The function of biological macromolecules is inherently tied to their precise three-dimensional structure. To accurately characterize these higher-order folds, particularly the secondary structures of proteins and polypeptides, biochemists rely on highly sensitive spectroscopic techniques. Circular Dichroism (CD) spectroscopy is one such essential analytical tool.
Circular Dichroism (CD) spectroscopy technique measures the differential absorption between left- and right-circularly polarized light, a phenomenon intrinsically linked to molecular chirality. Given that major biomolecules like proteins and DNA are inherently chiral, their unique CD signatures provide a powerful means of identification and structural analysis.
Furthermore, CD spectroscopy is invaluable for studying the stability of these structures. By monitoring structural loss upon heating, CD enables precise characterization of Thermal Denaturation (TD), yielding crucial parameters such as the denaturation transition midpoint, which serves as a metric of protein stability.
What is Circular Dichroism (CD)?
Circular dichroism is an extremely sensitive phenomenon to the secondary structure of proteins and polypeptides. One type of light absorption spectroscopy, circular dichroism (CD) spectroscopy, compares the absorbance of left- and right-circularly polarized light. The result of polarized light’s interaction with chiral molecules is circular dichroism. Most compounds
found in biology are chiral. For example, the higher structures of proteins, DNA, and RNA, as well as a variety of other physiologically significant molecules, are chiral, as are 19 of the 20 common amino acids that make up proteins. Circular dichroism is an excellent analytical tool for biological molecules because of their highly chiral chemistry. As such, studying biological molecules is the primary use for this approach. The study of the higher-order structures of chiral macromolecules like proteins and DNA is a significant portion of the application of circular dichroism in biochemistry. This is because the CD spectrum of a protein or DNA molecule is heavily impacted by the macromolecule’s three-dimensional structure rather than being the sum of the CD spectra of its individual residues or nucleotides. Chiral macromolecules can be identified by their unique circular dichroism signatures, which can also be used to track structural alterations. The several secondary structural components of proteins, like the α-helix and the β sheet, are the most extensively researched circular dichroism signatures. This is understood to the extent that percentages of each secondary structural element in a protein’s structure may be predicted from CD spectra in the far-UV (below 260 nm). Other names for a circular dichroism spectrophotometer are circular dichroic graph or circular dichroism spectropolarimeter.
Instrumentation for Circular Dichroism (CD)
- Light Source: Typically operating in the visible or ultraviolet (UV) portions of the electromagnetic spectrum, the light source generates unpolarized light at a variety of wavelengths. Xenon arc lamps, deuterium lamps, and tungsten-halogen lamps are examples of common light sources.
- Monochromator: The monochromator turns the unpolarized light into polarized light by choosing a particular wavelength from the wide range that the light source emits. It enables accurate adjustment of the light wavelength utilized in the CD measurements.
- Photoelastic modulator: A source of monochromatic linearly polarized light can be converted into either left- or right-circularly polarized light by passing it through a quarter-wave plate, whose unique axis is at 45 degrees to the linear polarization plane. This is how the majority of modern circular dichroism instruments work. A circular dichroism spectrophotometer uses a specific optical element known as a photo-elastic modulator (PEM) in place of a static quarter-wave plate.
- Sample holder/CD medium active: holds the sample to be analyzed. It typically consists of a cuvette holder where the sample solution is placed. The cuvette is usually made of quartz or other optically transparent materials.
- Detector: A light detector is located across from the sample place. If the light path contains no circularly dichroic sample, the amount of light reaching the detector remains constant. If a circularly dichroic sample is present in the light path, the recorded light intensity for the left and right CPL will change.
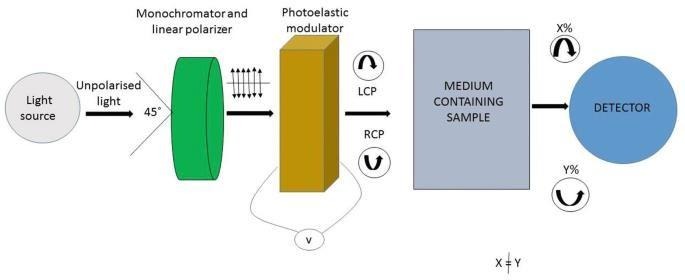
https://link.springer.com/chapter/10.1007/978-981-16-4550-1_2
Circular dichroism (CD) spectroscopy instrumentation includes a UV-visible light source, a monochromator to select specific wavelengths, and a polarizer combined with a modulator to produce alternating left- and right-circularly polarized light. The light passes through a sample in a temperature-controlled holder, and a detector measures the differential absorption, with data processed to generate a CD spectrum.
What is Thermal Denaturation (TD)?
One significant physical characteristic of proteins is their thermal stability. Using circular dichroism (CD) spectroscopy is an extremely popular technique for studying it. Thermal denaturation using circular dichroism (CD) spectroscopy is a technique employed to study the stability and unfolding transitions of proteins or other biomolecules as a function of temperature.
Protein denaturation is the structural alteration of a protein, resulting in the disruption of its native conformation and functional properties. This process involves the unfolding of the protein’s secondary, tertiary, and quaternary structures. Denaturation typically entails the disruption of non-covalent interactions, including hydrogen bonds, hydrophobic interactions, van der Waals forces, and electrostatic interactions. Since peptide bonds are not hampered by this, the primary structure remains undisturbed.
The process of a protein losing its biological activity and experiencing structural unfolding as a result of exposure to high temperatures is known as “thermal denaturation”. The process of heat denaturation entails the interference of non-covalent interactions that maintain the native structure of the protein. Hydrophobic interactions, van der Waals forces, electrostatic interactions, and hydrogen bonding are some examples of these interactions. Higher temperatures cause a rise in the kinetic energy of at crucial temperature, called the melting or denaturation temperature (Tm), is reached at which the energy released by the high temperature is more than the energy needed to break the stabilizing interactions that retain the protein in its native form.
Consequently, the protein starts to unfold, changing from a folded condition to a more disordered, denatured state. Plotting the thermal denaturation process versus temperature usually results in a sigmoidal curve, with the rate of denaturation rising gradually as the temperature gets closer to the Tm. The Tm, or denaturation transition midpoint, is a crucial metric for describing the thermal stability of proteins. Protein molecules, which increase molecular motion and vibrational energy.
The temperature at which 50% of a protein remains in its native structure, and the other 50% is in its maximal denatured or unfolded conformation, is known as the transition midpoint (Tm). The higher the Tm, the more stable the protein.
Instrumentation for Thermal Denaturation (TD)
While the CD spectrometer functions as the primary detection system, performing thermal denaturation requires an integrated thermoelectric assembly to execute precise, programmable shifts in the sample’s physical state.
- Peltier Temperature Controller: The core of TD instrumentation is the Peltier element. Unlike traditional water baths, a Peltier heater/cooler uses the thermoelectric effect to transfer heat between two different electrical junctions. It allows for rapid and highly stable temperature ramps (typically 0.1°C to 2.0°C per minute). Most modern units can span from -10°C to 110°C, allowing for the study of both cold and heat denaturation.
- Sample Cell & Stirring Mechanism: The cell holder usually includes a magnetic stirrer at the base. A tiny flea (stir bar) inside the quartz cuvette ensures the buffer temperature is homogenous. For TD, specialized capped quartz cuvettes are used to prevent evaporation at high temperatures, which would otherwise change the protein concentration and refractive index. Uniform heat distribution is critical to avoid hot spots that could skew the melting curve.
- Internal Temperature Monitoring: A fine, teflon-coated thermocouple probe is inserted directly into the sample buffer (away from the light path). This probe provides the software with the True Temperature, ensuring the ellipticity data is plotted against the actual sample temperature rather than the instrument’s setting.
- Heat Dissipation: Peltier elements generate significant waste heat while cooling or maintaining high temperatures, the system must be connected to a circulating water bath or a heat exchanger. This prevents the spectrometer’s internal components from overheating during long melt cycles.
Comparing CD Scans and TD Melt Curves
The fundamental distinction between these methods is not found in the hardware, but in the experimental dimension being probed. While the physical components remain identical, the software reconfigures the instrument’s operational logic to prioritize temperature-dependent transitions over wavelength-specific signals.
| Feature | Circular Dichroism (CD) Scan | Thermal Denaturation (TD) Melt |
| Primary Variable | Wavelength (λ): Scanning the UV spectrum (e.g., 190–260). | Temperature (T): Ramping heat (e.g., 20°C to 95°C |
| Data Output | Spectral Fingerprint: A snapshot of the fold at a specific temperature. | Transition Profile: A movie of the unfolding process over time. |
| Hardware State | Monochromator is active, Peltier is static. | Monochromator is fixed; Peltier is active. |
| Physical Insight | Conformation: Identifies if the protein is α-helical, β-sheet, or disordered. | Thermodynamics: Quantifies the energy barrier required to unfold the molecule. |
| Key Metric | Molar Ellipticity (θ): Specific to the protein’s secondary structure. | Melting Point (Tm): The midpoint of the structural collapse. |
| Practical Use | Checking if a protein is correctly folded after purification. | Comparing the shelf-life or toughness of different formulations. |
Interpreting the Data
The Sigmoidal Curve
A typical melt results in a sigmoidal (S-shaped) curve.
- The Pre-transition Baseline: The initial flat or slightly sloped region where the protein is fully folded and stable.
- The Transition Region: The steep part of the curve where the protein begins to unfold rapidly.
- The Post-transition Baseline: The final region where the protein is fully denatured (unfolded).
Determining the Melting Temperature (Tm)
The Tm is the most critical value derived from CDTD. It is the temperature at the midpoint of the transition region, where exactly 50% of the protein molecules are folded and 50% are unfolded.
- Higher Tm = Greater thermal stability.
- Calculation: This is usually found by taking the first derivative of the ellipticity with respect to temperature (dθ/dT). The peak of this derivative curve is the Tm.
Assessing Reversibility
After reaching the maximum temperature (e.g., 95°C), the instrumentation can be set to cool down back to 20°C.
- If the CD signal returns to its original state, the folding is reversible
- If the signal remains flat, the protein has undergone irreversible aggregation (common in many enzymes).
Applications
- Screens various buffers, pH levels, and excipients to determine the optimal conditions for long-term biopharmaceutical storage.
- Serves as a conformational fingerprint to ensure batch-to-batch consistency and structural identity during large-scale manufacturing.
- Compares mutant stability to wild-type structures to identify amino acid substitutions that enhance thermal robustness (Tm).
- Detects shifts in melting temperature to quantify the stabilizing effect of small molecules, ions, or drugs binding to a target protein.
- Evaluates the functional temperature range and denaturation kinetics of industrial catalysts under diverse operational stresses.
- Monitors the thermal transition of complex biomaterials and scaffolds to determine the temperature at which their ordered structure collapses.
Conclusion
In the current landscape of biophysical characterization, the ability to correlate conformational transitions with thermodynamic stability is a fundamental requirement for the successful development of biological entities. Circular Dichroism Thermal Denaturation (CDTD) serves as a critical analytical bridge in this process, offering a high-resolution window into the structural resilience of proteins and nucleic acids under thermal stress.
By synergizing the chiroptical sensitivity of the optical bench with the high-precision control of thermoelectric (Peltier) instrumentation, CDTD transcends simple structural identification. It provides a dynamic stress-strain profile that reveals the cooperative nature of molecular folding. Whether it is employed to validate the integrity of a recombinant protein, optimize the formulation of a therapeutic monoclonal antibody, or quantify the stabilizing effects of a novel ligand, the derived Melting Temperature (Tm) serves as a definitive biophysical benchmark.
References
- An introduction to circular dichroism spectroscopy. https://www.ntu.edu.sg/docs/librariesprovider12/default-document-library/circular-dichroism.pdf?sfvrsn=ec10f4d4_2
- Wallace, B. A., & Janes, R. W. (2003). Circular dichroism spectroscopy of membrane proteins. Chemical Society Reviews, 32(5), 302–311. https://doi.org/10.1039/B209934N
- Rodger A, Marrington R, Roper D, Windsor S. Circular dichroism spectroscopy for the study of protein-ligand interactions. Methods Mol Biol. 2005;305:343-64. doi: 10.1385/1-59259-912-5:343. PMID: 15940006.
- Circular Dichroism Spectroscopy. (2025, December 9). JASCO Inc. https://jascoinc.com/learning-center/theory/spectroscopy/circular-dichroism-spectroscopy/
- Mokashi, V. V., & Chaudhari, B. P. (2023). Unlocking insights into folding, structure, and function of proteins through circular dichroism spectroscopy—A short review. Applied Biochemistry and Biotechnology, 195(10), 6143–6161. https://doi.org/10.1007/s12010-023-04419-y
- Greenfield NJ. Using circular dichroism collected as a function of temperature to determine the thermodynamics of protein unfolding and binding interactions. Nat Protoc. 2006;1(6):2527-35. doi: 10.1038/nprot.2006.204. PMID: 17406506; PMCID: PMC2752288.
- Ramprakash, J & Doseeva, Victoria & Galkin, Andrey & Krajewski, W & Muthukumar, Lavanya & Pullalarevu, S & Demirkan, Elif & Herzberg, Osnat & Moult, John & Schwarz, F. (2008). Comparison of the chemical and thermal denaturation of proteins by a two-state transition model. Analytical biochemistry. 374. 221-30. 10.1016/j.ab.2007.10.005.
- https://www.youtube.com/watch?v=z-EqQffymOY
