strand results in a continuous, high-fidelity DNA molecule.
DNA Ligase: Sealing the Nicks in the Backbone
Acting as the final architect of the replication process, DNA ligase ensures the structural integrity of the genome by converting fragmented sequences into a unified strand. Its physical structure comprising a DNA-binding domain, an adenylation domain, and a flexible OB-fold enables it to wrap entirely around the DNA double helix to identify breaks in the sugar-phosphate backbone known as nicks.
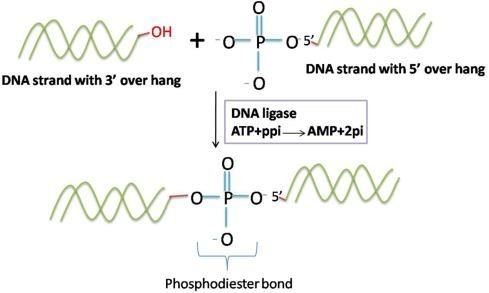
The enzyme facilitates a sophisticated chemical reaction that relies on either ATP (in eukaryotes) or NAD+ (in bacteria) for energy. The process begins when the ligase activates the 5′-phosphate group at the nick site, which subsequently allows the neighboring 3′-hydroxyl group to perform a nucleophilic attack. This results in the formation of a covalent phosphodiester bond. By permanently linking Okazaki fragments, DNA ligase eliminates the gaps left by the replication machinery, establishing a continuous and stable DNA molecule ready for cell division.
Leading Strand vs. Lagging Strand: A Detailed Comparison
| Feature | Leading strand | Lagging strand |
| Manner of Synthesis | Continuous: Synthesized as one long, uninterrupted polymer. | Discontinuous: Built in short bursts known as Okazaki fragments. |
| Direction of Growth | Forward: Moves toward the advancing replication fork. | Backward: Moves away from the replication fork. |
| Template Polarity | 3′ → 5′(as read by the polymerase). | 5’→3′ (as read by the polymerase). |
| RNA Primase Role | Minimal; used once at the origin. | High; used repeatedly for every fragment. |
| Primary Builder | DNA Pol III: Stays attached (high processivity). | DNA Pol III: Must constantly detach and re-attach. |
| Replacement Enzyme | DNA Pol I: Minimal involvement. | DNA Pol I: Essential for removing RNA primers and filling gaps. |
| Backbone Sealing | DNA Ligase: Minimal involvement. | DNA Ligase: Essential for joining Okazaki fragments together |
| Complexity | Simple, requires one loading event. | High, requires a looping mechanism (Trombone Model). |
The Trombone Model: Coordination at the Replication Fork
The Trombone Model describes the sophisticated mechanical coordination required to synthesize both the leading and lagging strands simultaneously at a single replication fork. Because the two strands are antiparallel, the lagging-strand template must physically loop backward through the DNA polymerase III holoenzyme complex. This creates a flexible, expanding loop of single-stranded DNA that grows as the polymerase synthesizes an Okazaki fragment.
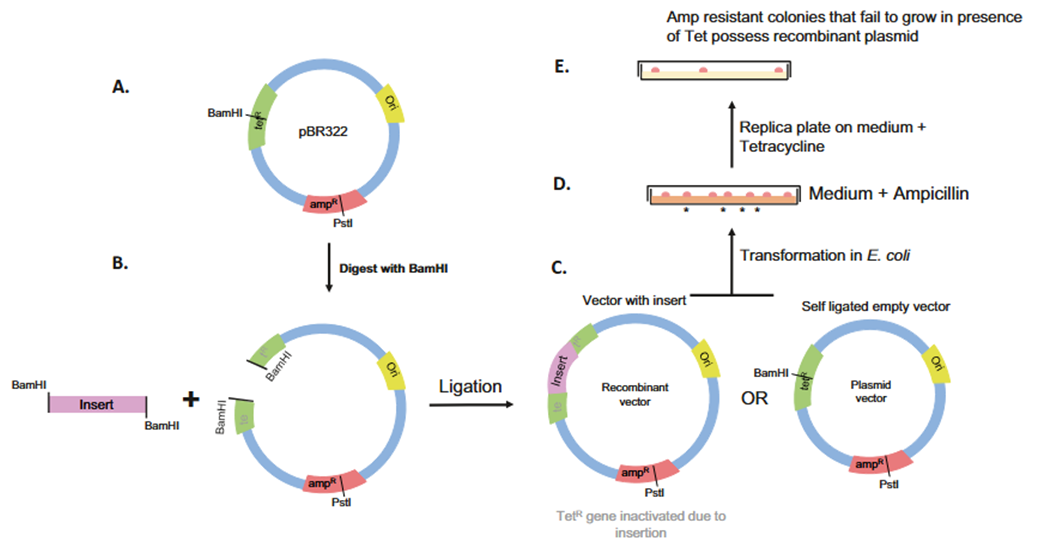
https://www.researchgate.net/figure/Model-of-T7-DNA-replicationThe-leading-strand-template-yellow-is-copied-continuously_fig1_38099857
As the replication fork progresses, the loop extends until the polymerase completes the fragment and hits the previous RNA primer. At this point, the slide of the trombone is released; the polymerase dissociates from the finished fragment and reattaches to a new primer closer to the fork, resetting the loop.
This cyclical looping and releasing reminiscent of the sliding motion of a trombone allows the entire replisome to move as a single, unified machine in the direction of the unwinding helicase. This ensures that despite the discontinuous nature of lagging strand synthesis, both strands are copied at the same overall rate without the replication fork stalling.
The End-Replication Problem: Telomeres and the Lagging Strand
The End-Replication Problem is a biological glitch inherent to linear chromosomes. It arises because DNA polymerase is structurally limited, that is it can only synthesize DNA in a 5’→3′ direction and requires an existing primer to begin. When the final RNA primer at the very tip of the lagging strand is removed, DNA polymerase cannot fill the resulting gap because there is no upstream 3′-OH group to attach to. Consequently, each round of replication leaves the new DNA strand slightly shorter than the original template. To prevent the loss of essential genetic data, chromosomes are capped with telomeres long, repetitive non-coding sequences (typically TTAGGG). These act as a protective buffer, ensuring that the inevitable shrinkage only consumes sacrificial DNA rather than vital genes. When telomeres become critically short, cells enter senescence and stop dividing, a process known as the Hayflick Limit. However, telomerase, a specialized enzyme containing its own RNA template can extend these ends.

Clinical Significance: Targeting Replication Enzymes in Disease
Targeting the enzymes responsible for DNA replication allows us to disrupt the growth of cancer, viruses, and bacteria with high precision.
- Topoisomerase I which prevents the DNA from supercoiling can be inhibited causing lethal DNA breaks that kill rapidly dividing tumour cells.
- Several drugs which act as a Decoy of nucleotides but lack the critical 3′-OH group acts as an inhibitor of DNA polymerase
- Telomerase Inhibition for Hematologic Malignancies Blocking the enzyme that maintains chromosome tips strips cancer cells of their immortality, forcing them into cellular senescence and death.
- Targeting the bacterial-specific enzyme DNA Gyrase, stops the untangling of the chromosome, leading to rapid cell death by stalling replication.
- Topoisomerase II Inhibition Prevents the resealing of DNA after the enzyme makes a necessary cut, leading to overwhelming chromosomal damage in cancer cells.
- Reverse Transcriptase Inhibition Blocks the viral enzyme from converting RNA into DNA, preventing retroviruses like HIV from integrating into the host’s genome.
Conclusion
DNA replication is defined by the directionality dilemma because DNA polymerase only synthesizes 5′ to 3′, the antiparallel lagging strand must be built discontinuously. Using a backstitching technique called the Trombone Model, the cell creates a loop to produce short Okazaki fragments.
This process requires a specialized enzymatic sequence RNA Primase that starts the fragments, DNA Pol III extends them, DNA Pol I replaces the primers, and DNA Ligase seals the backbone. However, this method leaves a gap at the chromosome tips the End-Replication Problem. Telomeres act as sacrificial buffers to protect vital genes from this progressive shortening.
Clinically, these enzymes are high-value targets. Inhibiting DNA Gyrase kills bacteria, while blocking Reverse Transcriptase stops viruses like HIV. Similarly, targeting Topoisomerases or Telomerase allows for the precise destruction of immortal cancer cells.
References
- Georgescu, R. E., Langston, L. D., Yao, N. Y., Yurieva, O., & O’Donnell, M. E. (2014). Replisome mechanics: Lagging strand events that influence speed and processivity. Nucleic Acids Research, 42(10), 6497–6510. https://doi.org/10.1093/nar/gku257
- Maslowska, K. H., Makiela-Dzbenska, K., Mo, J. Y., Fijalkowska, I. J., & Schaaper, R. M. (2018). High-accuracy lagging-strand DNA replication mediated by DNA polymerase dissociation. Proceedings of the National Academy of Sciences (PNAS), 115(17), 4437–4442. https://doi.org/10.1073/pnas.1720353115
- Balakrishnan, L., & Bambara, R. A. (2013). Okazaki fragment metabolism. Cold Spring Harbor Perspectives in Biology, 5(2), a010173. https://doi.org/10.1101/cshperspect.a010173
- Yao, N. Y., & O’Donnell, M. E. (2008). Replisome dynamics and use of DNA trombone loops to bypass replication blocks. Molecular BioSystems, 4(11), 1075–1084. https://doi.org/10.1039/b811097b
- Russell, P. J. (2010). DNA replication. In iGenetics: A molecular approach (3rd ed., pp. 43–72). Pearson Benjamin Cummings
- Nasheuer, H. P., & Onwubiko, N. O. (2023). Lagging strand initiation processes in DNA replication of eukaryotes—Strings of highly coordinated reactions governed by multiprotein complexes. Genes, 14(5), 1012. https://doi.org/10.3390/genes14051012
- Ye, T., Yuan, Q., Wu, S., Zhao, J. T., Wu, Z. J., Liu, J. C., Wu, W., & Zhou, J. Q. (2025). Asymmetrical end structures of leading and lagging telomeres in Saccharomyces cerevisiae dictate the nature of the end replication problem. Cell Reports, 44(12), 116622. https://doi.org/10.1016/j.celrep.2025.116622
