Haemophilus influenzae is a Gram-negative coccus-bacillus that was first isolated by Richard Pfeiffer during the influenza pandemic of 1890.
It was previously called Pfeiffer’s bacillus or Bacillus influenzae. It was mistakenly thought to be the cause of the disease influenza, and it was named accordingly. More than 19 Haemophilus species have been found so far. Some of them are Haemophilus haemolyticus, Haemophilus parainfluenzae, Haemophilus ducreyi, Haemophilus aprophilus, Haemophilus influenzae, and so on.
H. influenzae is the major pathogen of the genus Haemophilus. It may be either capsulated (typeable) or non-capsulated (non-typeable). Based on the capsular polysaccharide, it has six serotypes. Similarly, based on three biochemical tests, i.e., indole production, urease activity, and ornithine decarboxylase activity, H. influenzae is classified into eight biotypes. The majority of clinical isolates are biotypes I, II, and III; invasive Hib strains are biotype I.
Classification of Haemophilus influenzae
Table 1: Classification of Haemophilus influenzae
| Kingdom | Bacteria |
| Phylum | Proteobacteria |
| Class | Gamma Proteobacteria |
| Family | Pasteurellaceae |
| Genus | Haemophilus |
| Species | influenza |
Morphology and Microscopy of Haemophilus influenzae
- Gram-negative pleomorphic or coccobacilli
- Small in size, usually 0.3 μm to 1 μm
- Long, thread, and occasionally filamentous
- Non motile
- Non sporing
- Aerobic and Facultative anaerobe
- Capsulated (capsules play a major role in pathogenesis)
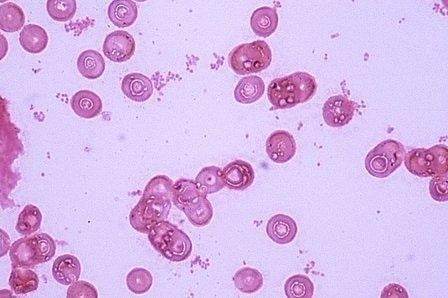
Image source:
https://microbiologylearning.weebly.com/gram-negative-coccobacilli.html
Cultural and Growth Characteristics of Haemophilus influenzae
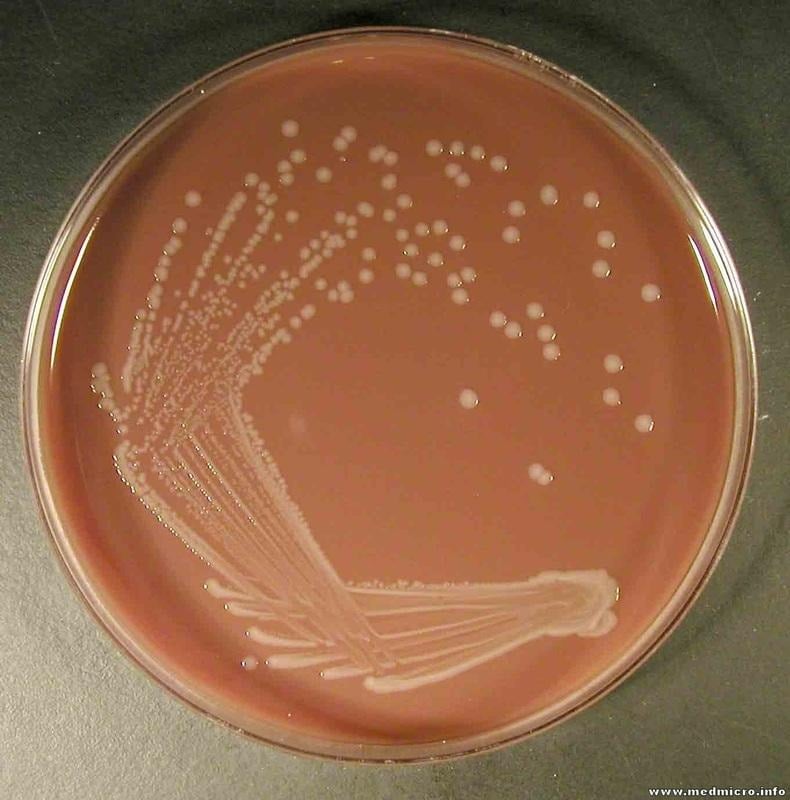
- H. influenzae grows well in chocolate agar, where colonies are moist, smooth, and gray.
- It produces colonies of 0.5mm in diameter on Horse Blood Agar when incubated at 37°C for 24 hrs.
- Also, when subcultured on a transparent medium such as Levinthal agar, colonies of encapsulated strains show iridescence. It appears in different shades of red, orange, green, and blue that change with the angle of observation.
- Opaque, mucoid, and smooth kinds of colonies are formed by the capsulated strains, and the diameter observed is 3-4mm.
- Fastidious and facultative anaerobes can also grow in 5% CO2.
- The optimum temperature needed for H. influenzae to grow is 35-37°C, and the optimum pH is 7.6.
Biochemical and Identification Tests of Haemophilus influenzae
Table 2: Biochemical tests for identification
| Test | Results |
| Catalase | Positive |
| Oxidase | Positive |
| Indole | Varies |
| Urease | Varies |
| Glucose | Positive |
| Fructose | Negative |
| Xylose | Positive |
| Lactose | Negative |
Virulence Factors and Pathogenesis of Haemophilus influenzae
Some of the virulence factors and their roles in pathogenesis are described below.
- Capsule:
In Hib, the capsular polysaccharide is composed of phosphoribosylribitol phosphate (PRP), which has antiphagocytic properties and is highly associated with virulence.
- Membrane Lipooligosaccharide:
It is responsible for bacterial attachment, invasiveness, and paralysis of ciliated respiratory epithelium.
- There are involvement of other cell envelope factors too in facilitating the attachment to host cells.
- Also, secretory antibodies are probably inactivated by IgA protease.
- Pili and other cell factors help in attachment to host cells.
The pathogenesis of H. influenzae involves two phases. They are:
- Portal of entry: Upper respiratory tract, especially the nasopharynx, is the entry route of the H. influenzae (usually all strains).
- Dissemination: When H. influenzae enters the body, its type b penetrates the nasopharyngeal epithelium. It either spreads through the bloodstream or directly to the meninges. It can spread from the sinuses to the meninges and is second only to Streptococcus pneumoniae as the most common cause of recurrent meningitis. People who suffer from chronic otitis media, head injury, or CSF leak are highly at risk of getting infected by H. influenzae.
Epidemiology and Transmission of Haemophilus influenzae
Usually, reservoirs of H. influenzae are humans. There is no animal reservoir, so we can say that H. influenzae are solely a human pathogen. Non-encapsulated strains of H. influenzae are carried by around 60-80% of healthy children, and by around 35% of healthy adults. Asymptomatic carriers of capsulated H. influenzae type b in healthy children are around 2-3%. It can be transmitted from person to person through contaminated respiratory droplets. Also, certain infections can be caused by a person’s endogenous strains.
Clinical Manifestations of Haemophilus influenzae
The diseases that are caused by H. influenzae can be divided into two groups, i.e., invasive and noninvasive. The common invasive and non-invasive groups are described below:
Invasive infection
It includes diseases such as meningitis, acute epiglottitis, bacteremia, suppurative lesions, and pneumonia. A few of them are described below:
- Meningitis: Most common in children between 6 months and 2 years of age.
- Acute epiglottitis: It is an acute inflammation of the epiglottis. The swelling of cellulitis and supraglottic tissues are seen, which makes it one of the life-threatening diseases that are found in small children (age 2-3 years).
- Bacteraemia: Children who become febrile may be infected with H. influenzae type b, which can cause bacteraemia. When these bacteria are transmitted from the blood, it may lead to suppurative lesions. Sometimes, neonates can also be infected by non-capsulated H. influenzae, causing septicaemia and meningitis.
- Pneumonia: Lobar pneumonia can be caused by this bacterium, mainly in older patients, chronic alcoholics, and smokers, etc.
Non-invasive infections
Many local infections, such as otitis media, sinusitis, and conjunctivitis, can be caused by both encapsulated and non-encapsulated strains of H. influenzae.
Laboratory Diagnosis of Haemophilus influenzae
The laboratory diagnosis of H. influenzae are listed below:
Specimens
Specimens depend on the site of infection and include CSF, Nasopharyngeal swab, Blood, Pus, Sputum, Urine (rarely in case of UTI), and Joint aspirates.
Note: All the specimens should be kept at room temperature and processed within 10 minutes of collection.
Direct microscopy
Gram stain: Gram-negative coccobacilli
- Gram-stained smear of sediment obtained after centrifugation of CSF at 2000rpm for 10 min reveals that H. influenzae.
- Capsules of encapsulated strains are not easily seen on these tiny organisms, except serologically by the Quellung reaction.
Culture:
- Specimens should be cultured as soon as possible.
- It grows better aerobically (CO2-enriched) than anaerobically.
- Culture media that can be used for H. influenzae are Chocolate agar, Levinthal agar, Levinthal broth, BHI agar supplemented with X and V factor, and BHI broth supplemented with X and V factor.
- After inoculating the specimen into CA, the plate should be incubated in a 5-10% CO2-enriched atmosphere (commonly using a candle jar) at 37°C for 24 hours.
- It gives round, convex colonies with an opaque zone near the centre.
Satellitism phenomena:
- H. influenzae requires X and V factors for its growth.
- X factor is for the synthesis of respiratory enzymes like: cytochrome, catalase and peroxidase, a protoporphyrin IX (haematin, a precursor of haemin) is required.
- V factor- NAD is used as an electron carrier in the organism’s oxidation-reduction system.
- Blood agar, which is commonly used, contains X factor but lacks V factor because intact cells are present in the medium.
- When blood agar is heated to 75°C, more V factors are released, so chocolate agar is best for the growth of this organism.
- H. influenzae produces colonies on horse BA of about o.5mm in diameter. However, if S. aureus, which produces V factor in excess of its own need, is cultured on a BA plate with H. influenzae, other substances and V factor produced by S. aureus help the growth of H. influenzae, which forms the basis of the satellitism test, which is the simplest way of recognizing H. influenzae regularly.
- S. pneumoniae also produces V factor and causes H. influenzae to show satellism.
- A loopful of suspected Haemophilus colonies is mixed in about 2ml of sterile peptone water or normal saline.
- Then, using a sterile swab, inoculate the organism suspension onto plates of NA and BA.
- A pure culture of S. aureus is streaked across each inoculated plate.
- Both plates are incubated in CO2 enriched atmosphere at 35-37°C overnight.
- If there is growth on the BA plate but not on the NA plate, and colonies near the columns of S. aureus growth are larger than those farthest from it, then it is assumed to be H.influenzae.
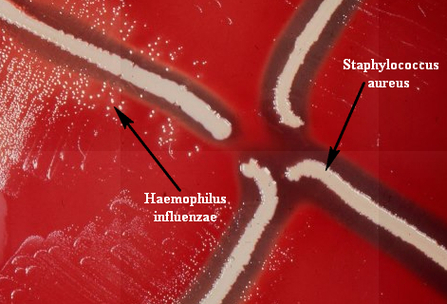
https://microbiologylearning.weebly.com/gram-negative-coccobacilli.html
Table 3: Satellitism Phenomenon
| Haemophilus | X factor | V factor | Xylose |
| H. influenzae | positive | positive | positive |
| H. aegyptius | positive | positive | negative |
| H. parainfluenzae | negative | positive | negative |
Quelling Test
- The test is performed by placing a drop of CSF, broth, or saline suspension of the test organism on each end of a slide.
- A loopful of H. influenzae type b antiserum is mixed with one drop of saline.
- Both preparations are then covered with a coverslip and viewed under oil immersion.
- The organism can be identified as Hib if rods or coccobacilli surrounded by clearly defined capsules are seen in the prepared antiserum when compared with a saline preparation.
- Because the antibody is deposited on the bacterial cell wall, the capsules are seen in the antiserum preparation.
- If each oil-immersion field contains 10-50 orgs, the best reactions are obtained.
Antigen detection in Haemophilus influenzae
Type b polysaccharide antigen (PRP capsules) of H. influenzae in body fluids (e.g., CSF or pus) can be detected using latex agglutination and coagglutination. Less than 1 ng/mL of PRP can be detected by latex agglutination tests. Some molecular techniques, such as the polymerase chain reaction, are widely used today for detecting H. influenzae.
Treatments of Haemophilus influenzae Infections
Antibiotics such as Ampicillin, Chloramphenicol, Cefotaxime, Ceftazidime, etc. are effective against H. infuenzae. However, 25-30% of invasive strains showed resistance to most antibiotics, so Antibiotic susceptibility testing must be performed before prescribing. Cephalosporins and Chloramphenicol (third generation) can be used in case of penicillin resistance.
Prevention and Control of Haemophilus influenzae Infections
- PRP-conjugate vaccines (capsular antigen) can provide long-term protection.
- The major control measures should be used by healthcare professionals by strictly following hospital safety protocols.
- Contact between infected patients and other individuals should be maintained.
Conclusion
In conclusion, H. influenzae has a major clinical impact due to its virulence, persistence, and antibiotic resistance, particularly in vulnerable patient groups; therefore, it is an important gram-negative coccobacillus that warrants our attention.
References
- Khattak ZE, Anjum F. Haemophilus influenzae Infection. [Updated 2023 Apr 27]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK562176/
- Mardh P A, Taylor-Robinson D eds 1984 Bacterial Vaginosis. Almqvist and Wiksell.
- Jordens J Z, Slack MPE 1995 Haemophilus influenzae then and now. Eur J Clin Microbiol 14:935-948.
